Key Points
-
During cell division, DNA sequence along with its distinct organization in chromatin must be transmitted to daughter cells in order to maintain genome stability and keep the memory of established transcriptional states that are central to cell fate and identity.
-
Local chromatin structure and three-dimensional chromosomal organization influence where and when DNA replication initiates. The mechanisms that control replication timing are still enigmatic, but there are strong correlations to developmental decisions as well as the spectra of genome rearrangements observed in cancer.
-
In every S phase, chromatin undergoes disruption as replication forks travel through the genome and sophisticated mechanisms operate in concert with the replisome to reproduce chromatin organization on the new daughter strands. In recent years, many of these mechanisms have been discovered, but how these activities are directed to specific types of chromatin remains a key question.
-
How specific chromatin marks are restored on newly replicated chromatin is seminal to understanding epigenetic inheritance of transcriptional states. Specific self-perpetuating mechanisms have been identified for individual marks, and insights into restoration kinetics have revealed unexpected fluctuation of methylation states through the cell cycle.
-
Cancer development is characterized by genetic and epigenetic alterations. Emerging evidence suggests that replication stress, a known source of genome instability, may also fuel epigenome aberrations and challenge chromatin maintenance.
Abstract
Stability and function of eukaryotic genomes are closely linked to chromatin structure and organization. During cell division the entire genome must be accurately replicated and the chromatin landscape reproduced on new DNA. Chromatin and nuclear structure influence where and when DNA replication initiates, whereas the replication process itself disrupts chromatin and challenges established patterns of genome regulation. Specialized replication-coupled mechanisms assemble new DNA into chromatin, but epigenome maintenance is a continuous process taking place throughout the cell cycle. If DNA synthesis is perturbed, cells can suffer loss of both genome and epigenome integrity with severe consequences for the organism.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Probst, A. V., Dunleavy, E & Almouzni G. Epigenetic inheritance during the cell cycle. Nature Rev. Mol. Cell Biol. 10, 192–206 (2009).
Groth, A., Rocha, W., Verreault, A. & Almouzni, G. Chromatin challenges during DNA replication and repair. Cell 128, 721–733 (2007).
Meister, P., Mango, S. E. & Gasser, S. M. Locking the genome: nuclear organization and cell fate. Curr. Opin. Genet. Dev. 21, 167–174 (2011).
Blow, J. J., Ge, X. Q. & Jackson, D. A. How dormant origins promote complete genome replication. Trends Biochem. Sci. 36, 405–414 (2011).
Branzei, D. & Foiani, M. Maintaining genome stability at the replication fork. Nature Rev. Mol. Cell Biol. 11, 208–219 (2010).
Halazonetis, T. D., Gorgoulis, V. G. & Bartek, J. An oncogene-induced DNA damage model for cancer development. Science 319, 1352–1355 (2008).
Sharma, S., Kelly, T. K. & Jones, P. A. Epigenetics in cancer. Carcinogenesis 31, 27–36 (2010).
Jasencakova, Z. & Groth, A. Replication stress, a source of epigenetic aberrations in cancer? Bioessays 32, 847–855 (2010).
Kouzarides, T. Chromatin modifications and their function. Cell 128, 693–705 (2007).
Mechali, M. Eukaryotic DNA replication origins: many choices for appropriate answers. Nature Rev. Mol. Cell Biol. 11, 728–738 (2010).
Remus, D. & Diffley, J. F. Eukaryotic DNA replication control: lock and load, then fire. Curr. Opin. Cell Biol. 21, 771–777 (2009).
Letessier, A. et al. Cell type specific replication initiation programs set fragility of the FRA3B fragile site. Nature 470, 120–123 (2011).
MacAlpine, H. K., Gordan, R., Powell, S. K., Hartemink, A. J. & MacAlpine, D. M. Drosophila ORC localizes to open chromatin and marks sites of cohesin complex loading. Genome Res. 20, 201–211 (2010).
Lubelsky, Y. et al. Pre-replication complex proteins assemble at regions of low nucleosome occupancy within the Chinese hamster dihydrofolate reductase initiation zone. Nucleic Acids Res. 39, 3141–3155 (2011).
Cayrou, C. et al. Genome-scale analysis of metazoan replication origins reveals their organization in specific but flexible sites defined by conserved features. Genome Res. 12, 1438–1449 (2011). Provides, together with references 13 and 14, the first genome-wide maps of ORC localization, revealing its preference for nucleosome-free regions.
Thomae, A. W. et al. Interaction between HMGA1a and the origin recognition complex creates site-specific replication origins. Proc. Natl Acad. Sci. USA 105, 1692–1697 (2008).
Atanasiu, C., Deng, Z., Wiedmer, A., Norseen, J. & Lieberman, P. M. ORC binding to TRF2 stimulates OriP replication. EMBO Rep. 7, 716–721 (2006).
Schwaiger, M., Kohler, H., Oakeley, E. J., Stadler, M. B. & Schubeler, D. Heterochromatin protein 1 (HP1) modulates replication timing of the Drosophila genome. Genome Res. 20, 771–780 (2010).
Tardat, M. et al. The histone H4 Lys 20 methyltransferase PRSet7 regulates replication origins in mammalian cells. Nature Cell Biol. 12, 1086–1093 (2010).
Jorgensen, S. et al. SET8 is degraded via PCNA-coupled CRL4(CDT2) ubiquitylation in S phase and after UV irradiation. J. Cell Biol. 192, 43–54 (2011).
Centore, R. C. et al. CRL4(Cdt2)-mediated destruction of the histone methyltransferase Set8 prevents premature chromatin compaction in S phase. Mol. Cell 40, 22–33 (2010).
Rice, J. C. et al. Mitotic-specific methylation of histone H4 Lys 20 follows increased PRSet7 expression and its localization to mitotic chromosomes. Genes Dev. 16, 2225–2230 (2002).
Miotto, B. & Struhl, K. HBO1 histone acetylase activity is essential for DNA replication licensing and inhibited by Geminin. Mol. Cell 37, 57–66 (2010).
