Key Points
-
Biofilms are communities of microbial cells adhered to surfaces or present at liquid–air interfaces. Cells within biofilms have properties distinct from their planktonic counterparts.
-
Candida albicans can form biofilms on a wide range of implanted medical devices and host surfaces. These biofilms serve as drug-resistant reservoirs of cells that can multiply and seed bloodstream infections.
-
C. albicans biofilms contain a mixture of yeast-form cells, pseudohyphal cells and hyphal cells surrounded by an extracellular matrix.
-
More than 50 transcriptional regulators have been implicated in biofilm formation to date, and a core network of nine regulators is required for biofilm development in both in vitro and in vivo models. Approximately 1,000 genes are targets of this network.
-
Among the factors contributing to the resistance of C. albicans biofilms to antifungal drugs are the increased expression of drug efflux pumps, the protective features of the extracellular matrix, and the existence of 'persister' cells in the biofilm.
-
C. albicans is the fungal pathogen most frequently isolated from mixed bacterial–fungal infections. These mixed-species biofilms can increase virulence and protect one or multiple species from environmental hazards.
Abstract
Candida albicans is among the most prevalent fungal species of the human microbiota and asymptomatically colonizes healthy individuals. However, it is also an opportunistic pathogen that can cause severe, and often fatal, bloodstream infections. The medical impact of C. albicans typically depends on its ability to form biofilms, which are closely packed communities of cells that attach to surfaces, such as tissues and implanted medical devices. In this Review, we provide an overview of the processes involved in the formation of C. albicans biofilms and discuss the core transcriptional network that regulates biofilm development. We also consider some of the advantages that biofilms provide to C. albicans in comparison with planktonic growth and explore polymicrobial biofilms that are formed by C. albicans and certain bacterial species.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Nobile, C. J. & Johnson, A. D. Candida albicans biofilms and human disease. Annu. Rev. Microbiol. 69, 71–92 (2015).
Davey, M. E. & O'Toole, G. A. Microbial biofilms: from ecology to molecular genetics. Microbiol. Mol. Biol. Rev. 64, 847–867 (2000).
Kolter, R. & Greenberg, E. P. Microbial sciences: The superficial life of microbes. Nature 441, 300–302 (2006).
Hall-Stoodley, L., Costerton, J. W. & Stoodley, P. Bacterial biofilms: from the natural environment to infectious diseases. Nat. Rev. Microbiol. 2, 95–108 (2004).
Wolcott, R., Costerton, J. W., Raoult, D. & Culter, S. J. The polymicrobial nature of biofilm infection. Clin. Microbiol. Infect. 19, 107–112 (2013).
Fox, E. P. & Nobile, C. J. A sticky situation: untangling the transcriptional network controlling biofilm development in Candida albicans. Transcription 3, 315–322 (2012).
Achkar, J. M. & Fries, B. C. Candida infections of the genitourinary tract. Clin. Microbiol. Rev. 23, 253–273 (2010).
Ganguly, S. & Mitchell, A. P. Mucosal biofilms of Candida albicans. Curr. Opin. Microbiol. 14, 380–385 (2011).
Kennedy, M. J. & Volz, P. A. Ecology of Candida albicans gut colonization: inhibition of Candida adhesion, colonization, and dissemination from the gastrointestinal tract by bacterial antagonism. Infect. Immun. 49, 654–663 (1985).
Kumamoto, C. A. Candida biofilms. Curr. Opin. Microbiol. 5, 608–611 (2002).
Kumamoto, C. A. Inflammation and gastrointestinal Candida colonization. Curr. Opin. Microbiol. 14, 386–391 (2011).
Calderone, R. A. & Fonzi, W. A. Virulence factors of Candida albicans. Trends Microbiol. 9, 327–335 (2001).
Pappas, P. G. et al. Guidelines for treatment of candidiasis. Clin. Infect. Dis. 38, 161–189 (2004).
Wenzel, R. P. Nosocomial candidemia: risk factors and attributable mortality. Clin. Infect. Dis. 20, 1531–1534 (1995).
Kullberg, B. J. & Oude Lashof, A. M. Epidemiology of opportunistic invasive mycoses. Eur. J. Med. Res. 7, 183–191 (2002).
Weig, M., Gross, U. & Mühlschlegel, F. Clinical aspects and pathogenesis of Candida infection. Trends Microbiol. 6, 468–470 (1998).
Chandra, J. et al. Biofilm formation by the fungal pathogen Candida albicans: development, architecture, and drug resistance. J. Bacteriol. 183, 5385–5394 (2001).
Ramage, G., Mowat, E., Jones, B., Williams, C. & Lopez-Ribot, J. Our current understanding of fungal biofilms. Crit. Rev. Microbiol. 35, 340–355 (2009).
Kojic, E. M. & Darouiche, R. O. Candida infections of medical devices. Clin. Microbiol. Rev. 17, 255–267 (2004).
