Key Points
-
Ribosome profiling is a deep-sequencing-based tool that allows the detailed measurement of translation globally and in vivo.
-
The method provides quantification of levels of new protein synthesis, as well as information about ribosome positions that can be used to infer details about translation mechanism or to identify translated open reading frames (ORFs).
-
Ribosome profiling enables instantaneous rather than steady-state measurement and is thus a particularly valuable tool for the study of gene expression over dynamic processes.
-
Proximity-specific ribosome profiling is based on localized labelling of ribosome populations within cells and enables in vivo measurement of translation at specific organelles or subcellular structures.
-
Ribosome profiling is the first tool available for the experimental annotation of translated ORFs and has enabled the discovery of a wide range of new translation products. These include novel short peptides and alternative isoforms of characterized proteins, the vast majority of which are currently of unknown function.
Abstract
Ribosome profiling, which involves the deep sequencing of ribosome-protected mRNA fragments, is a powerful tool for globally monitoring translation in vivo. The method has facilitated discovery of the regulation of gene expression underlying diverse and complex biological processes, of important aspects of the mechanism of protein synthesis, and even of new proteins, by providing a systematic approach for experimental annotation of coding regions. Here, we introduce the methodology of ribosome profiling and discuss examples in which this approach has been a key factor in guiding biological discovery, including its prominent role in identifying thousands of novel translated short open reading frames and alternative translation products.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
McCann, K. L. & Baserga, S. J. Mysterious ribosomopathies. Science 341, 849–850 (2013).
Cleary, J. D. & Ranum, L. P. W. Repeat-associated non-ATG (RAN) translation in neurological disease. Hum. Mol. Genet. 22, R45–R51 (2013).
Ellis, S. R. Nucleolar stress in Diamond Blackfan anemia pathophysiology. Biochim. Biophys. Acta 1842, 765–768 (2014).
Trainor, P. A. & Merrill, A. E. Ribosome biogenesis in skeletal development and the pathogenesis of skeletal disorders. Biochim. Biophys. Acta 1842, 769–778 (2014).
Bolze, A. et al. Ribosomal protein SA haploinsufficiency in humans with isolated congenital asplenia. Science 340, 976–978 (2013).
Kondrashov, N. et al. Ribosome-mediated specificity in Hox mRNA translation and vertebrate tissue patterning. Cell 145, 383–397 (2011).
Ingolia, N. T., Ghaemmaghami, S., Newman, J. R. S. & Weissman, J. S. Genome-wide analysis in vivo of translation with nucleotide resolution using ribosome profiling. Science 324, 218–223 (2009). This work defines the ribosome profiling method and details its specificity, precision and utility.
Wolin, S. L. & Walter, P. Ribosome pausing and stacking during translation of a eukaryotic mRNA. EMBO J. 7, 3559–3569 (1988).
Steitz, J. A. Polypeptide chain initiation: nucleotide sequences of the three ribosomal binding sites in bacteriophage R17 RNA. Nature 224, 957–964 (1969).
Ingolia, N. T., Lareau, L. F. & Weissman, J. S. Ribosome profiling of mouse embryonic stem cells reveals the complexity and dynamics of mammalian proteomes. Cell 147, 789–802 (2011).
Stern-Ginossar, N. et al. Decoding human cytomegalovirus. Science 338, 1088–1093 (2012).
Brar, G. A. et al. High-resolution view of the yeast meiotic program revealed by ribosome profiling. Science 335, 552–557 (2012). References 11 and 12 describe the application of ribosome profiling to physiological dynamic cellular processes, the HCMV infection cycle in human cells and meiosis in budding yeast, respectively. In these disparate systems, both studies identified many new examples of translational control, uORF translation and the translation of many sORFs and alternative ORFs in genomic regions that were thought to be non-coding.
Bazzini, A. A. et al. Identification of small ORFs in vertebrates using ribosome footprinting and evolutionary conservation. EMBO J. 33, 981–993 (2014).
Pauli, A. et al. Toddler: an embryonic signal that promotes cell movement via Apelin receptors. Science 343, 1248636 (2014).
Dunn, J. G., Foo, C. K., Belletier, N. G., Gavis, E. R. & Weissman, J. S. Ribosome profiling reveals pervasive and regulated stop codon readthrough in Drosophila melanogaster. eLife 2, e01179 (2013).
Aspden, J. L. et al. Extensive translation of small open reading frames revealed by Poly-Ribo-Seq. eLife 3, e03528 (2014).