Miotto, B. & Struhl, K. HBO1 histone acetylase is a coactivator of the replication licensing factor Cdt1. Genes Dev. 22, 2633–2638 (2008).
Iizuka, M., Matsui, T., Takisawa, H. & Smith, M. M. Regulation of replication licensing by acetyltransferase Hbo1. Mol. Cell. Biol. 26, 1098–1108 (2006).
Remus, D. et al. Concerted loading of Mcm2–7 double hexamers around DNA during DNA replication origin licensing. Cell 139, 719–730 (2009).
Gambus, A. et al. GINS maintains association of Cdc45 with MCM in replisome progression complexes at eukaryotic DNA replication forks. Nature Cell Biol. 8, 358–366 (2006). Applies an elegant two-component strategy, tagging Sld5 and Mcm4, to purify and characterize the yeast replication progression complex.
Ge, X. Q., Jackson, D. A. & Blow, J. J. Dormant origins licensed by excess Mcm2–7 are required for human cells to survive replicative stress. Genes Dev. 21, 3331–3341 (2007).
Ibarra, A., Schwob, E. & Mendez, J. Excess MCM proteins protect human cells from replicative stress by licensing backup origins of replication. Proc. Natl Acad. Sci. USA 105, 8956–8961 (2008).
Courbet, S. et al. Replication fork movement sets chromatin loop size and origin choice in mammalian cells. Nature 455, 557–560 (2008). Demonstrates, together with reference 34, the interdependency between chromatin three-dimensional organization and spacing of replication origins.
Jackson, D. A. & Pombo, A. Replicon clusters are stable units of chromosome structure: evidence that nuclear organization contributes to the efficient activation and propagation of S phase in human cells. J. Cell Biol. 140, 1285–1295 (1998).
Gillespie, P. J. & Blow, J. J. Clusters, factories and domains: the complex structure of S-phase comes into focus. Cell Cycle 9, 3218–3226 (2010).
Buongiorno-Nardelli, M., Micheli, G., Carri, M. T. & Marilley, M. A relationship between replicon size and supercoiled loop domains in the eukaryotic genome. Nature 298, 100–102 (1982).
Guillou, E. et al. Cohesin organizes chromatin loops at DNA replication factories. Genes Dev. 24, 2812–2822 (2010).
Gilbert, D. M. et al. Space and time in the nucleus: developmental control of replication timing and chromosome architecture. Cold Spring Harb. Symp. Quant. Biol. 75, 143–153 (2010).
Hiratani, I. et al. Global reorganization of replication domains during embryonic stem cell differentiation. PLoS Biol. 6, 2220–2236 (2008). First genome-wide profiling of DNA replication timing during embryonic stem cell differentiation, illustrating the link between replication timing and cell fate decisions.
Filion, G. J. et al. Systematic protein location mapping reveals five principal chromatin types in Drosophila cells. Cell 143, 212–224 (2010). Shows that the genome is assembled into five distinctive chromatin types in D. melanogaster using genome-wide location maps for chromatin proteins and histone modifications.
Bell, O. et al. Accessibility of the Drosophila genome discriminates PcG repression, H4K16 acetylation and replication timing. Nature Struct. Mol. Biol. 17, 894–900 (2010).
Hansen, R. S. et al. Sequencing newly replicated DNA reveals widespread plasticity in human replication timing. Proc. Natl Acad. Sci. USA 107, 139–144 (2010).
Schwaiger, M. et al. Chromatin state marks cell-type and gender-specific replication of the Drosophila genome. Genes Dev. 23, 589–601 (2009).
Lieberman-Aiden, E. et al. Comprehensive mapping of long-range interactions reveals folding principles of the human genome. Science 326, 289–293 (2009). Provides genome-wide maps of long-range chromatin interactions generated by high resolution chromatin conformation capture methods (Hi-C).
Ryba, T. et al. Evolutionarily conserved replication timing profiles predict long-range chromatin interactions and distinguish closely related cell types. Genome Res. 20, 761–770 (2010). Reveals a strong correlation between replication timing and Hi-C-defined three-dimensional organization of the genome, supporting the theory that replication timing domains are spatially compartmentalized functional three-dimensional chromosomal units.
De, S. & Michor, F. DNA replication timing and long-range DNA interactions predict mutational landscapes of cancer genomes. Nature Biotech. 29, 1103–1108 (2011). Integration of data on cancer-associated somatic copy number alterations with DNA replication timing and DNA long-range interactions (Hi-C data), suggesting that the genome-wide mutational landscape in cancer in part reflects that spontaneous breaks arising during replication are erroneously repaired using regions of microhomology in three-dimensional proximity.
Francis, N. J., Follmer, N. E., Simon, M. D., Aghia, G. & Butler, J. D. Polycomb proteins remain bound to chromatin and DNA during DNA replication in vitro. Cell 137, 110–122 (2009).
Kuipers, M. A. et al. Highly stable loading of Mcm proteins onto chromatin in living cells requires replication to unload. J. Cell Biol. 192, 29–41 (2011).
Alexandrow, M. G. & Hamlin, J. L. Chromatin decondensation in S-phase involves recruitment of Cdk2 by Cdc45 and histone H1 phosphorylation. J. Cell Biol. 168, 875–886 (2005).
Thomson, A. M., Gillespie, P. J. & Blow, J. J. Replication factory activation can be decoupled from the replication timing program by modulating Cdk levels. J. Cell Biol. 188, 209–221 (2010).
Contreras, A. et al. The dynamic mobility of histone H1 is regulated by cyclin/CDK phosphorylation. Mol. Cell. Biol. 23, 8626–8636 (2003).
Thiriet, C. & Hayes, J. J. Linker histone phosphorylation regulates global timing of replication origin firing. J. Biol. Chem. 284, 2823–2829 (2009).
Cardoso, M. C., Leonhardt, H. & Nadal-Ginard, B. Reversal of terminal differentiation and control of DNA replication: cyclin A and Cdk2 specifically localize at subnuclear sites of DNA replication. Cell 74, 979–992 (1993).