Ramage, G., Martínez, J. P. & López-Ribot, J. L. Candida biofilms on implanted biomaterials: a clinically significant problem. FEMS Yeast Res. 6, 979–986 (2006).
Donlan, R. M. Biofilm formation: a clinically relevant microbiological process. Clin. Infect. Dis. 33, 1387–1392 (2001).
Tumbarello, M. et al. Biofilm production by Candida species and inadequate antifungal therapy as predictors of mortality for patients with candidemia. J. Clin. Microbiol. 45, 1843–1850 (2007).
Tumbarello, M. et al. Risk factors and outcomes of candidemia caused by biofilm-forming isolates in a tertiary care hospital. PLoS ONE 7, e33705 (2012).
Lebeaux, D., Ghigo, J. M. & Beloin, C. Biofilm-related infections: bridging the gap between clinical management and fundamental aspects of recalcitrance toward antibiotics. Microbiol. Mol. Biol. Rev. 78, 510–543 (2014).
Fox, E. P., Singh-Babak, S. D., Hartooni, N. & Nobile, C. J. in Antifungals: From genomics to resistance and the development of Novel Agents (eds Coste, A. T. & Vandeputte, P.) 71–90 (Caister Academic Press, 2015).
Andes, D. R. et al. Impact of treatment strategy on outcomes in patients with candidemia and other forms of invasive candidiasis: a patient-level quantitative review of randomized trials. Clin. Infect. Dis. 54, 1110–1122 (2012).
Douglas, L. J. Candida biofilms and their role in infection. Trends Microbiol. 11, 30–36 (2003).
Uppuluri, P. et al. Dispersion as an important step in the Candida albicans biofilm developmental cycle. PLoS Pathog. 6, e1000828 (2010). This work provides an examination of cell dispersion from biofilms, including regulation, nutrient dependence and the characteristics of dispersed cells.
Hawser, S. P. & Douglas, L. J. Biofilm formation by Candida species on the surface of catheter materials in vitro. Infect. Immun. 62, 915–921 (1994).
Nobile, C. J. & Mitchell, A. P. Genetics and genomics of Candida albicans biofilm formation. Cell. Microbiol. 8, 1382–1391 (2006).
Baillie, G. S. & Douglas, L. J. Role of dimorphism in the development of Candida albicans biofilms. J. Med. Microbiol. 48, 671–679 (1999).
Gulati, M. & Nobile, C. J. Candida albicans biofilms: development, regulation, and molecular mechanisms. Microbes Infect. 18, 310–321 (2016).
Andes, D. et al. Development and characterization of an in vivo central venous catheter Candida albicans biofilm model. Infect. Immun 72, 6023–6031 (2004). This study describes a commonly used and well-established in vivo model of biofilm formation.
Johnson, C. C., Yu, A., Lee, H., Fidel, P. L. & Noverr, M. C. Development of a contemporary animal model of Candida albicans-associated denture stomatitis using a novel intraoral denture system. Infect. Immun. 80, 1736–1743 (2012).
Nett, J. E. et al. Rat indwelling urinary catheter model of Candida albicans biofilm infection. Infect. Immun. 82, 4931–4940 (2014).
Harriott, M. M. et al. Candida albicans forms biofilms on the vaginal mucosa. Microbiology 156, 3635–3644 (2010).
Dongari-Bagtzoglou, A., Kashleva, H., Dwivedi, P., Diaz, P. & Vasilakos, J. Characterization of mucosal Candida albicans biofilms. PLoS ONE 4, e7967 (2009).
Řičicová, M. et al. Candida albicans biofilm formation in a new in vivo rat model. Microbiology 156, 909–919 (2010).
Schinabeck, M. K. et al. Rabbit model of Candida albicans biofilm infection: liposomal amphotericin B antifungal lock therapy. Antimicrob. Agents Chemother. 48, 1727–1732 (2004).
Shuford, J. A., Rouse, M. S., Piper, K. E., Steckelberg, J. M. & Patel, R. Evaluation of caspofungin and amphotericin B deoxycholate against Candida albicans biofilms in an experimental intravascular catheter infection model. J. Infect. Dis. 194, 710–713 (2006).
Ramage, G., Tomsett, K., Wickes, B. L., López-Ribot, J. L. & Redding, S. W. Denture stomatitis: a role for Candida biofilms. Oral Surg. Oral Med. Oral Pathol. Oral Radiol. Endod. 98, 53–59 (2004).
Lazzell, A. L. et al. Treatment and prevention of Candida albicans biofilms with caspofungin in a novel central venous catheter murine model of candidiasis. J. Antimicrob. Chemother. 64, 567–570 (2009).
Kakade, P., Sadhale, P., Sanyal, K. & Nagaraja, V. ZCF32, a fungus specific Zn(II)2 Cys6 transcription factor, is a repressor of the biofilm development in the human pathogen Candida albicans. Sci. Rep. 6, 31124 (2016).