Smith, J. E. et al. Translation of small open reading frames within unannotated RNA transcripts in Saccharomyces cerevisiae. Cell Rep. 7, 1858–1866 (2014).
Lee, S. et al. Global mapping of translation initiation sites in mammalian cells at single-nucleotide resolution. Proc. Natl Acad. Sci. USA 109, E2424–E2432 (2012).
Andreev, D. E. et al. Translation of 5′ leaders is pervasive in genes resistant to eIF2 repression. eLife 4, e03971 (2014).
Van Dijk, E. L., Auger, H., Jaszczyszyn, Y. & Thermes, C. Ten years of next-generation sequencing technology. Trends Genet. 30, 418–426 (2014).
Ingolia, N. T., Brar, G. A., Rouskin, S., McGeachy, A. M. & Weissman, J. S. The ribosome profiling strategy for monitoring translation in vivo by deep sequencing of ribosome-protected mRNA fragments. Nat. Protoc. 7, 1534–1550 (2012).
Kuersten, S., Radek, A., Vogel, C. & Penalva, L. O. Translation regulation gets its 'omics' moment. Wiley Interdiscip. Rev. RNA 4, 617–630 (2013).
Oh, E. et al. Selective ribosome profiling reveals the cotranslational chaperone action of trigger factor in vivo. Cell 147, 1295–1308 (2011).
Arias, C. et al. KSHV 2.0: a comprehensive annotation of the Kaposi's sarcoma-associated herpesvirus genome using next-generation sequencing reveals novel genomic and functional features. PLoS Pathog. 10, e1003847 (2014).
Stadler, M. & Fire, A. Wobble base-pairing slows in vivo translation elongation in metazoans. RNA 17, 2063–2073 (2011).
Bazzini, A. A., Lee, M. T. & Giraldez, A. J. Ribosome profiling shows that miR-430 reduces translation before causing mRNA decay in zebrafish. Science 336, 233–237 (2012).
Juntawong, P., Girke, T., Bazin, J. & Bailey-Serres, J. Translational dynamics revealed by genome-wide profiling of ribosome footprints in Arabidopsis. Proc. Natl Acad. Sci. USA 111, E203–E212 (2014).
Jensen, B. C. et al. Extensive stage-regulation of translation revealed by ribosome profiling of Trypanosoma brucei. BMC Genomics 15, 911 (2014).
Caro, F., Ahyong, V., Betegon, M. & DeRisi, J. L. Genome-wide regulatory dynamics of translation in the Plasmodium falciparum asexual blood stages. eLife 3, e04106 (2014).
Schafer, S. et al. Translational regulation shapes the molecular landscape of complex disease phenotypes. Nat. Commun. 6, 7200 (2015).
Rooijers, K., Loayza-Puch, F., Nijtmans, L. G. & Agami, R. Ribosome profiling reveals features of normal and disease-associated mitochondrial translation. Nat. Commun. 4, 2886 (2013).
Zoschke, R., Watkins, K. P. & Barkan, A. A rapid ribosome profiling method elucidates chloroplast ribosome behavior in vivo. Plant Cell 25, 2265–2275 (2013).
Michel, A. M. et al. GWIPS-viz: development of a ribo-seq genome browser. Nucleic Acids Res. 42, D859–D864 (2014).
Liu, X., Jiang, H., Gu, Z. & Roberts, J. W. High-resolution view of bacteriophage lambda gene expression by ribosome profiling. Proc. Natl Acad. Sci. USA 110, 11928–11933 (2013).
Gerashchenko, M. V., Lobanov, A. V. & Gladyshev, V. N. Genome-wide ribosome profiling reveals complex translational regulation in response to oxidative stress. Proc. Natl Acad. Sci. USA 109, 17394–17399 (2012).
Li, G.-W., Oh, E. & Weissman, J. S. The anti-Shine–Dalgarno sequence drives translational pausing and codon choice in bacteria. Nature 484, 538–541 (2012).
Woolstenhulme, C. J., Guydosh, N. R., Green, R. & Buskirk, A. R. High-precision analysis of translational pausing by ribosome profiling in bacteria lacking EFP. Cell Rep. 11, 13–21 (2015).
Chew, G.-L. et al. Ribosome profiling reveals resemblance between long non-coding RNAs and 5′ leaders of coding RNAs. Development 140, 2828–2834 (2013).
Ingolia, N. T. et al. Ribosome profiling reveals pervasive translation outside of annotated protein-coding genes. Cell Rep. 8, 1365–1379 (2014).