Koundrioukoff, S. et al. A direct interaction between proliferating cell nuclear antigen (PCNA) and Cdk2 targets PCNA-interacting proteins for phosphorylation. J. Biol. Chem. 275, 22882–22887 (2000).
Chibazakura, T. et al. Cyclin A promotes S-phase entry via interaction with the replication licensing factor Mcm7. Mol. Cell Biol. 31, 248–255 (2011).
Katsuno, Y. et al. Cyclin A–Cdk1 regulates the origin firing program in mammalian cells. Proc. Natl Acad. Sci. USA 106, 3184–3189 (2009).
Sogo, J. M., Stahl, H., Koller, T. & Knippers, R. Structure of replicating simian virus 40 minichromosomes. The replication fork, core histone segregation and terminal structures. J. Mol. Biol. 189, 189–204 (1986).
Gasser, R., Koller, T. & Sogo, J. M. The stability of nucleosomes at the replication fork. J. Mol. Biol. 258, 224–239 (1996).
Groth, A. Replicating chromatin: a tale of histones. Biochem. Cell Biol. 87, 51–63 (2009).
Xu, M. et al. Partitioning of histone H3H4 tetramers during DNA replication-dependent chromatin assembly. Science 328, 94–98 (2010). Quantitative mass spectrometry analysis of cell lines conditionally expressing tagged H3.1 and H3.3, demonstrating that splitting of old (H3.1–H4) 2 tetramers is rare in HeLa cells.
Jackson, V. & Chalkley, R. A re-evaluation of new histone deposition on replicating chromatin. J. Biol. Chem. 256, 5095–5103 (1981).
Radman-Livaja, M. et al. Patterns and mechanisms of ancestral histone protein inheritance in budding yeast. PLoS Biol. 9, e1001075 (2011).
Ishimi, Y., Ichinose, S., Omori, A., Sato, K. & Kimura, H. Binding of human minichromosome maintenance proteins with histone H3. J. Biol. Chem. 271, 24115–24122 (1996).
Ramsperger, U. & Stahl, H. Unwinding of chromatin by the SV40 large T antigen DNA helicase. EMBO J. 14, 3215–3225 (1995).
Groth, A. et al. Regulation of replication fork progression through histone supply and demand. Science 318, 1928–1931 (2007).
Jasencakova, Z. et al. Replication stress interferes with histone recycling and predeposition marking of new histones. Mol. Cell 37, 736–743 (2010). Provides the PTM profile of histones associated with ASF1 in S phase and reveals that replication stress accelerates mono-methylation of H3K9 on new histones and impairs histone recycling.
English, C. M., Adkins, M. W., Carson, J. J., Churchill, M. E. & Tyler, J. K. Structural basis for the histone chaperone activity of Asf1. Cell 127, 495–508 (2006).
Natsume, R. et al. Structure and function of the histone chaperone CIA/ASF1 complexed with histones H3 and H4. Nature 446, 338–341 (2007).
Duro, E. et al. Identification of the MMS22L-TONSL complex that promotes homologous recombination. Mol. Cell 40, 632–644 (2010).
Tan, B. C., Chien, C. T., Hirose, S. & Lee, S. C. Functional cooperation between FACT and MCM helicase facilitates initiation of chromatin DNA replication. EMBO J. 25, 3975–3985 (2006).
VanDemark, A. P. et al. The structure of the yFACT Pob3-M domain, its interaction with the DNA replication factor RPA, and a potential role in nucleosome deposition. Mol. Cell 22, 363–374 (2006).
Wittmeyer, J. & Formosa, T. The Saccharomyces cerevisiae DNA polymerase α catalytic subunit interacts with Cdc68/Spt16 and with Pob3, a protein similar to an HMG1-like protein. Mol. Cell. Biol. 17, 4178–4190 (1997).
Formosa, T. The role of FACT in making and breaking nucleosomes. Biochim. Biophys. Acta 23 Jul 2011 (doi:10.1016/j.bbagrm.2011.07.009).
Abe, T. et al. The histone chaperone FACT maintains normal replication fork rates. J. Biol. Chem. 286, 30504–30512 (2011).
Winkler, D. D., Muthurajan, U. M., Hieb, A. R. & Luger, K. Histone chaperone FACT coordinates nucleosome interaction through multiple synergistic binding events. J. Biol. Chem. 286, 41883–41892 (2011).
Marzluff, W. F., Wagner, E. J. & Duronio, R. J. Metabolism and regulation of canonical histone mRNAs: life without a poly(A) tail. Nature Rev. Genet. 9, 843–854 (2008).
Annunziato, A. T. Assembling chromatin: the long and winding road. Biochim. Biophys. Acta 18 Jul 2011 (doi:10.1016/j.bbagrm.2011.07.005).
Smith, S. & Stillman, B. Purification and characterization of CAFI, a human cell factor required for chromatin assembly during DNA replication in vitro. Cell 58, 15–25 (1989).
Campos, E. I. et al. The program for processing newly synthesized histones H3.1 and H4. Nature Struct. Mol. Biol. 17, 1343–1351 (2010).
Alvarez, F. et al. Sequential establishment of marks on soluble histones h3 and h4. J. Biol. Chem. 286, 17714–17721 (2011).
Cook, A. J., Gurard-Levin, Z. A., Vassias, I. & Almouzni, G. A specific function for the histone chaperone NASP to fine-tune a reservoir of soluble H3H4 in the histone supply chain. Mol. Cell 44, 918–927 (2011).
Sobel, R. E., Cook, R. G., Perry, C. A., Annunziato, A. T. & Allis, C. D. Conservation of deposition-related acetylation sites in newly synthesized histones H3 and H4. Proc. Natl Acad. Sci. USA 92, 1237–1241 (1995).
Loyola, A., Bonaldi, T., Roche, D., Imhof, A. & Almouzni, G. PTMs on H3 variants before chromatin assembly potentiate their final epigenetic state. Mol. Cell 24, 309–316 (2006).
Tyler, J. K. et al. The RCAF complex mediates chromatin assembly during DNA replication and repair. Nature 402, 555–560 (1999).
