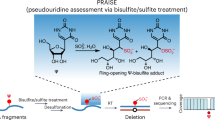
Abstract
Pseudouridylation is the most abundant internal post-transcriptional modification of stable RNAs, with fundamental roles in the biogenesis and function of spliceosomal small nuclear RNAs (snRNAs) and ribosomal RNAs (rRNAs). Recently, the first transcriptome-wide maps of RNA pseudouridylation were published, greatly expanding the catalogue of known pseudouridylated RNAs. These data have further implicated RNA pseudouridylation in the cellular stress response and, moreover, have established that mRNAs are also targets of pseudouridine synthases, potentially representing a novel mechanism for expanding the complexity of the cellular proteome.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Cohn, W. E. Some results of the applications of ion-exchange chromatography to nucleic acid chemistry. J. Cell. Physiol. Suppl. 38, 21–40 (1951).
Cohn, W. E. 5-ribosyl uracil, a carbon-carbon ribofuranosyl nucleoside in ribonucleic acids. Biochim. Biophys. Acta 32, 569–571 (1959).
Cohn, W. E. Pseudouridine, a carbon-carbon linked ribonucleoside in ribonucleic acids: isolation, structure, and chemical characteristics. J. Biol. Chem. 235, 1488–1498 (1960).
Arnez, J. G. & Steitz, T. A. Crystal structure of unmodified tRNAGln complexed with glutaminyl-tRNA synthetase and ATP suggests a possible role for pseudo-uridines in stabilization of RNA structure. Biochemistry 33, 7560–7567 (1994).
Charette, M. & Gray, M. W. Pseudouridine in RNA: what, where, how, and why. IUBMB Life 49, 341–351 (2000).
Karijolich, J., Kantartzis, A. & Yu, Y.-T. RNA modifications: a mechanism that modulates gene expression. Methods Mol. Biol. 629, 1–19 (2010).
Spenkuch, F., Motorin, Y. & Helm, M. Pseudouridine: still mysterious, but never a fake (uridine)! RNA Biol. 11, 1540–1554 (2014).
Ganot, P., Bortolin, M. L. & Kiss, T. Site-specific pseudouridine formation in preribosomal RNA is guided by small nucleolar RNAs. Cell 89, 799–809 (1997).
Ni, J., Tien, A. L. & Fournier, M. J. Small nucleolar RNAs direct site-specific synthesis of pseudouridine in ribosomal RNA. Cell 89, 565–573 (1997).
Wu, G., Xiao, M., Yang, C. & Yu, Y.-T. U2 snRNA is inducibly pseudouridylated at novel sites by Pus7p and snR81 RNP. EMBO J. 30, 79–89 (2011).
Meier, U. T. Pseudouridylation goes regulatory. EMBO J. 30, 3–4 (2011).
Basak, A. & Query, C. C. A pseudouridine residue in the spliceosome core is part of the filamentous growth program in yeast. Cell Rep. 8, 966–973 (2014).
Carlile, T. M. et al. Pseudouridine profiling reveals regulated mRNA pseudouridylation in yeast and human cells. Nature 515, 143–146 (2014).
Lovejoy, A. F., Riordan, D. P. & Brown, P. O. Transcriptome-wide mapping of pseudouridines: pseudouridine synthases modify specific mRNAs in S. cerevisiae. PLoS ONE 9, e110799 (2014).
Schwartz, S. et al. Transcriptome-wide mapping reveals widespread dynamic-regulated pseudouridylation of ncRNA and mRNA. Cell 159, 148–162 (2014).
Darzacq, X. et al. Cajal body-specific small nuclear RNAs: a novel class of 2′-O-methylation and pseudouridylation guide RNAs. EMBO J. 21, 2746–2756 (2002).
Deryusheva, S. & Gall, J. G. Small Cajal body-specific RNAs of Drosophila function in the absence of Cajal bodies. Mol. Biol. Cell 20, 5250–5259 (2009).
Zhao, X., Li, Z.-H., Terns, R. M., Terns, M. P. & Yu, Y.-T. An H/ACA guide RNA directs U2 pseudouridylation at two different sites in the branchpoint recognition region in Xenopus oocytes. RNA 8, 1515–1525 (2002).
Huttenhofer, A. et al. RNomics: an experimental approach that identifies 201 candidates for novel, small, non-messenger RNAs in mouse. EMBO J. 20, 2943–2953 (2001).
Vitali, P. et al. Identification of 13 novel human modification guide RNAs. Nucleic Acids Res. 31, 6543–6551 (2003).