Ghosh, A. K., Wangsanut, T., Fonzi, W. A. & Rolfes, R. J. The GRF10 homeobox gene regulates filamentous growth in the human fungal pathogen Candida albicans. FEMS Yeast Res. 15, fov093 (2015).
Chen, H. F. & Lan, C. Y. Role of SFP1 in the regulation of Candida albicans biofilm formation. PLoS ONE 10, e0129903 (2015).
Nobile, C. J. et al. A recently evolved transcriptional network controls biofilm development in Candida albicans. Cell 148, 126–138 (2012). This study defines the core transcriptional network regulating C. albicans biofilms.
Fox, E. P. et al. An expanded regulatory network temporally controls Candida albicans biofilm formation. Mol. Microbiol. 96, 1226–1239 (2015). This work expands the core biofilm regulatory circuit while also providing insight into the transcriptional changes that occur temporally between different stages of biofilm development.
Nobile, C. J. & Mitchell, A. P. Regulation of cell-surface genes and biofilm formation by the C. albicans transcription factor Bcr1p. Curr. Biol. 15, 1150–1155 (2005).
Hernday, A. D. et al. Structure of the transcriptional network controlling white-opaque switching in Candida albicans. Mol. Microbiol. 90, 22–35 (2013).
Borneman, A. R. et al. Target hub proteins serve as master regulators of development in yeast. Genes Dev. 20, 435–438 (2006).
Sorrells, T. R. & Johnson, A. D. Making sense of transcription networks. Cell 161, 714–723 (2015).
Dranginis, A. M., Rauceo, J. M., Coronado, J. E. & Lipke, P. N. A biochemical guide to yeast adhesins: glycoproteins for social and antisocial occasions. Microbiol. Mol. Biol. Rev. 71, 282–294 (2007).
Alves, C. T. et al. Effect of progesterone on Candida albicans vaginal pathogenicity. Int. J. Med. Microbiol. 304, 1011–1017 (2014).
Sandini, S. et al. The MP65 gene is required for cell wall integrity, adherence to epithelial cells and biofilm formation in Candida albicans. BMC Microbiol. 11, 106 (2011).
Frade, J. P. & Arthington-Skaggs, B. A. Effect of serum and surface characteristics on Candida albicans biofilm formation. Mycoses 54, e154–e162 (2011).
Salgado, P. S. et al. Structural basis for the broad specificity to host-cell ligands by the pathogenic fungus Candida albicans. Proc. Natl Acad. Sci. USA 108, 15775–15779 (2011).
Li, F. & Palecek, S. P. Distinct domains of the Candida albicans adhesin Eap1p mediate cell-cell and cell-substrate interactions. Microbiology 154, 1193–1203 (2008).
Parsek, M. R. & Greenberg, E. P. Sociomicrobiology: the connections between quorum sensing and biofilms. Trends Microbiol. 13, 27–33 (2005).
Desai, J. V. & Mitchell, A. P. Candida albicans biofilm development and its genetic control. Microbiol. Spectr. http://dx.doi.org/10.1128/microbiolspec.MB-0005-2014 (2015).
Sundstrom, P. Adhesion in Candida spp. Cell. Microbiol. 4, 461–469 (2002).
Nobile, C. J. et al. Critical role of Bcr1-dependent adhesins in C. albicans biofilm formation in vitro and in vivo. PLoS Pathog. 2, e63 (2006).
Zhao, X., Oh, S. H., Yeater, K. M. & Hoyer, L. L. Analysis of the Candida albicans Als2p and Als4p adhesins suggests the potential for compensatory function within the Als family. Microbiology 151, 1619–1630 (2005).
Li, F. et al. Eap1p, an adhesin that mediates Candida albicans biofilm formation in vitro and in vivo. Eukaryot. Cell 6, 931–939 (2007).
Finkel, J. S. et al. Portrait of Candida albicans adherence regulators. PLoS Pathog. 8, e1002525 (2012). This study provides a comprehensive examination of the regulation of adherence, the first step in biofilm formation.
Swidergall, M. & Filler, S. G. Oropharyngeal candidiasis: fungal invasion and epithelial cell responses. PLoS Pathog. 13, e1006056 (2017).
Kempf, M. et al. Disruption of Candida albicans IFF4 gene involves modifications of the cell electrical surface properties. Colloids Surf. B. Biointerfaces 58, 250–255 (2007).
Hoyer, L. L. & Cota, E. Candida albicans agglutinin-like sequence (Als) family vignettes: a review of Als protein structure and function. Front. Microbiol. 7, 280 (2016).
Nobile, C. J., Nett, J. E., Andes, D. R. & Mitchell, A. P. Function of Candida albicans adhesin Hwp1 in biofilm formation. Eukaryot. Cell 5, 1604–1610 (2006).