Andreev, D. E. et al. Oxygen and glucose deprivation induces widespread alterations in mRNA translation within 20 minutes. Genome Biol. 16, 90 (2015).
Guydosh, N. R. & Green, R. Dom34 rescues ribosomes in 3′ untranslated regions. Cell 156, 950–962 (2014).
Shalgi, R. et al. Widespread regulation of translation by elongation pausing in heat shock. Mol. Cell 49, 439–452 (2013).
Han, Y. et al. Ribosome profiling reveals sequence-independent post-initiation pausing as a signature of translation. Cell Res. 24, 842–851 (2014).
Liu, B., Han, Y. & Qian, S.-B. Cotranslational response to proteotoxic stress by elongation pausing of ribosomes. Mol. Cell 49, 453–463 (2013).
Subramaniam, A. R., Zid, B. M. & O'Shea, E. K. An integrated approach reveals regulatory controls on bacterial translation elongation. Cell 159, 1200–1211 (2014). This work probes position-specific changes in ribosome distribution among various cellular conditions, concluding that tRNA abundances do not account for elongation rates for most codons, and that pausing of ribosomes during starvation may result in translation abortion.
Lareau, L. F., Hite, D. H., Hogan, G. J. & Brown, P. O. Distinct stages of the translation elongation cycle revealed by sequencing ribosome-protected mRNA fragments. eLife 3, e01257 (2014). This work identifies a class of short ribosome footprints that may be enriched by treatment with translation elongation inhibitors and that are likely to represent a distinct conformation of the ribosome at a specific stage of the elongation cycle.
Siegel, A. F., van den Engh, G., Hood, L., Trask, B. & Roach, J. C. Modeling the feasibility of whole genome shotgun sequencing using a pairwise end strategy. Genomics 68, 237–246 (2000).
Roberts, A., Schaeffer, L. & Pachter, L. Updating RNA-Seq analyses after re-annotation. Bioinformatics 29, 1631–1637 (2013).
Saliba, A.-E., Westermann, A. J., Gorski, S. A. & Vogel, J. Single-cell RNA-seq: advances and future challenges. Nucleic Acids Res. 42, 8845–8860 (2014).
Green, R. & Noller, H. F. Ribosomes and translation. Annu. Rev. Biochem. 66, 679–716 (1997).
Li, G.-W., Burkhardt, D., Gross, C. & Weissman, J. S. Quantifying absolute protein synthesis rates reveals principles underlying allocation of cellular resources. Cell 157, 624–635 (2014).
Aris, J. P., Klionsky, D. J. & Simoni, R. D. The Fo subunits of the Escherichia coli F1Fo-ATP synthase are sufficient to form a functional proton pore. J. Biol. Chem. 260, 11207–11215 (1985).
Humphryes, N. et al. The Ecm11–Gmc2 complex promotes synaptonemal complex formation through assembly of transverse filaments in budding yeast. PLoS Genet. 9, e1003194 (2013).
Lee, M. T. et al. Nanog, Pou5f1 and SoxB1 activate zygotic gene expression during the maternal-to-zygotic transition. Nature 503, 360–364 (2013).
Kronja, I. et al. Widespread changes in the posttranscriptional landscape at the Drosophila oocyte-to-embryo transition. Cell Rep. 7, 1495–1508 (2014).
Vasquez, J.-J., Hon, C.-C., Vanselow, J. T., Schlosser, A. & Siegel, T. N. Comparative ribosome profiling reveals extensive translational complexity in different Trypanosoma brucei life cycle stages. Nucleic Acids Res. 42, 3623–3637 (2014).
Stumpf, C. R., Moreno, M. V., Olshen, A. B., Taylor, B. S. & Ruggero, D. The translational landscape of the mammalian cell cycle. Mol. Cell 52, 574–582 (2013).
Stadler, M. & Fire, A. Conserved translatome remodeling in nematode species executing a shared developmental transition. PLoS Genet. 9, e1003739 (2013).
Liu, B. & Qian, S.-B. Translational reprogramming in cellular stress response. Wiley Interdiscip. Rev. RNA 5, 301–305 (2014).
Ingolia, N. T. Ribosome profiling: new views of translation, from single codons to genome scale. Nat. Rev. Genet. 15, 205–213 (2014).
Michel, A. M. & Baranov, P. V. Ribosome profiling: a Hi-Def monitor for protein synthesis at the genome-wide scale. Wiley Interdiscip. Rev. RNA 4, 473–490 (2013).
