Abstract
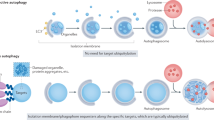
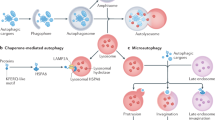
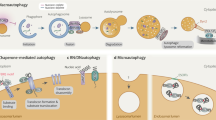
Autophagy not only recycles intracellular components to compensate for nutrient deprivation but also selectively eliminates organelles to regulate their number and maintain quality control. Mitophagy, the specific autophagic elimination of mitochondria, has been identified in yeast, mediated by autophagy-related 32 (Atg32), and in mammals during red blood cell differentiation, mediated by NIP3-like protein X (NIX; also known as BNIP3L). Moreover, mitophagy is regulated in many metazoan cell types by parkin and PTEN-induced putative kinase protein 1 (PINK1), and mutations in the genes encoding these proteins have been linked to forms of Parkinson's disease.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Nakatogawa, H., Suzuki, K., Kamada, Y. & Ohsumi, Y. Dynamics and diversity in autophagy mechanisms: lessons from yeast. Nature Rev. Mol. Cell Biol. 10, 458–467 (2009).
Yang, Z. & Klionsky, D. J. Eaten alive: a history of macroautophagy. Nature Cell Biol. 12, 814–822 (2010).
Mari, M. et al. An Atg9-containing compartment that functions in the early steps of autophagosome biogenesis. J. Cell Biol. 190, 1005–1022 (2010).
Axe, E. L. et al. Autophagosome formation from membrane compartments enriched in phosphatidylinositol 3-phosphate and dynamically connected to the endoplasmic reticulum. J. Cell Biol. 182, 685–701 (2008).
Hailey, D. W. et al. Mitochondria supply membranes for autophagosome biogenesis during starvation. Cell 141, 656–667 (2010).
Ravikumar, B., Moreau, K., Jahreiss, L., Puri, C. & Rubinsztein, D. C. Plasma membrane contributes to the formation of pre-autophagosomal structures. Nature Cell Biol. 12, 747–757 (2010).
Klionsky, D. J. et al. A comprehensive glossary of autophagy-related molecules and processes. Autophagy 6, 438–448 (2010).
Oku, M. & Sakai, Y. Pexophagy in Pichia pastoris. Methods Enzymol. 451, 217–228 (2008).
De Duve, C. & Wattiaux, R. Functions of lysosomes. Annu. Rev. Physiol. 28, 435–492 (1966).
Kim, I., Rodriguez-Enriquez, S. & Lemasters, J. J. Selective degradation of mitochondria by mitophagy. Arch. Biochem. Biophys. 462, 245–253 (2007).
Nakatogawa, H., Ichimura, Y. & Ohsumi, Y. Atg8, a ubiquitin-like protein required for autophagosome formation, mediates membrane tethering and hemifusion. Cell 130, 165–178 (2007).
Noda, N. N., Ohsumi, Y. & Inagaki, F. Atg8-family interacting motif crucial for selective autophagy. FEBS Lett. 584, 1379–1385 (2010).
Tolkovsky, A. M., Xue, L., Fletcher, G. C. & Borutaite, V. Mitochondrial disappearance from cells: a clue to the role of autophagy in programmed cell death and disease? Biochimie 84, 233–240 (2002).
Nowikovsky, K., Reipert, S., Devenish, R. J. & Schweyen, R. J. Mdm38 protein depletion causes loss of mitochondrial K+/H+ exchange activity, osmotic swelling and mitophagy. Cell Death Differ. 14, 1647–1656 (2007).
Twig, G. et al. Fission and selective fusion govern mitochondrial segregation and elimination by autophagy. EMBO J. 27, 433–446 (2008).
Westermann, D. Mitochondrial fusion and fission in cell life and death. Nature Rev. Mol. Cell Biol. 11, 872–884 (2010).
Tal, R., Winter, G., Ecker, N., Klionsky, D. J. & Abeliovich, H. Aup1p, a yeast mitochondrial protein phosphatase homolog, is required for efficient stationary phase mitophagy and cell survival. J. Biol. Chem. 282, 5617–5624 (2007).