Nobile, C. J. et al. Complementary adhesin function in C. albicans biofilm formation. Curr. Biol. 18, 1017–1024 (2008).
Zarnowski, R. et al. Novel entries in a fungal biofilm matrix encyclopedia. mBio 5, e01333-4 (2014). This study provides a detailed breakdown of the composition of the biofilm extracellular matrix.
Martins, M. et al. Presence of extracellular DNA in the Candida albicans biofilm matrix and its contribution to biofilms. Mycopathologia 169, 323–331 (2010).
Nobile, C. J. et al. Biofilm matrix regulation by Candida albicans Zap1. PLoS Biol. 7, e1000133 (2009).
Nett, J. E., Sanchez, H., Cain, M. T., Ross, K. M. & Andes, D. R. Interface of Candida albicans biofilm matrix-associated drug resistance and cell wall integrity regulation. Eukaryot. Cell 10, 1660–1669 (2011).
Mitchell, K. F. et al. Community participation in biofilm matrix assembly and function. Proc. Natl Acad. Sci. USA 112, 4092–4097 (2015). This study examines the role of many polysaccharide components of the biofilm extracellular matrix.
Sudbery, P. E. Growth of Candida albicans hyphae. Nat. Rev. Microbiol. 9, 737–748 (2011).
Ramage, G., Vande Walle, K., López-Ribot, J. L. & Wickes, B. L. The filamentation pathway controlled by the Efg1 regulator protein is required for normal biofilm formation and development in Candida albicans. FEMS Microbiol. Lett. 214, 95–100 (2002).
Schweizer, A., Rupp, S., Taylor, B. N., Röllinghoff, M. & Schröppel, K. The TEA/ATTS transcription factor CaTec1p regulates hyphal development and virulence in Candida albicans. Mol. Microbiol. 38, 435–445 (2000).
Konstantinidou, N. & Morrissey, J. P. Co-occurence of filamentation defects and impaired biofilms in Candida albicans protein kinase mutants. FEMS Yeast Res. 15, fov092 (2015).
Rajendran, R. et al. Integrating Candida albicans metabolism with biofilm heterogeneity by transcriptome mapping. Sci. Rep. 6, 35436 (2016).
Holland, L. M. et al. Comparative phenotypic analysis of the major fungal pathogens Candida parapsilosis and Candida albicans. PLoS Pathog. 10, e1004365 (2014).
Nobile, C. J. et al. A histone deacetylase complex mediates biofilm dispersal and drug resistance in Candida albicans. mBio 5, e01201-14 (2014).
Uppuluri, P. et al. The transcriptional regulator Nrg1p controls Candida albicans biofilm formation and dispersion. Eukaryot. Cell 9, 1531–1537 (2010).
Hnisz, D. et al. A Histone deacetylase adjusts transcription kinetics at coding sequences during Candida albicans morphogenesis. PLoS Genet. 8, e1003118 (2012).
Robbins, N. et al. Hsp90 governs dispersion and drug resistance of fungal biofilms. PLoS Pathog. 7, e1002257 (2011).
Shapiro, R. S. et al. Hsp90 orchestrates temperature-dependent Candida albicans morphogenesis via Ras1-PKA signaling. Curr. Biol. 19, 621–629 (2009).
Granger, B. L. Insight into the antiadhesive effect of yeast wall protein 1 of Candida albicans. Eukaryot. Cell 11, 795–805 (2012).
Blankenship, J. R. & Mitchell, A. P. How to build a biofilm: a fungal perspective. Curr. Opin. Microbiol. 9, 588–594 (2006).
Sellam, A. et al. A Candida albicans early stage biofilm detachment event in rich medium. BMC Microbiol. 9, 25 (2009).
Martinez-Gomariz, M. et al. Proteomic analysis of cytoplasmic and surface proteins from yeast cells, hyphae, and biofilms of Candida albicans. Proteomics 9, 2230–2252 (2009).
Richard, M. L., Nobile, C. J., Bruno, V. M. & Mitchell, A. P. Candida albicans biofilm-defective mutants. Eukaryot. Cell 4, 1493–1502 (2005).
Winter, M. B. et al. Global identification of biofilm-specific proteolysis in Candida albicans. mBio 7, e01514-16 (2016).
Li, P., Seneviratne, C. J., Alpi, E., Vizcaino, J. A. & Jin, L. Delicate metabolic control and coordinated stress response critically determine antifungal tolerance of Candida albicans biofilm persisters. Antimicrob. Agents Chemother. 59, 6101–6112 (2015).
Fiori, B. et al. In vitro activities of anidulafungin and other antifungal agents against biofilms formed by clinical isolates of different Candida and Aspergillus species. Antimicrob. Agents Chemother. 55, 3031–3035 (2011).