Cox, J. S. & Walter, P. A novel mechanism for regulating activity of a transcription factor that controls the unfolded protein response. Cell 87, 391–404 (1996).
Kannan, K. et al. The general mode of translation inhibition by macrolide antibiotics. Proc. Natl Acad. Sci. USA 111, 15958–15963 (2014).
Kannan, K., Vázquez-Laslop, N. & Mankin, A. S. Selective protein synthesis by ribosomes with a drug-obstructed exit tunnel. Cell 151, 508–520 (2012).
Davis, A. R., Gohara, D. W. & Yap, M.-N. F. Sequence selectivity of macrolide-induced translational attenuation. Proc. Natl Acad. Sci. USA 111, 15379–15384 (2014).
Jung, H., Gkogkas, C. G., Sonenberg, N. & Holt, C. E. Remote control of gene function by local translation. Cell 157, 26–40 (2014).
Williams, C. C., Jan, C. H. & Weissman, J. S. Targeting and plasticity of mitochondrial proteins revealed by proximity-specific ribosome profiling. Science 346, 748–751 (2014).
Jan, C. H., Williams, C. C. & Weissman, J. S. Principles of ER cotranslational translocation revealed by proximity-specific ribosome profiling. Science 346, 1257521 (2014).
Arava, Y. et al. Genome-wide analysis of mRNA translation profiles in Saccharomyces cerevisiae. Proc. Natl Acad. Sci. USA 100, 3889–3894 (2003).
Heiman, M., Kulicke, R., Fenster, R. J., Greengard, P. & Heintz, N. Cell type-specific mRNA purification by translating ribosome affinity purification (TRAP). Nat. Protoc. 9, 1282–1291 (2014).
Heiman, M. et al. A translational profiling approach for the molecular characterization of CNS cell types. Cell 135, 738–748 (2008).
Inada, T. et al. One-step affinity purification of the yeast ribosome and its associated proteins and mRNAs. RNA 8, 948–958 (2002).
Zanetti, M. E., Chang, I.-F., Gong, F., Galbraith, D. W. & Bailey-Serres, J. Immunopurification of polyribosomal complexes of Arabidopsis for global analysis of gene expression. Plant Physiol. 138, 624–635 (2005).
Mustroph, A., Zanetti, M. E., Girke, T. & Bailey-Serres, J. Isolation and analysis of mRNAs from specific cell types of plants by ribosome immunopurification. Methods Mol. Biol. 959, 277–302 (2013).
Thomas, A. et al. A versatile method for cell-specific profiling of translated mRNAs in Drosophila. PLoS ONE 7, e40276 (2012).
Housley, M. P. et al. Translational profiling through biotinylation of tagged ribosomes in zebrafish. Development 141, 3988–3993 (2014).
Reinhardt, J. A. & Jones, C. D. Two rapidly evolving genes contribute to male fitness in Drosophila. J. Mol. Evol. 77, 246–259 (2013).
Starck, S. R. et al. Leucine-tRNA initiates at CUG start codons for protein synthesis and presentation by MHC class I. Science 336, 1719–1723 (2012).
Rebbapragada, I. & Lykke-Andersen, J. Execution of nonsense-mediated mRNA decay: what defines a substrate? Curr. Opin. Cell Biol. 21, 394–402 (2009).
Pauli, A., Valen, E. & Schier, A. F. Identifying (non-)coding RNAs and small peptides: challenges and opportunities. BioEssays 37, 103–112 (2015).
Ulitsky, I. & Bartel, D. P. lincRNAs: genomics, evolution, and mechanisms. Cell 154, 26–46 (2013).
Kondo, T. et al. Small peptides switch the transcriptional activity of Shavenbaby during Drosophila embryogenesis. Science 329, 336–339 (2010). This paper identifies key roles in fly development for several short peptides (from 11 to 32 amino acids) translated from sORFs on a transcript that was previously thought to be non-coding.
Magny, E. G. et al. Conserved regulation of cardiac calcium uptake by peptides encoded in small open reading frames. Science 341, 1116–1120 (2013).
Anderson, D. M. et al. A micropeptide encoded by a putative long noncoding RNA regulates muscle performance. Cell 160, 595–606 (2015).
Ruiz-Orera, J., Messeguer, X., Subirana, J. A. & Alba, M. M. Long non-coding RNAs as a source of new peptides. eLife 3, e03523 (2014).