Mello, J. A. et al. Human Asf1 and CAF1 interact and synergize in a repair-coupled nucleosome assembly pathway. EMBO Rep. 3, 329–334 (2002).
Hu, H. et al. Structure of a CENP-A-histone H4 heterodimer in complex with chaperone HJURP. Genes Dev. 25, 901–906 (2011).
Tagami, H., Ray-Gallet, D., Almouzni, G. & Nakatani, Y. Histone H3.1 and H3.3 complexes mediate nucleosome assembly pathways dependent or independent of DNA synthesis. Cell 116, 51–61 (2004).
Groth, A. et al. Human Asf1 regulates the flow of S phase histones during replicational stress. Mol. Cell 17, 301–311 (2005).
Ray-Gallet, D. et al. Dynamics of histone h3 deposition in vivo reveal a nucleosome gap-filling mechanism for h3.3 to maintain chromatin integrity. Mol. Cell 44, 928–941 (2011).
Ahmad, K. & Henikoff, S. The histone variant H3.3 marks active chromatin by replication-independent nucleosome assembly. Mol. Cell 9, 1191–1200 (2002).
Kang, B. et al. Phosphorylation of H4 Ser 47 promotes HIRA-mediated nucleosome assembly. Genes Dev. 25, 1359–1364 (2011).
Masumoto, H., Hawke, D., Kobayashi, R. & Verreault, A. A role for cell-cycle-regulated histone H3 lysine 56 acetylation in the DNA damage response. Nature 436, 294–298 (2005).
Burgess, R. J., Zhou, H., Han, J. & Zhang, Z. A role for Gcn5 in replication-coupled nucleosome assembly. Mol. Cell 37, 469–480 (2010).
Li, Q. et al. Acetylation of histone H3 lysine 56 regulates replication-coupled nucleosome assembly. Cell 134, 244–255 (2008).
Das, C., Lucia, M. S., Hansen, K. C. & Tyler, J. K. CBP/p300-mediated acetylation of histone H3 on lysine 56. Nature 459, 113–117 (2009).
Loyola, A. et al. The HP1α–CAF1–SetDB1-containing complex provides H3K9me1 for Suv39-mediated K9me3 in pericentric heterochromatin. EMBO Rep. 10, 769–775 (2009).
Quivy, J. P., Gerard, A., Cook, A. J., Roche, D. & Almouzni, G. The HP1p150/CAF1 interaction is required for pericentric heterochromatin replication and S-phase progression in mouse cells. Nature Struct. Mol. Biol. 15, 972–979 (2008).
Murzina, N., Verreault, A., Laue, E. & Stillman, B. Heterochromatin dynamics in mouse cells: interaction between chromatin assembly factor 1 and HP1 proteins. Mol. Cell 4, 529–540 (1999).
Probst, A. V. & Almouzni, G. Heterochromatin establishment in the context of genome-wide epigenetic reprogramming. Trends Genet. 27, 177–185 (2011).
Margueron, R. & Reinberg, D. Chromatin structure and the inheritance of epigenetic information. Nature Rev. Genet. 11, 285–296 (2010).
Johnson, A. & O'Donnell, M. Cellular DNA replicases: components and dynamics at the replication fork. Annu. Rev. Biochem. 74, 283–315 (2005).
Shibahara, K. & Stillman, B. Replication-dependent marking of DNA by PCNA facilitates CAF-1-coupled inheritance of chromatin. Cell 96, 575–585 (1999).
Moggs, J. G. et al. A CAF-1-PCNA-mediated chromatin assembly pathway triggered by sensing DNA damage. Mol. Cell Biol. 20, 1206–1218 (2000).
Chafin, D. R., Vitolo, J. M., Henricksen, L. A., Bambara, R. A. & Hayes, J. J. Human DNA ligase I efficiently seals nicks in nucleosomes. EMBO J. 19, 5492–5501 (2000).
Huggins, C. F. et al. Flap endonuclease 1 efficiently cleaves base excision repair and DNA replication intermediates assembled into nucleosomes. Mol. Cell 10, 1201–1211 (2002).
Beattie, T. R. & Bell, S. D. The role of the DNA sliding clamp in Okazaki fragment maturation in archaea and eukaryotes. Biochem. Soc. Trans. 39, 70–76 (2011).
Andrews, A. J., Chen, X., Zevin, A., Stargell, L. A. & Luger, K. The histone chaperone Nap1 promotes nucleosome assembly by eliminating nonnucleosomal histone DNA interactions. Mol. Cell 37, 834–842 (2010).
Nasmyth, K. Cohesin: a catenase with separate entry and exit gates? Nature Cell Biol. 13, 1170–1177 (2011).
Lengronne, A. et al. Establishment of sister chromatid cohesion at the S. cerevisiae replication fork. Mol. Cell 23, 787–799 (2006).
Bermudez, V. P. et al. The alternative Ctf18-Dcc1-Ctf8-replication factor C complex required for sister chromatid cohesion loads proliferating cell nuclear antigen onto DNA. Proc. Natl Acad. Sci. USA 100, 10237–10242 (2003).
Terret, M. E., Sherwood, R., Rahman, S., Qin, J. & Jallepalli, P. V. Cohesin acetylation speeds the replication fork. Nature 462, 231–234 (2009).
Sporbert, A., Gahl, A., Ankerhold, R., Leonhardt, H. & Cardoso, M. C. DNA polymerase clamp shows little turnover at established replication sites but sequential de novo assembly at adjacent origin clusters. Mol. Cell 10, 1355–1365 (2002).
Nakano, S., Stillman, B. & Horvitz, H. R. Replication-coupled chromatin assembly generates a neuronal bilateral asymmetry in C. elegans. Cell 147, 1525–1536 (2011). Study of neural asymmetry in C. elegans showing that replication-coupled nucleosome assembly is required for maintenance and/or establishment of specific gene expression patterns through sequential cell divisions.
Annunziato, A. T. & Seale, R. L. Histone deacetylation is required for the maturation of newly replicated chromatin. J. Biol. Chem. 258, 12675–12684 (1983).