Jacobson, M. J., Steckelberg, K. E., Piper, K. E., Steckelberg, J. M. & Patel, R. In vitro activity of micafungin against planktonic and sessile Candida albicans isolates. Antimicrob. Agents Chemother. 53, 2638–2639 (2009).
Pappas, P. G. et al. Clinical practice guidelines for the management of candidiasis: 2009 update by the Infectious Diseases Society of America. Clin. Infect. Dis. 48, 503–535 (2009).
Cowen, L. E. The evolution of fungal drug resistance: modulating the trajectory from genotype to phenotype. Nat. Rev. Microbiol. 6, 187–198 (2008).
Anderson, J. B. Evolution of antifungal-drug resistance: mechanisms and pathogen fitness. Nat. Rev. Microbiol. 3, 547–556 (2005).
Ramage, G., Bachmann, S., Patterson, T. F., Wickes, B. L. & Lopez-Ribot, J. L. Investigation of multidrug efflux pumps in relation to fluconazole resistance in Candida albicans biofilms. J. Antimicrob. Chemother. 49, 973–980 (2002).
Mukherjee, P. K., Chandra, J., Kuhn, D. M. & Ghannoum, M. A. Mechanism of fluconazole resistance in Candida albicans biofilms: phase-specific role of efflux pumps and membrane sterols. Infect. Immun. 71, 4333–4340 (2003).
Prasad, R., Shah, A. H. & Rawal, M. K. Antifungals: mechanism of action and drug resistance. Adv. Exp. Med. Biol. 892, 327–349 (2016).
Mateus, C., Crow, S. A. & Ahearn, D. G. Adherence of Candida albicans to silicone induces immediate enhanced tolerance to fluconazole. Antimicrob. Agents Chemother. 48, 3358–3366 (2004).
Nett, J. E., Lepak, A. J., Marchillo, K. & Andes, D. R. Time course global gene expression analysis of an in vivo Candida biofilm. J. Infect. Dis. 200, 307–313 (2009).
Yeater, K. M. et al. Temporal analysis of Candida albicans gene expression during biofilm development. Microbiology 153, 2373–2385 (2007).
Nett, J. et al. Putative role of β-1,3 glucans in Candida albicans biofilm resistance. Antimicrob. Agents Chemother. 51, 510–520 (2007).
Nett, J. E., Sanchez, H., Cain, M. T. & Andes, D. R. Genetic basis of Candida biofilm resistance due to drug-sequestering matrix glucan. J. Infect. Dis. 202, 171–175 (2010).
Taff, H. T. et al. A Candida biofilm-induced pathway for matrix glucan delivery: implications for drug resistance. PLoS Pathog. 8, e1002848 (2012).
Vediyappan, G., Rossignol, T. & D'Enfert, C. Interaction of Candida albicans biofilms with antifungals: transcriptional response and binding of antifungals to beta-glucans. Antimicrob. Agents Chemother. 54, 2096–2111 (2010).
LaFleur, M. D., Kumamoto, C. A. & Lewis, K. Candida albicans biofilms produce antifungal-tolerant persister cells. Antimicrob. Agents Chemother. 50, 3839–3846 (2006).
Lewis, K. Persister cells, dormancy and infectious disease. Nat. Rev. Microbiol. 5, 48–56 (2007).
Mathe, L. & Van Dijck, P. Recent insights into Candida albicans biofilm resistance mechanisms. Curr. Genet. 59, 251–264 (2013).
Khot, P. D., Suci, P. A., Miller, R. L., Nelson, R. D. & Tyler, B. J. A small subpopulation of blastospores in Candida albicans biofilms exhibit resistance to amphotericin B associated with differential regulation of ergosterol and β-1,6-glucan pathway genes. Antimicrob. Agents Chemother. 50, 3708–3716 (2006).
Truong, T. et al. Comparative ploidy proteomics of Candida albicans biofilms unraveled the role of the AHP1 gene in the biofilm persistence against Amphotericin B. Mol. Cell. Proteom. 15, 3488–3500 (2016).
Ghosh, S. et al. Arginine-induced germ tube formation in Candida albicans is essential for escape from murine macrophage line RAW 264.7. Infect. Immun. 77, 1596–1605 (2009).
Gropp, K. et al. The yeast Candida albicans evades human complement attack by secretion of aspartic proteases. Mol. Immunol. 47, 465–475 (2009).
Szafranski-Schneider, E. et al. Msb2 shedding protects Candida albicans against antimicrobial peptides. PLoS Pathog. 8, e1002501 (2012).
Swidergall, M., Ernst, A. M. & Ernst, J. F. Candida albicans mucin Msb2 is a broad-range protectant against antimicrobial peptides. Antimicrob. Agents Chemother. 57, 3917–3922 (2013).
Hirschfeld, J. Dynamic interactions of neutrophils and biofilms. J. Oral Microbiol. 6, 26102 (2014).