Xu, Y. & Ganem, D. Making sense of antisense: seemingly noncoding RNAs antisense to the master regulator of Kaposi's sarcoma-associated herpesvirus lytic replication do not regulate that transcript but serve as mRNAs encoding small peptides. J. Virol. 84, 5465–5475 (2010).
Guttman, M., Russell, P., Ingolia, N. T., Weissman, J. S. & Lander, E. S. Ribosome profiling provides evidence that large noncoding RNAs do not encode proteins. Cell 154, 240–251 (2013).
Carvunis, A.-R. et al. Proto-genes and de novo gene birth. Nature 487, 370–374 (2012). In this work, the authors present evidence to support the protogene hypothesis, according to which new proteins can evolve through the selection and elongation of ORFs encoding peptides translated from putative intergenic transcripts.
Brubaker, S. W., Gauthier, A. E., Mills, E. W., Ingolia, N. T. & Kagan, J. C. A bicistronic MAVS transcript highlights a class of truncated variants in antiviral immunity. Cell 156, 800–811 (2014).
Michel, A. M. et al. Observation of dually decoded regions of the human genome using ribosome profiling data. Genome Res. 22, 2219–2229 (2012).
Noderer, W. L. et al. Quantitative analysis of mammalian translation initiation sites by FACS-seq. Mol. Syst. Biol. 10, 748 (2014).
Schwaid, A. G. et al. Chemoproteomic discovery of cysteine-containing human short open reading frames. J. Am. Chem. Soc. 135, 16750–16753 (2013).
Slavoff, S. A. et al. Peptidomic discovery of short open reading frame-encoded peptides in human cells. Nat. Chem. Biol. 9, 59–64 (2013).
Ma, J. et al. Discovery of human sORF-encoded polypeptides (SEPs) in cell lines and tissue. J. Proteome Res. 13, 1757–1765 (2014).
Crappé, J. et al. Combining in silico prediction and ribosome profiling in a genome-wide search for novel putatively coding sORFs. BMC Genomics 14, 648 (2013).
Menschaert, G. et al. Deep proteome coverage based on ribosome profiling aids mass spectrometry-based protein and peptide discovery and provides evidence of alternative translation products and near-cognate translation initiation events. Mol. Cell. Proteomics 12, 1780–1790 (2013).
Vanderperre, B. et al. Direct detection of alternative open reading frames translation products in human significantly expands the proteome. PLoS ONE 8, e70698 (2013).
Lin, M. F., Jungreis, I. & Kellis, M. PhyloCSF: a comparative genomics method to distinguish protein coding and non-coding regions. Bioinformatics 27, i275–i282 (2011).
Huang, H.-Y., Tang, H.-L., Chao, H.-Y., Yeh, L.-S. & Wang, C.-C. An unusual pattern of protein expression and localization of yeast alanyl-tRNA synthetase isoforms. Mol. Microbiol. 60, 189–198 (2006).
Chang, K.-J. & Wang, C.-C. Translation initiation from a naturally occurring non-AUG codon in Saccharomyces cerevisiae. J. Biol. Chem. 279, 13778–13785 (2004).
Wan, J. & Qian, S.-B. TISdb: a database for alternative translation initiation in mammalian cells. Nucleic Acids Res. 42, D845–D850 (2014). This work presents a database of alternative translation initiation sites that have been identified by ribosome profiling in mammalian cells.
Artieri, C. G. & Fraser, H. B. Evolution at two levels of gene expression in yeast. Genome Res. 24, 411–421 (2014).
Jungreis, I. et al. Evidence of abundant stop codon readthrough in Drosophila and other metazoa. Genome Res. 21, 2096–2113 (2011).
Schueren, F. et al. Peroxisomal lactate dehydrogenase is generated by translational readthrough in mammals. eLife 3, e03640 (2014).
Jinek, M. et al. A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science 337, 816–821 (2012).
Zalatan, J. G. et al. Engineering complex synthetic transcriptional programs with CRISPR RNA scaffolds. Cell 160, 339–350 (2015).
Gilbert, L. A. et al. CRISPR-mediated modular RNA-guided regulation of transcription in eukaryotes. Cell 154, 442–451 (2013).
Gilbert, L. A. et al. Genome-scale CRISPR-mediated control of gene repression and activation. Cell 159, 647–661 (2014).
Qi, L. S. et al. Repurposing CRISPR as an RNA-guided platform for sequence-specific control of gene expression. Cell 152, 1173–1183 (2013).