Kissova, I., Deffieu, M., Manon, S. & Camougrand, N. Uth1p is involved in the autophagic degradation of mitochondria. J. Biol. Chem. 279, 39068–39074 (2004).
Schweers, R. L. et al. NIX is required for programmed mitochondrial clearance during reticulocyte maturation. Proc. Natl Acad. Sci. USA 104, 19500–19505 (2007).
Kundu, M. et al. Ulk1 plays a critical role in the autophagic clearance of mitochondria and ribosomes during reticulocyte maturation. Blood 112, 1493–1502 (2008).
Okamoto, K., Kondo-Okamoto, N. & Ohsumi, Y. Mitochondria-anchored receptor Atg32 mediates degradation of mitochondria via selective autophagy. Dev. Cell 17, 87–97 (2009).
Kanki, T., Wang, K. & Klionsky, D. J. A genomic screen for yeast mutants defective in mitophagy. Autophagy 6, 278–280 (2010).
Kanki, T., Wang, K., Cao, Y., Baba, M. & Klionsky, D. J. Atg32 is a mitochondrial protein that confers selectivity during mitophagy. Dev. Cell 17, 98–109 (2009).
Mortensen, M. et al. Loss of autophagy in erythroid cells leads to defective removal of mitochondria and severe anemia in vivo. Proc. Natl Acad. Sci. USA 107, 832–837 (2010).
Zhang, J. et al. Mitochondrial clearance is regulated by Atg7-dependent and -independent mechanisms during reticulocyte maturation. Blood 114, 157–164 (2009).
Aerbajinai, W., Giattina, M., Lee, Y. T., Raffeld, M. & Miller, J. L. The proapoptotic factor Nix is coexpressed with Bcl-xL during terminal erythroid differentiation. Blood 102, 712–717 (2003).
Sandoval, H. et al. Essential role for Nix in autophagic maturation of erythroid cells. Nature 454, 232–235 (2008).
Aouacheria, A., Brunet, F. & Gouy, M. Phylogenomics of life-or-death switches in multicellular animals: Bcl-2, BH3-only, and BNip families of apoptotic regulators. Mol. Biol. Evol. 22, 2395–2416 (2005).
Schwarten, M. et al. Nix directly binds to GABARAP: a possible crosstalk between apoptosis and autophagy. Autophagy 5, 690–698 (2009).
Novak, I. et al. Nix is a selective autophagy receptor for mitochondrial clearance. EMBO Rep. 11, 45–51 (2010).
Valente, E. M. et al. Hereditary early-onset Parkinson's disease caused by mutations in PINK1. Science 304, 1158–1160 (2004).
Kitada, T. et al. Mutations in the parkin gene cause autosomal recessive juvenile parkinsonism. Nature 392, 605–608 (1998).
Greene, J. C. et al. Mitochondrial pathology and apoptotic muscle degeneration in Drosophila parkin mutants. Proc. Natl Acad. Sci. USA 100, 4078–4083 (2003).
Park, J. et al. Mitochondrial dysfunction in Drosophila PINK1 mutants is complemented by parkin. Nature 441, 1157–1161 (2006).
Clark, I. E. et al. Drosophila pink1 is required for mitochondrial function and interacts genetically with parkin. Nature 441, 1162–1166 (2006).
Poole, A. C. et al. The PINK1/Parkin pathway regulates mitochondrial morphology. Proc. Natl Acad. Sci. USA 105, 1638–1643 (2008).
Yang, Y. et al. Pink1 regulates mitochondrial dynamics through interaction with the fission/fusion machinery. Proc. Natl Acad. Sci. USA 105, 7070–7075 (2008).
Deng, H., Dodson, M. W., Huang, H. & Guo, M. The Parkinson's disease genes pink1 and parkin promote mitochondrial fission and/or inhibit fusion in Drosophila. Proc. Natl Acad. Sci. USA 105, 14503–14508 (2008).