Xie, Z. et al. Candida albicans biofilms do not trigger reactive oxygen species and evade neutrophil killing. J. Infect. Dis. 206, 1936–1945 (2012).
Johnson, C. J. et al. The extracellular matrix of Candida albicans biofilms impairs formation of neutrophil extracellular traps. PLoS Pathog. 12, e1005884 (2016).
Peters, B. M., Jabra-Rizk, M. A., O'May, G. A., Costerton, J. W. & Shirtliff, M. E. Polymicrobial interactions: impact on pathogenesis and human disease. Clin. Microbiol. Rev. 25, 193–213 (2012).
Peleg, A. Y., Hogan, D. A. & Mylonakis, E. Medically important bacterial-fungal interactions. Nat. Rev. Microbiol. 8, 340–349 (2010).
Pammi, M., Zhong, D., Johnson, Y., Revell, P. & Versalovic, J. Polymicrobial bloodstream infections in the neonatal intensive care unit are associated with increased mortality: a case-control study. BMC Infect. Dis. 14, 390 (2014).
Peters, B. M. & Noverr, M. C. Candida albicans-Staphylococcus aureus polymicrobial peritonitis modulates host innate immunity. Infect. Immun. 81, 2178–2189 (2013). This study demonstrates that the combination of C. albicans and S. aureus results in dramatic increases in microbial burden and mortality in a mouse model of peritonitis.
Jarosz, L. M., Deng, D. M., van der Mei, H. C., Crielaard, W. & Krom, B. P. Streptococcus mutans competence-stimulating peptide inhibits Candida albicans hypha formation. Eukaryot. Cell 8, 1658–1664 (2009).
Bamford, C. V. et al. Streptococcus gordonii modulates Candida albicans biofilm formation through intergeneric communication. Infect. Immun. 77, 3696–3704 (2009).
Jack, A. A. et al. Streptococcus gordonii comCDE (competence) operon modulates biofilm formation with Candida albicans. Microbiology 161, 411–421 (2015).
Lindsay, A. K. & Hogan, D. A. Candida albicans: molecular interactions with Pseudomonas aeruginosa and Staphylococcus aureus. Fungal Biol. Rev. 28, 85–96 (2014).
Bamford, C. V., Nobbs, A. H., Barbour, M. E., Lamont, R. J. & Jenkinson, H. F. Functional regions of Candida albicans hyphal cell wall protein Als3 that determine interaction with the oral bacterium Streptococcus gordonii. Microbiology 161, 18–29 (2015).
Beaussart, A. et al. Single-cell force spectroscopy of the medically important Staphylococcus epidermidis-Candida albicans interaction. Nanoscale 5, 10894–10900 (2013).
Hoyer, L. L., Oh, S. H., Jones, R. & Cota, E. A proposed mechanism for the interaction between the Candida albicans Als3 adhesin and streptococcal cell wall proteins. Front. Microbiol. 5, 564 (2014).
Peters, B. M. et al. Staphylococcus aureus adherence to Candida albicans hyphae is mediated by the hyphal adhesin Als3p. Microbiology 158, 2975–2986 (2012).
Schlecht, L. M. et al. Systemic Staphylococcus aureus infection mediated by Candida albicans hyphal invasion of mucosal tissue. Microbiology 161, 168–181 (2015).
Förster, T. M. et al. Enemies and brothers in arms: Candida albicans and gram-positive bacteria. Cell. Microbiol. 28, 1709–1715 (2016).
Harriott, M. M. & Noverr, M. C. Candida albicans and Staphylococcus aureus form polymicrobial biofilms: effects on antimicrobial resistance. Antimicrob. Agents Chemother. 53, 3914–3922 (2009). This work shows that co-incubation of C. albicans and S. aureus results in increased biofilm formation and vancomycin resistance in S. aureus.
Harriott, M. M. & Noverr, M. C. Ability of Candida albicans mutants to induce Staphylococcus aureus vancomycin resistance during polymicrobial biofilm formation. Antimicrob. Agents Chemother. 54, 3746–3755 (2010).
Kong, E. F. et al. Commensal protection of Staphylococcus aureus against antimicrobials by Candida albicans biofilm matrix. mBio 7, e01365-16 (2016). This study identifies the component of the C. albicans extracellular matrix that protects S. aureus from vancomycin.
Fox, E. P. et al. Anaerobic bacteria grow within Candida albicans biofilms and induce biofilm formation in suspension cultures. Curr. Biol. 24, 2411–2416 (2014). This study demonstrates that bacteria can induce C. albicans biofilm formation and use the resulting biofilms for protection from the environment.
Hogan, D. A. & Kolter, R. Pseudomonas-Candida interactions: an ecological role for virulence factors. Science 296, 2229–2232 (2002). This is a foundational study of interactions between C. albicans and bacterial species.