Acknowledgements
The authors wish to thank C. Jan and E. Ünal for helpful comments on this manuscript and N. Ingolia for development of the original ribosome profiling protocol and helpful discussions. This work was partially supported by the Winkler Family Biological Sciences Award to G.A.B. and Howard Hughes Medical Institute and Center for RNA Systems Biology funding to J.S.W.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
J.S.W. is an inventor on a patent application for ribosome profiling.
Supplementary information
Supplementary information S1 (Figure)
FLOSS analysis enables post-hoc curation of ribosome profiling data. (PDF 197 kb)
Glossary
- Ribosome footprints
-
mRNA fragments of ∼30 nucleotides that result from nuclease treatment of translating ribosomes. These are mRNA regions that are protected by the ribosome as the mRNA is decoded to a protein sequence.
- Upstream open reading frames
-
(uORFs). ORFs in the 5′ leader region of a characterized mRNA transcript. Translation of uORFs may regulate translation of a downstream ORF. Ribosome profiling allows for the empirical identification of all translated uORFs in vivo under a condition of interest. Although uORFs are short, here we do not include them in the class of 'short ORFs', which are on an mRNA that was not previously thought to encode a protein.
- Polysome gradients
-
A method for fractionating ribosomes that are bound to mRNAs by velocity centrifugation of cell extract on sucrose gradients, allowing for the separation of mRNAs that are associated with one ribosome (monosome) from those being translated by multiple ribosomes (polysome). Sucrose gradient fractionation facilitates qualitative analysis of the translation status of cells.
- Ribosome P site
-
The site within an actively translating ribosome that is usually associated with the tRNA attached to the growing peptide chain.
- Codon periodicity
-
The three-nucleotide pattern of ribosome occupancy, reflecting mRNA translocation in the ribosome by codon as translation occurs.
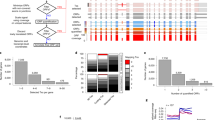
- Fragment length organization similarity score (FLOSS) analysis
-
A metric for determining the probability that ribosome footprints over a given region (or set of regions) result from translation. This analysis involves comparing size distributions of footprints over a query region and over validated coding regions and is based on the concept that the biophysical properties of translating ribosomes result in characteristic signatures in ribosome footprint sizes.
- Translocon
-
The proteinaceous tunnel through which nascent proteins cross the endoplasmic reticulum membrane.
- Translating ribosome affinity capture
-
(TRAP). A method that allows identification of translated mRNAs on the basis of their in vivo association with a tagged ribosomal subunit that is expressed in a cell type-specific manner. This method is a valuable tool for assaying tissue-specific translation in animal and plant systems.
- Nonsense-mediated decay
-
mRNA degradation, which has traditionally been thought to result from stop codons that terminate translation more 5′ than is usual on an mRNA.
- Short ORFs
-
(sORFs). Open reading frames of fewer than 100 codons on mRNAs that are not known to encode a canonical (long) protein. sORFs are a class of ORF that have not traditionally been thought to be frequently translated, although ribosome profiling and other approaches have recently validated the translation of thousands of sORFs in a range of organisms.
- ORFs encoding alternative isoforms of known proteins
-
Open reading frames (ORFs) that differ from another ORF at the same locus in either the start codon or the stop codon position but share the same reading frame. Translation of these ORFs may result in, for example, different subcellular targeting for a similar protein.
- Signatures of protein-coding conservation
-
Purifying evolutionary selection results in higher levels of synonymous than nonsynonymous substitutions, specifically among homologous coding sequences. The pattern of nonsynonymous to synonymous differences among homologous regions compared in a phylogenetic group can be used to predict the likelihood that a genomic locus encodes a translated open reading frame (ORF).
Rights and permissions
About this article
Cite this article
Brar, G., Weissman, J. Ribosome profiling reveals the what, when, where and how of protein synthesis. Nat Rev Mol Cell Biol 16, 651–664 (2015). https://doi.org/10.1038/nrm4069
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrm4069
This article is cited by
-
Ribosome profiling: a powerful tool in oncological research
Biomarker Research (2024)
-
Threonine fuels glioblastoma through YRDC-mediated codon-biased translational reprogramming
Nature Cancer (2024)
-
Dissecting 16p11.2 hemi-deletion to study sex-specific striatal phenotypes of neurodevelopmental disorders
Molecular Psychiatry (2024)
-
The distinct translational landscapes of gram-negative Salmonella and gram-positive Listeria
Nature Communications (2023)
-
Start codon-associated ribosomal frameshifting mediates nutrient stress adaptation
Nature Structural & Molecular Biology (2023)