Suen, D. F., Narendra, D. P., Tanaka, A., Manfredi, G. & Youle, R. J. Parkin overexpression selects against a deleterious mtDNA mutation in heteroplasmic cybrid cells. Proc. Natl Acad. Sci. USA 107, 11835–11840 (2010).
Narendra, D., Tanaka, A., Suen, D. F. & Youle, R. J. Parkin is recruited selectively to impaired mitochondria and promotes their autophagy. J. Cell Biol. 183, 795–803 (2008).
Narendra, D. P. et al. PINK1 is selectively stabilized on impaired mitochondria to activate Parkin. PLoS Biol. 8, e1000298 (2010).
Vives-Bauza, C. et al. PINK1-dependent recruitment of Parkin to mitochondria in mitophagy. Proc. Natl Acad. Sci. USA 107, 378–383 (2010).
Geisler, S. et al. PINK1/Parkin-mediated mitophagy is dependent on VDAC1 and p62/SQSTM1. Nature Cell Biol. 12, 119–131 (2010).
Matsuda, N. et al. PINK1 stabilized by mitochondrial depolarization recruits Parkin to damaged mitochondria and activates latent Parkin for mitophagy. J. Cell Biol. 189, 211–221 (2010).
Kim, Y. et al. PINK1 controls mitochondrial localization of Parkin through direct phosphorylation. BBRC 377, 975–980 (2008).
Kawajiri, S. et al. PINK1 is recruited to mitochondria with parkin and associates with LC3 in mitophagy. FEBS Lett. 584, 1073–1079 (2010).
Yang, Y. et al. Mitochondrial pathology and muscle and dopaminergic neuron degeneration caused by inactivation of Drosophila Pink1 is rescued by Parkin. Proc. Natl Acad. Sci. USA 103, 10793–10798 (2006).
Ding, W. X. et al. Nix is critical to two distinct phases of mitophagy, reactive oxygen species-mediated autophagy induction and Parkin–ubiquitin–p62- mediated mitochondrial priming. J. Biol. Chem. 285, 27879–27890 (2010).
Lin, W. & Kang, U. J. Characterization of PINK1 processing, stability, and subcellular localization. J. Neurochem. 106, 464–474 (2008).
Whitworth, A. J. et al. Rhomboid-7 and HtrA2/Omi act in a common pathway with the Parkinson's disease factors Pink1 and Parkin. Dis. Model Mech. 1, 168–174 (2008).
Shiba, K. et al. Parkin stabilizes PINK1 through direct interaction. Biochem. Biophys. Res. Commun. 383, 331–335 (2009).
Um, J. W., Stichel-Gunkel, C., Lübbert, H., Lee, G. & Chung, K. C. Molecular interaction between Parkin and Pink1 in mammalian neuronal cells. Mol. Cell Neurosci. 40, 421–432 (2009).
Sha, D., Chin, L. S. & Li, L. Phosphorylation of parkin by Parkinson disease-linked kinase PINK1 activates parkin E3 ligase function and NF-κB signaling. Hum. Mol. Genet. 19, 352–363 (2010).
Lee, J. Y., Nagano, Y., Taylor, J. P., Lim, K. L. & Yao, T. P. Disease-causing mutations in Parkin impair mitochondrial ubiquitination, aggregation, and HDAC6-dependent mitophagy. J. Cell Biol. 189, 671–679 (2010).
Okatsu, K. et al. p62/SQSTM1 cooperates with Parkin for perinuclear clustering of depolarized mitochondria. Genes Cells 15, 887–900 (2010).
Narendra, D., Kane, L., Hauser, D., Fearnley, I. & Youle, R. p62/SQSTM1 is required for Parkin-induced mitochondrial clustering but not mitophagy; VDAC1 is dispensible for both. Autophagy 6, 1090–1106 (2010).
Pankiv, S. et al. p62/SQSTM1 binds directly to Atg8/LC3 to facilitate degradation of ubiquitinated protein aggregates by autophagy. J. Biol. Chem. 282, 24131–24145 (2007).
Dawson, T. M. & Dawson, V. L. The role of parkin in familial and sporadic Parkinson's disease. Mov. Disord. 25 (Suppl. 1), S32–S39 (2010).