Strus, M. et al. The in vitro activity of vaginal Lactobacillus with probiotic properties against Candida. Infect. Dis. Obstet. Gynecol. 13, 69–75 (2005).
Hogan, D. A., Vik, A. & Kolter, R. A. Pseudomonas aeruginosa quorum-sensing molecule influences Candida albicans morphology. Mol. Microbiol. 54, 1212–1223 (2004).
McAlester, G., O'Gara, F. & Morrissey, J. P. Signal-mediated interactions between Pseudomonas aeruginosa and Candida albicans. J. Med. Microbiol. 57, 563–569 (2008).
Graham, C. E., Cruz, M. R., Garsin, D. A. & Lorenz, M. C. Enterococcus faecalis bacteriocin EntV inhibits hyphal morphogenesis, biofilm formation, and virulence of Candida albicans. Proc. Natl Acad. Sci. USA 114, 4507–4512 (2017).
Seth, E. C. & Taga, M. E. Nutrient cross-feeding in the microbial world. Front. Microbiol. 5, 350 (2014).
Shrestha, P. M. & Rotaru, A. E. Plugging in or going wireless: strategies for interspecies electron transfer. Front. Microbiol. 5, 237 (2014).
Stams, A. J. & Plugge, C. M. Electron transfer in syntrophic communities of anaerobic bacteria and archaea. Nat. Rev. Microbiol. 7, 568–577 (2009).
Morales, D. K. et al. Control of Candida albicans metabolism and biofilm formation by Pseudomonas aeruginosa phenazines. mBio 4, e00526-12 (2013).
Chen, A. I. et al. Candida albicans ethanol stimulates Pseudomonas aeruginosa WspR-controlled biofilm formation as part of a cyclic relationship involving phenazines. PLoS Pathog. 10, e1004480 (2014).
Diaz, P. I. et al. Synergistic interaction between Candida albicans and commensal oral streptococci in a novel in vitro mucosal model. Infect. Immun 80, 620–632 (2012).
Vande Velde, G., Kucharíková, S., Schrevens, S., Himmelreich, U. & Van Dijck, P. Towards non-invasive monitoring of pathogen-host interactions during Candida albicans biofilm formation using in vivo bioluminescence. Cell. Microbiol 16, 115–130 (2014). This study describes an in vivo live-animal imaging model of infection.
Vande Velde, G., Kucharíková, S., Van Dijck, P. & Himmelreich, U. Bioluminescence imaging of fungal biofilm development in live animals. Methods Mol. Biol. 1098, 153–167 (2014).
Tournu, H. & Van Dijck, P. Candida biofilms and the host: models and new concepts for eradication. Int. J. Microbiol. 2012, 845352 (2012).
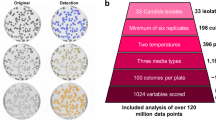
Lohse, M. B. et al. Assessment and optimizations of Candida albicans in vitro biofilm assays. Antimicrob. Agents Chemother. 61, e02749-16 (2017).
Ramage, G., Vande Walle, K., Wickes, B. L. & López-Ribot, J. L. Standardized method for in vitro antifungal susceptibility testing of Candida albicans biofilms. Antimicrob. Agents Chemother. 45, 2475–2479 (2001).
Nett, J. E., Cain, M. T., Crawford, K. & Andes, D. R. Optimizing a Candida biofilm microtiter plate model for measurement of antifungal susceptibility by tetrazolium salt assay. J. Clin. Microbiol. 49, 1426–1433 (2011).
Jin, Y., Yip, H. K., Samaranayake, Y. H., Yau, J. Y. & Samaranayake, L. P. Biofilm-forming ability of Candida albicans is unlikely to contribute to high levels of oral yeast carriage in cases of human immunodeficiency virus infection. J. Clin. Microbiol. 41, 2961–2967 (2003).
Nett, J. E., Marchillo, K., Spiegel, C. A. & Andes, D. R. Development and validation of an in vivo Candida albicans biofilm denture model. Infect. Immun. 78, 3650–3659 (2010).
Wang, X. & Fries, B. C. A murine model for catheter-associated candiduria. J. Med. Microbiol. 60, 1523–1529 (2011).
Doyle, T. C., Nawotka, K. A., Kawahara, C. B., Francis, K. P. & Contag, P. R. Visualizing fungal infections in living mice using bioluminescent pathogenic Candida albicans strains transformed with the firefly luciferase gene. Microb. Pathog. 40, 82–90 (2006).
Kelly, M. T. et al. The Candida albicans CaACE2 gene affects morphogenesis, adherence and virulence. Mol. Microbiol. 53, 969–983 (2004).
Askew, C. et al. The zinc cluster transcription factor Ahr1p directs Mcm1p regulation of Candida albicans adhesion. Mol. Microbiol. 79, 940–953 (2011).