Kiss, A. M., Jady, B. E., Bertrand, E. & Kiss, T. Human box H/ACA pseudouridylation guide RNA machinery. Mol. Cell. Biol. 24, 5797–5807 (2004).
Chen, C., Zhao, X., Kierzek, R. & Yu, Y.-T. A flexible RNA backbone within the polypyrimidine tract is required for U2AF65 binding and pre-mRNA splicing in vivo. Mol. Cell. Biol. 30, 4108–4119 (2010).
Karijolich, J. & Yu, Y.-T. Converting nonsense codons into sense codons by targeted pseudouridylation. Nature 474, 395–398 (2011).
Fernandez, I. S. et al. Unusual base pairing during the decoding of a stop codon by the ribosome. Nature 500, 107–110 (2013).
Parisien, M., Yi, C. & Pan, T. Rationalization and prediction of selective decoding of pseudouridine-modified nonsense and sense codons. RNA 18, 355–367 (2012).
Kariko, K. et al. Incorporation of pseudouridine into mRNA yields superior nonimmunogenic vector with increased translational capacity and biological stability. Mol. Ther. 16, 1833–1840 (2008).
Kishore, S. et al. Insights into snoRNA biogenesis and processing from PAR-CLIP of snoRNA core proteins and small RNA sequencing. Genome Biol. 14, R45 (2013).
Ofengand, J. & Bakin, A. Mapping to nucleotide resolution of pseudouridine residues in large subunit ribosomal RNAs from representative eukaryotes, prokaryotes, archaebacteria, mitochondria and chloroplasts. J. Mol. Biol. 266, 246–268 (1997).
Dominissini, D. et al. Topology of the human and mouse m6A RNA methylomes revealed by m6A-seq. Nature 485, 201–206 (2012).
Meyer, K. D. et al. Comprehensive analysis of mRNA methylation reveals enrichment in 3′ UTRs and near stop codons. Cell 149, 1635–1646 (2012).
Wang, X. et al. N6-methyladenosine-dependent regulation of messenger RNA stability. Nature 505, 117–120 (2014).
Meier, U. T. Dissecting dyskeratosis. Nat. Genet. 33, 116–117 (2003).
Ruggero, D. et al. Dyskeratosis congenita and cancer in mice deficient in ribosomal RNA modification. Science 299, 259–262 (2003).
Bellodi, C. et al. H/ACA small RNA dysfunctions in disease reveal key roles for noncoding RNA modifications in hematopoietic stem cell differentiation. Cell Rep. 3, 1493–1502 (2013).
Mitchell, J. R., Wood, E. & Collins, K. A telomerase component is defective in the human disease dyskeratosis congenita. Nature 402, 551–555 (1999).
Bellodi, C. et al. Loss of function of the tumor suppressor DKC1 perturbs p27 translation control and contributes to pituitary tumorigenesis. Cancer Res. 70, 6026–6035 (2010).
Bykhovskaya, Y., Casas, K., Mengesha, E., Inbal, A. & Fischel-Ghodsian, N. Missense mutation in pseudouridine synthase 1 (PUS1) causes mitochondrial myopathy and sideroblastic anemia (MLASA). Am. J. Hum. Genet. 74, 1303–1308 (2004).
Murray, J. L., Sheng, J. & Rubin, D. H. A role for H/ACA and C/D small nucleolar RNAs in viral replication. Mol. Biotechnol. 56, 429–437 (2014).
Acknowledgements
The authors would like to thank members of the Yu laboratory for insightful discussions.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Karijolich, J., Yi, C. & Yu, YT. Transcriptome-wide dynamics of RNA pseudouridylation. Nat Rev Mol Cell Biol 16, 581–585 (2015). https://doi.org/10.1038/nrm4040
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrm4040
This article is cited by
-
HIF1A-repressed PUS10 regulates NUDC/Cofilin1 dependent renal cell carcinoma migration by promoting the maturation of miR-194-5p
Cell & Bioscience (2023)
-
RNA modifications—a regulatory dimension yet to be deciphered in immunity
Genes & Immunity (2023)
-
RNA modification in cardiovascular disease: implications for therapeutic interventions
Signal Transduction and Targeted Therapy (2023)
-
The emerging role of epitranscriptome in shaping stress responses in plants
Plant Cell Reports (2023)
-
Mechanism of Dcp2/RNCR3/Dkc1/Snora62 axis regulating neuronal apoptosis in chronic cerebral ischemia
Cell Biology and Toxicology (2023)