Fisher, D. & Mechali, M. Vertebrate HoxB gene expression requires DNA replication. EMBO J. 22, 3737–3748 (2003).
Kloc, A., Zaratiegui, M., Nora, E. & Martienssen, R. RNA interference guides histone modification during the S phase of chromosomal replication. Curr. Biol. 18, 490–495 (2008).
Chen, E. S. et al. Cell cycle control of centromeric repeat transcription and heterochromatin assembly. Nature 451, 734–737 (2008).
Perry, C. A. & Annunziato, A. T. Influence of histone acetylation on the solubility, H1 content and DNase I sensitivity of newly assembled chromatin. Nucleic Acids Res. 17, 4275–4291 (1989).
Taddei, A., Maison, C., Roche, D. & Almouzni, G. Reversible disruption of pericentric heterochromatin and centromere function by inhibiting deacetylases. Nature Cell Biol. 3, 114–120 (2001).
Conti, C. et al. Inhibition of histone deacetylase in cancer cells slows down replication forks, activates dormant origins, and induces DNA damage. Cancer Res. 70, 4470–4480 (2010).
Bhaskara, S. et al. Hdac3 is essential for the maintenance of chromatin structure and genome stability. Cancer Cell 18, 436–447 (2010). Demonstrates that failure in heterochromatin maintenance can promote genomic instability and tumour development in Hdac3 -null mice.
Sirbu, B. M. et al. Analysis of protein dynamics at active, stalled, and collapsed replication forks. Genes Dev. 25, 1320–1327 (2011). Describes, together with reference 177, a powerful method to biochemically isolate proteins on newly replicated DNA. In both techniques (iPOND and DNA-mediated chromatin pull-down) DNA is labelled by EdU, which after cell lysis is coupled to biotin by Click-iT chemistry.
Milutinovic, S., Zhuang, Q. & Szyf, M. Proliferating cell nuclear antigen associates with histone deacetylase activity, integrating DNA replication and chromatin modification. J. Biol. Chem. 277, 20974–20978 (2002).
Rowbotham, S. P. et al. Maintenance of silent chromatin through replication requires SWI/SNF-like chromatin remodeler SMARCAD1. Mol. Cell 42, 285–296 (2011).
Taddei, A., Roche, D., Sibarita, J. B., Turner, B. M. & Almouzni, G. Duplication and maintenance of heterochromatin domains. J. Cell Biol. 147, 1153–1166 (1999).
Lande-Diner, L., Zhang, J. & Cedar, H. Shifts in replication timing actively affect histone acetylation during nucleosome reassembly. Mol. Cell 34, 767–774 (2009).
Jones, P. A. & Liang, G. Rethinking how DNA methylation patterns are maintained. Nature Rev. Genet. 10, 805–811 (2009).
Egger, G. et al. Identification of DNMT1 (DNA methyltransferase 1) hypomorphs in somatic knockouts suggests an essential role for DNMT1 in cell survival. Proc. Natl Acad. Sci. USA 103, 14080–14085 (2006).
Schermelleh, L. et al. Dynamics of Dnmt1 interaction with the replication machinery and its role in postreplicative maintenance of DNA methylation. Nucleic Acids Res. 35, 4301–4312 (2007).
Wu, H. & Zhang, Y. Mechanisms and functions of Tet protein-mediated 5-methylcytosine oxidation. Genes Dev. 25, 2436–2452 (2011).
Poot, R. A. et al. The Williams syndrome transcription factor interacts with PCNA to target chromatin remodelling by ISWI to replication foci. Nature Cell Biol. 6, 1236–1244 (2004).
Scharf, A. N., Barth, T. K. & Imhof, A. Establishment of histone modifications after chromatin assembly. Nucleic Acids Res. 37, 5032–5040 (2009).
Xu, M., Wang, W., Chen, S. & Zhu, B. A model for mitotic inheritance of histone lysine methylation. EMBO Rep. 13, 60–67 (2011). References 129 and 130 conducted global analysis of histone PTMs during cell division by quantitative proteomics. Reference 129 provided the first indication that some histone H3 marks (K9me3 and K27me3) are not restored prior to mitosis. Reference 130 establishes the kinetics by which key H3 marks (K9me1, K9me2 and K9me3 and K27me1, K27me2 and K27me3) are restored and proposes that fluctuation of methylation over a silenced domain is tolerated.
Esteve, P. O. et al. Direct interaction between DNMT1 and G9a coordinates DNA and histone methylation during replication. Genes Dev. 20, 3089–3103 (2006).
Sarraf, S. A. & Stancheva, I. Methyl-CpG binding protein MBD1 couples histone H3 methylation at lysine 9 by SETDB1 to DNA replication and chromatin assembly. Mol. Cell 15, 595–605 (2004).
Raynaud, C. et al. Two cell-cycle regulated SET-domain proteins interact with proliferating cell nuclear antigen (PCNA) in Arabidopsis. Plant J. 47, 395–407 (2006).
Jacob, Y. et al. Regulation of heterochromatic DNA replication by histone H3 lysine 27 methyltransferases. Nature 466, 987–991 (2010).
Zaratiegui, M. et al. RNAi promotes heterochromatic silencing through replication-coupled release of RNA Pol II. Nature 479, 135–138 (2011).
Li, F., Martienssen, R. & Cande, W. Z. Coordination of DNA replication and histone modification by the Rik1-Dos2 complex. Nature 475, 244–248 (2011). Shows that the replisome itself can directly recruit a repressor complex required for heterochromatin maintenance in S. pombe.
Pesavento, J. J., Yang, H., Kelleher, N. L. & Mizzen, C. A. Certain and progressive methylation of histone H4 at lysine 20 during the cell cycle. Mol. Cell. Biol. 28, 468–486 (2008).
Lanzuolo, C., Lo Sardo, F., Diamantini, A. & Orlando, V. PcG complexes set the stage for epigenetic inheritance of gene silencing in early S phase before replication. PLoS Genet. 7, e1002370 (2011).
Dodd, I. B., Micheelsen, M. A., Sneppen, K. & Thon, G. Theoretical analysis of epigenetic cell memory by nucleosome modification. Cell 129, 813–822 (2007).