Ziviani, E., Tao, R. N. & Whitworth, A. J. Drosophila parkin requires PINK1 for mitochondrial translocation and ubiquitinates mitofusin. Proc. Natl Acad. Sci. USA 107, 5018–5023 (2010).
Poole, A. C., Thomas, R. E., Yu, S., Vincow, E. S. & Pallanck, L. The mitochondrial fusion-promoting factor mitofusin is a substrate of the PINK1/parkin pathway. PLoS ONE 5, e10054 (2010).
Gegg, M. E. et al. Mitofusin 1 and mitofusin 2 are ubiquitinated in a PINK1/parkin-dependent manner upon induction of mitophagy. Hum. Mol. Genet. 19, 4861–4870 (2010).
Tanaka, A. et al. Proteasome and p97 mediate mitophagy and degradation of mitofusins induced by Parkin. J. Cell Biol. (in the press; doi:10.1083/jcb.201007013).
Geisler, S. et al. The PINK1/Parkin-mediated mitophagy is compromised by PD-associated mutations. Autophagy 6, 871–878 (2010).
Cesari, R. et al. Parkin, a gene implicated in autosomal recessive juvenile parkinsonism, is a candidate tumor suppressor gene on chromosome 6q25-q27. Proc. Natl Acad. Sci. USA 100, 5956–5961 (2003).
Poulogiannis, G. et al. PARK2 deletions occur frequently in sporadic colorectal cancer and accelerate adenoma development in Apc mutant mice. Proc. Natl Acad. Sci. USA 107, 15145–15150 (2010).
Klein, C. & Schlossmacher, M. G. The genetics of Parkinson disease: implications for neurological care. Nature Clin. Pract. Neurol. 2, 136–146 (2006).
Schapira, A. H. Mitochondria in the aetiology and pathogenesis of Parkinson's disease. Lancet Neurol. 7, 97–109 (2008).
Kraytsberg, Y. et al. Mitochondrial DNA deletions are abundant and cause functional impairment in aged human substantia nigra neurons. Nature Genet. 38, 518–520 (2006).
Bender, A. et al. High levels of mitochondrial DNA deletions in substantia nigra neurons in aging and Parkinson disease. Nature Genet. 38, 515–517 (2006).
Luoma, P. et al. Parkinsonism, premature menopause, and mitochondrial DNA polymerase γ mutations: clinical and molecular genetic study. Lancet 364, 875–882 (2004).
Reeve, A. K. et al. Nature of mitochondrial DNA deletions in substantia nigra neurons. Am. J. Hum. Genet. 82, 228–235 (2008).
Baloh, R. H., Salavaggione, E., Milbrandt, J. & Pestronk, A. Familial parkinsonism and ophthalmoplegia from a mutation in the mitochondrial DNA helicase twinkle. Arch. Neurol. 64, 998–1000 (2007).
Acknowledgements
We thank H. Abeliovich for valuable discussions and D. Suen for the movie that is used in Supplementary information 1. D.P.N. is a member of the US National Institutes of Health–Oxford–Cambridge Scholars Program.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Related links
Rights and permissions
About this article
Cite this article
Youle, R., Narendra, D. Mechanisms of mitophagy. Nat Rev Mol Cell Biol 12, 9–14 (2011). https://doi.org/10.1038/nrm3028
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrm3028
This article is cited by
-
Intracellular delivery of Parkin-RING0-based fragments corrects Parkin-induced mitochondrial dysfunction through interaction with SLP-2
Journal of Translational Medicine (2024)
-
Transcriptomics analysis revealed that TAZ regulates the proliferation of KIRC cells through mitophagy
BMC Cancer (2024)
-
DJ-1 protects cell death from a mitochondrial oxidative stress due to GBA1 deficiency
Genes & Genomics (2024)
-
Emerging roles of mitochondrial functions and epigenetic changes in the modulation of stem cell fate
Cellular and Molecular Life Sciences (2024)
-
Therapeutic Strategies Targeting Mitochondrial Dysfunction in Sepsis-induced Cardiomyopathy
Cardiovascular Drugs and Therapy (2024)