García-Sánchez, S. et al. Candida albicans biofilms: a developmental state associated with specific and stable gene expression patterns. Eukaryot. Cell 3, 536–545 (2004).
Tsai, P. W. et al. The role of Mss11 in Candida albicans biofilm formation. Mol. Genet. Genom. 289, 807–819 (2014).
Sellam, A. et al. Role of transcription factor CaNdt80p in cell separation, hyphal growth, and virulence in Candida albicans. Eukaryot. Cell 9, 634–644 (2010).
Bonhomme, J. et al. Contribution of the glycolytic flux and hypoxia adaptation to efficient biofilm formation by Candida albicans. Mol. Microbiol. 80, 995–1013 (2011).
Ganguly, S. et al. Zap1 control of cell-cell signaling in Candida albicans biofilms. Eukaryot. Cell 10, 1448–1454 (2011).
Acknowledgements
The authors thank Sheena Singh-Babak for comments on the manuscript. Research in the authors' laboratories related to this work is supported by NIH grants R41AI112038 (to C.J.N.) and R01AI083311 (to A.D.J.), and by a Pew Biomedical Scholar Award (to C.J.N.) from the Pew Charitable Trusts. The funders had no role in planning and writing the Review or in the decision to submit the work for publication.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
C.J.N. and A.D.J. are cofounders of BioSynesis, Inc., a company developing inhibitors and diagnostics of Candida albicans biofilm formation, and M.L. is a consultant of BioSynesis, Inc. M.G. does not declare competing interests.
Supplementary information
Supplementary information S1 (box)
The host immune response to C. albicans (DOC 29 kb)
Glossary
- Yeast-form cells
-
Spherical fungal cells that form daughter cells, which bud off from the parent cell.
- Pseudohyphal cells
-
Ovoid chains of fungal cells that contain constrictions (rather than septa) at the cell junctions.
- Hyphal cells
-
Elongated, cylindrical fungal cells that contain complete septa at the cell junctions.
- Extracellular matrix
-
A protective physical barrier that surrounds cells in a biofilm and is composed of proteins, carbohydrates, lipids and nucleic acids.
- Persister cells
-
Non-dividing fungal cells with decreased metabolic activity that are resistant to antimicrobial agents.
- White–opaque switching
-
The ability for Candida albicans cells to switch between the 'white' and 'opaque' phenotypic cell types. The switch occurs epigenetically; that is, without a change in the primary DNA sequence of the genome.
- Horizontal gene transfer
-
The process through which genetic material is transferred between microorganisms through mechanisms such as transformation, conjugation and transduction. This process is distinct from vertical gene transfer, in which genetic material is transferred from mother cells to daughter cells.
- Quorum sensing
-
A method of communication that allows microorganisms to sense cell density and microbial community composition and respond as a group. The process involves the production and detection of soluble quorum sensing molecules.
- Glycosylphosphatidylinositol (GPI) anchors
-
Post-translational modifications of proteins in which a glycolipid is covalently attached and anchors the protein in the plasma membrane.
- Flow cell
-
A light microscopy method for observing biofilm formation in vitro under laminar flow conditions.
- Glucoamylase
-
An enzyme that catalyses the hydrolysis of glucosidic linkages in starch, which releases glucose.
- Glucan synthase
-
A glucosyltransferase enzyme that catalyses the synthesis of glucans, which are critical polysaccharide components of the fungal cell wall and the extracellular matrix.
- Chromatin-modifying complex
-
A protein complex that alters the chromatin structure.
- Complement system
-
A group of proteins that, when activated, mediate the innate immune response and inflammatory response to a pathogen.
- Mucin
-
A glycosylated protein that is the major component of mucus.
- Ergosterol
-
A sterol component of the fungal cell membrane necessary for membrane fluidity.
- Bacteriocin
-
A pore-forming peptide produced by some bacterial and archaeal species that is toxic to other microorganisms.
Rights and permissions
About this article
Cite this article
Lohse, M., Gulati, M., Johnson, A. et al. Development and regulation of single- and multi-species Candida albicans biofilms. Nat Rev Microbiol 16, 19–31 (2018). https://doi.org/10.1038/nrmicro.2017.107
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrmicro.2017.107
This article is cited by
-
Unveiling the gastric microbiota: implications for gastric carcinogenesis, immune responses, and clinical prospects
Journal of Experimental & Clinical Cancer Research (2024)
-
Antifungal activity against Candida albicans of methyl 3,5-dinitrobenzoate loaded nanoemulsion
Brazilian Journal of Microbiology (2024)
-
Streptococcus mutans supernatant affects the virulence of Candida albicans
Brazilian Journal of Microbiology (2024)
-
Exploring the Potential of Farnesol as a Novel Antifungal Drug and Related Challenges
Current Infectious Disease Reports (2024)
-
Optimization of in vitro Biofilm Growth of Candida albicans
Indian Journal of Microbiology (2024)