Aagaard, L. et al. Functional mammalian homologues of the Drosophila PEV-modifier Su(var)3–9 encode centromere-associated proteins which complex with the heterochromatin component M31. EMBO J. 18, 1923–1938 (1999).
Hansen, K. H. et al. A model for transmission of the H3K27me3 epigenetic mark. Nature Cell Biol. 10, 1291–1300 (2008).
Margueron, R. et al. Role of the polycomb protein EED in the propagation of repressive histone marks. Nature 461, 762–767 (2009). Shows, together with reference 141, that PRC2 can be recruited to H3K27me3, suggesting that this mark can be transmitted by a self-reinforcing loop.
De Vos, D. et al. Progressive methylation of ageing histones by Dot1 functions as a timer. EMBO Rep. 12, 956–962 (2011).
Sweet, S. M., Li, M., Thomas, P. M., Durbin, K. R. & Kelleher, N. L. Kinetics of re-establishing H3K79 methylation marks in global human chromatin. J. Biol. Chem. 285, 32778–32786 (2010).
Xu, G. L. et al. Chromosome instability and immunodeficiency syndrome caused by mutations in a DNA methyltransferase gene. Nature 402, 187–191 (1999).
Peters, A. H. et al. Loss of the Suv39h histone methyltransferases impairs mammalian heterochromatin and genome stability. Cell 107, 323–337 (2001).
Gaudet, F. et al. Induction of tumors in mice by genomic hypomethylation. Science 300, 489–492 (2003).
Hansen, K. D. et al. Increased methylation variation in epigenetic domains across cancer types. Nature Genet. 43, 768–775 (2011).
Fraga, M. F. et al. Loss of acetylation at Lys16 and trimethylation at Lys20 of histone H4 is a common hallmark of human cancer. Nature Genet. 37, 391–400 (2005).
Wen, B., Wu, H., Shinkai, Y., Irizarry, R. A. & Feinberg, A. P. Large histone H3 lysine 9 dimethylated chromatin blocks distinguish differentiated from embryonic stem cells. Nature Genet. 41, 246–250 (2009).
Clemente-Ruiz, M. & Prado, F. Chromatin assembly controls replication fork stability. EMBO Rep. 10, 790–796 (2009).
Myung, K., Pennaneach, V., Kats, E. S. & Kolodner, R. D. Saccharomyces cerevisiae chromatin-assembly factors that act during DNA replication function in the maintenance of genome stability. Proc. Natl Acad. Sci. USA 100, 6640–6645 (2003).
Prado, F., Cortes-Ledesma, F. & Aguilera, A. The absence of the yeast chromatin assembly factor Asf1 increases genomic instability and sister chromatid exchange. EMBO Rep. 5, 497–502 (2004).
Yang, J. H. & Freudenreich, C. H. The Rtt109 histone acetyltransferase facilitates error-free replication to prevent CAG/CTG repeat contractions. DNA Repair (Amst.) 9, 414–420 (2010).
Feser, J. et al. Elevated histone expression promotes life span extension. Mol. Cell 39, 724–735 (2010).
O'Sullivan, R. J., Kubicek, S., Schreiber, S. L. & Karlseder, J. Reduced histone biosynthesis and chromatin changes arising from a damage signal at telomeres. Nature Struct. Mol. Biol. 17, 1218–1225 (2010).
Sanchez-Molina, S. et al. Role for hACF1 in the G2/M damage checkpoint. Nucleic Acids Res. 39, 8445–8456 (2011).
Bester, A. C. et al. Nucleotide deficiency promotes genomic instability in early stages of cancer development. Cell 145, 435–446 (2011).
Sarkies, P., Reams, C., Simpson, L. J. & Sale, J. E. Epigenetic instability due to defective replication of structured DNA. Mol. Cell 40, 703–713 (2010). This study shows that G4 DNA structures that interfere with the fork progression can jeopardize chromatin maintenance and challenge epigenetic silencing.
Nyce, J., Liu, L. & Jones, P. A. Variable effects of DNA-synthesis inhibitors upon DNA methylation in mammalian cells. Nucleic Acids Res. 14, 4353–4367 (1986).
Lopes, M., Foiani, M. & Sogo, J. M. Multiple mechanisms control chromosome integrity after replication fork uncoupling and restart at irreparable UV lesions. Mol. Cell 21, 15–27 (2006).
Daigaku, Y., Davies, A. A. & Ulrich, H. D. Ubiquitin-dependent DNA damage bypass is separable from genome replication. Nature 465, 951–955 (2010).
Polo, S. E., Roche, D. & Almouzni, G. New histone incorporation marks sites of UV repair in human cells. Cell 127, 481–493 (2006).
Singh, G. & Klar, A. J. Mutations in deoxyribonucleotide biosynthesis pathway cause spreading of silencing across heterochromatic barriers at the mating-type region of the fission yeast. Yeast 25, 117–128 (2008).
Zaratiegui, M. et al. CENP-B preserves genome integrity at replication forks paused by retrotransposon LTR. Nature 469, 112–115 (2011).
Dubarry, M., Loiodice, I., Chen, C. L., Thermes, C. & Taddei, A. Tight protein-DNA interactions favor gene silencing. Genes Dev. 25, 1365–1370 (2011). Demonstrates that replication fork pausing can facilitate gene silencing by promoting ectopic recruitment of the SIR proteins to chromatin.
Saveliev, A., Everett, C., Sharpe, T., Webster, Z. & Festenstein, R. DNA triplet repeats mediate heterochromatin-protein-1-sensitive variegated gene silencing. Nature 422, 909–913 (2003).
Mirkin, S. M. Expandable DNA repeats and human disease. Nature 447, 932–940 (2007).
Di Micco, R. et al. Interplay between oncogene-induced DNA damage response and heterochromatin in senescence and cancer. Nature Cell Biol. 13, 292–302 (2011).
Shimada, K. et al. Ino80 chromatin remodeling complex promotes recovery of stalled replication forks. Curr. Biol. 18, 566–575 (2008).
Falbo, K. B. et al. Involvement of a chromatin remodeling complex in damage tolerance during DNA replication. Nature Struct. Mol. Biol. 16, 1167–1172 (2009).
Papamichos-Chronakis, M. & Peterson, C. L. The Ino80 chromatin-remodeling enzyme regulates replisome function and stability. Nature Struct. Mol. Biol. 15, 338–345 (2008).
O'Donnell, L. et al. The MMS22L-TONSL complex mediates recovery from replication stress and homologous recombination. Mol. Cell 40, 619–631 (2010).
O'Connell, B. C. et al. A genome-wide camptothecin sensitivity screen identifies a mammalian MMS22L-NFKBIL2 complex required for genomic stability. Mol. Cell 40, 645–657 (2010).
Piwko, W. et al. RNAi-based screening identifies the Mms22L-Nfkbil2 complex as a novel regulator of DNA replication in human cells. EMBO J. 29, 4210–4222 (2010).
Takeda, S. et al. BRU1, a novel link between responses to DNA damage and epigenetic gene silencing in Arabidopsis. Genes Dev. 18, 782–793 (2004).
Kliszczak, A. E., Rainey, M. D., Harhen, M., Boisvert, F. M. & Santocanale, C. DNA mediated chromatin pull-down for the study of chromatin replication. Scientific Reports 1, 95 (2011).
Bochman, M. L. & Schwacha, A. The Mcm complex: unwinding the mechanism of a replicative helicase. Microbiol. Mol. Biol. Rev. 73, 652–683 (2009).
Costa, A. et al. The structural basis for MCM2–7 helicase activation by GINS and Cdc45. Nature Struct. Mol. Biol. 18, 471–477 (2011).
Fu, Y. V. et al. Selective bypass of a lagging strand roadblock by the eukaryotic replicative DNA helicase. Cell 146, 931–941 (2011).
Errico, A. & Costanzo, V. Differences in the DNA replication of unicellular eukaryotes and metazoans: known unknowns. EMBO Rep. 11, 270–278 (2010).
Pak, D. T. et al. Association of the origin recognition complex with heterochromatin and HP1 in higher eukaryotes. Cell 91, 311–323 (1997).
Prasanth, S. G., Shen, Z., Prasanth, K. V. & Stillman, B. Human origin recognition complex is essential for HP1 binding to chromatin and heterochromatin organization. Proc. Natl Acad. Sci. USA 107, 15093–15098 (2010).
Triolo, T. & Sternglanz, R. Role of interactions between the origin recognition complex and SIR1 in transcriptional silencing. Nature 381, 251–253 (1996).
Iizuka, M. & Stillman, B. Histone acetyltransferase HBO1 interacts with the ORC1 subunit of the human initiator protein. J. Biol. Chem. 274, 23027–23034 (1999).
Burke, T. W., Cook, J. G., Asano, M. & Nevins, J. R. Replication factors MCM2 and ORC1 interact with the histone acetyltransferase HBO1. J. Biol. Chem. 276, 15397–15408 (2001).
Shen, Z. et al. A WD-repeat protein stabilizes ORC binding to chromatin. Mol. Cell 40, 99–111 (2010).
Dhar, S. K. et al. Replication from oriP of Epstein-Barr virus requires human ORC and is inhibited by geminin. Cell 106, 287–296 (2001).
Li, P. C., Chretien, L., Cote, J., Kelly, T. J. & Forsburg, S. L. S. pombe replication protein Cdc18 (Cdc6) interacts with Swi6 (HP1) heterochromatin protein: region specific effects and replication timing in the centromere. Cell Cycle 10, 323–336 (2011).
Doyon, Y. et al. ING tumor suppressor proteins are critical regulators of chromatin acetylation required for genome expression and perpetuation. Mol. Cell 21, 51–64 (2006).
Ryu, M. J. et al. Direct interaction between cohesin complex and DNA replication machinery. Biochem. Biophys. Res. Commun. 341, 770–775 (2006).
Franco, A. A., Lam, W. M., Burgers, P. M. & Kaufman, P. D. Histone deposition protein Asf1 maintains DNA replisome integrity and interacts with replication factor C. Genes Dev. 19, 1365–1375 (2005).
Chuang, L. S. et al. Human DNA-(cytosine-5) methyltransferase-PCNA complex as a target for p21WAF1. Science 277, 1996–2000 (1997).
Shumaker, D. K. et al. The highly conserved nuclear lamin Ig-fold binds to PCNA: its role in DNA replication. J. Cell Biol. 181, 269–280 (2008).
Moldovan, G. L., Pfander, B. & Jentsch, S. PCNA controls establishment of sister chromatid cohesion during S phase. Mol. Cell 23, 723–732 (2006).
Huen, M. S., Sy, S. M., van Deursen, J. M. & Chen, J. Direct interaction between SET8 and proliferating cell nuclear antigen couples H4K20 methylation with DNA replication. J. Biol. Chem. 283, 11073–11077 (2008).
Jorgensen, S. et al. The histone methyltransferase SET8 is required for S-phase progression. J. Cell Biol. 179, 1337–1345 (2007).
Liang, Z., Diamond, M., Smith, J. A., Schnell, M. & Daniel, R. Proliferating cell nuclear antigen is required for loading of the SMCX/KMD5C histone demethylase onto chromatin. Epigenetics Chromatin 4, 18 (2011).
Hasan, S., Hassa, P. O., Imhof, R. & Hottiger, M. O. Transcription coactivator p300 binds PCNA and may have a role in DNA repair synthesis. Nature 410, 387–391 (2001).
Hasan, S. et al. Regulation of human flap endonuclease1 activity by acetylation through the transcriptional coactivator p300. Mol. Cell 7, 1221–1231 (2001).
Balakrishnan, L., Stewart, J., Polaczek, P., Campbell, J. L. & Bambara, R. A. Acetylation of Dna2 endonuclease/helicase and flap endonuclease 1 by p300 promotes DNA stability by creating long flap intermediates. J. Biol. Chem. 285, 4398–4404 (2010).
Acknowledgements
We apologize to those whose publications we were unable to cite owing to space limitations. We would like to thank Z. Jasencakova and R. Pocock for critical reading of this manuscript, and our anonymous reviewers, I. Kamalyukova, C. B. Stromme and R. Martienssen for useful comments. C.A. is supported by a Human Frontiers Scientific Program long-term fellowship and the Danish Medical Research council (FSS). The A.G. laboratory is supported by a European Research Council Starting Grant (ERC2011StG, no. 281,765), the Lundbeck Foundation, the Danish Cancer Society, the Novo Nordisk Foundation, the Danish Medical Research Council and European Commission ITN FP7-PEOPLE2008 'Nucleosome4D'.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Related links
Related links
FURTHER INFORMATION
Glossary
- Epigenetics
-
The studies of heritable changes in genome function that occur without a change in DNA sequence.
- Replication stress
-
General term referring to deregulation of replication. This can include fork problems (change of speed, stalling or collapse) and replication initiation defects.
- Epigenome
-
The epigenome refers to the overall epigenetic state of a cell, including histone and DNA marks, histone variants, nucleosome positioning and higher-order structures.
- Chromosomal architecture
-
Three-dimensional organization of chromosomes in the nucleus. For example, each chromosome occupies a territory in the nucleus and will take up a specific higher-order structure of open and compact domains that is partly cell type specific.
- Nucleosome assembly
-
A stepwise process starting with the deposition of two H3–H4 dimers or potentially a (H3–H4)2 tetramer onto DNA to form a tetrasome. This is followed by the incorporation of two H2A–H2B dimers to form a nucleosome core particle.
- Histone variants
-
Replacement histones differ in amino acid sequence from canonical S phase histones to varying extents. They are often incorporated by dedicated pathways to serve specialized functions.
- Replication origins
-
Sites in the genome where replication initiates, giving rise to two forks that progress away from the origin in opposite directions.
- Nucleosome-free regions
-
(NFRs). Sites of reduced nucleosome occupancy compared with the immediate surrounding regions. NFRs display sensitivity to DNase I, which is likely to result from high histone exchange or DNA structures that resist nucleosome formation.
- Origin decision point
-
(ODP). Transition point in late G1phase that specifies the origins that will fire in the following S phase. It probably represents a change at specific pre-replication complexes (pre-RCs), which potentiates some pre-RCs while preventing others from initiating.
- Replicons
-
Stretches of DNA replicated from a single origin.
- DNA halo assay
-
An approach to visualize DNA loops in interphase nuclei. Nuclei are permeabilized and depleted of histone and soluble proteins on slides, allowing unwinding of supercoiled DNA loops to form a halo around an insoluble scaffold.
- Cohesin complex
-
Ring-shaped multi-protein complex (composed of SMC1, SMC3, RAD21 and sister-chromatid cohesion protein 3 (SCC3)) that by embracing chromatin fibres mediates sister chromatid cohesion and has roles in DNA repair and transcription.
- DNA superhelicity
-
Positive or negative supercoiling of DNA molecules.
- SILAC
-
'Stable isotope labelling with amino acids in cell culture' is an approach for in vivo metabolic labelling of proteins with amino acids containing light or heavy isotopes and is used for quantitative mass spectrometry.
- Histone chaperone
-
Factor that associates with histones and stimulates a reaction involving histone transfer without being part of the final product.
- Sister chromatid cohesion
-
The joining of two sister chromatids upon chromosome replication that enables proper chromosome segregation.
- Okazaki fragment maturation
-
Okazaki fragments are short DNA molecules of approximately 100 to 200 nucleotides in eukaryotes. They are initiated by primase–Pol α (DNA polymerase α) on lagging strands by the synthesis of an RNA primer with a short DNA extension, which is then further extended by Pol δ. The primer and part of the DNA is removed as two Okazaki fragments are ligated together.
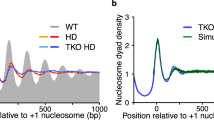
- Nucleosome dyads
-
Axes of symmetry in the nucleosome.
- iPOND
-
'Isolation of proteins on nascent DNA' is a technology to isolate proteins on newly synthesized DNA by combining EdU labelling with Click-iT chemistry.
- RNA interference
-
(RNAi). Processing of transcripts into small double-stranded RNAs that can silence gene expression. Small RNAs can work by interfering with translation to induce post-transcriptional gene silencing or induce chromatin-dependent gene silencing by interacting with nascent transcripts and targeting chromatin-modifying complexes.
- Senescence
-
A state of irreversible cell cycle arrest that occurs as a consequence of continued cell division in primary mammalian cells in part owing to erosion of telomeres. Senescence contributes to organismal ageing and at the same time provides a barrier to carcinogenesis.
- Replicative ageing
-
Accumulation of genetic and epigenetic defects at each round of replication during life span in yeast.
- G-quadruplex structures
-
(G4 structures). Guanine-rich DNA sequences capable of forming four-stranded secondary structures by square arrangement of guanines.
- Chromatin remodeller
-
A large multi-protein machine that, through ATP hydrolysis, enables access to nucleosomal DNA by altering the structure, composition and/or position of nucleosomes.
Rights and permissions
About this article
Cite this article
Alabert, C., Groth, A. Chromatin replication and epigenome maintenance. Nat Rev Mol Cell Biol 13, 153–167 (2012). https://doi.org/10.1038/nrm3288
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrm3288
This article is cited by
-
H2B ubiquitination recruits FACT to maintain a stable altered nucleosome state for transcriptional activation
Nature Communications (2023)
-
H4S47 O-GlcNAcylation regulates the activation of mammalian replication origins
Nature Structural & Molecular Biology (2023)
-
The COMPASS subunit Spp1 protects nascent DNA at the Tus/Ter replication fork barrier by limiting DNA availability to nucleases
Nature Communications (2023)
-
NFIB facilitates replication licensing by acting as a genome organizer
Nature Communications (2023)
-
TOF1 and RRM3 reveal a link between gene silencing and the pausing of replication forks
Current Genetics (2023)