Key Points
-
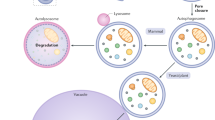
Eukaryotic cells have evolved a degradative pathway called autophagy that can deliver a large amount of cytoplasmic proteins and even whole organelles into lytic compartments, such as lysosomes in mammals or vacuoles in plant and yeast cells. Autophagy has vital roles in various physiological situations and is also involved in several pathological processes.
-
A number of physiological signals and stresses can induce autophagy. Following its induction, double membrane-bound vesicles called autophagosomes are newly formed in the cytoplasm to sequester materials (cargoes) to be degraded.
-
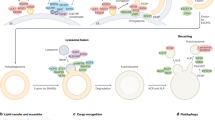
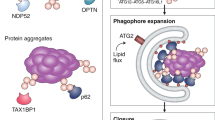
Two modes of cargo sequestration by autophagosomal membranes are suggested: non-selective (starvation-induced) autophagy, in which a portion of the cytoplasm is randomly engulfed, and selective autophagy, in which specific cargoes, such as toxic protein aggregates and superfluous or damaged organelles, are recognized and in many cases exclusively enwrapped by the membranes.
-
Studies in yeast have identified a unique subset of proteins called autophagy-related (Atg) proteins, which contain core components that are commonly required for membrane formation in all types of autophagy. These components constitute several subgroups, such as a protein kinase complex, a lipid kinase complex and two ubiquitin-like conjugation systems.
-
Atg proteins are concentrated at the site for membrane formation and organize a dynamic assembly called the preautophagosomal structure, in which Atg proteins specific for each type of autophagy (which differ in their induction signals and cargoes) are thought to serve as conductors that regulate the localization and activity of core machinery and determine the site and the mode of vesicle formation according to various situations.
Abstract
Autophagy is a fundamental function of eukaryotic cells and is well conserved from yeast to humans. The most remarkable feature of autophagy is the synthesis of double membrane-bound compartments that sequester materials to be degraded in lytic compartments, a process that seems to be mechanistically distinct from conventional membrane traffic. The discovery of autophagy in yeast and the genetic tractability of this organism have allowed us to identify genes that are responsible for this process, which has led to the explosive growth of this research field seen today. Analyses of autophagy-related (Atg) proteins have unveiled dynamic and diverse aspects of mechanisms that underlie membrane formation during autophagy.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Deter, R. L., Baudhuin, P. & De Duve, C. Participation of lysosomes in cellular autophagy induced in rat liver by glucagon. J. Cell Biol. 35, C11–C16 (1967).
Klionsky, D. J. Autophagy: from phenomenology to molecular understanding in less than a decade. Nature Rev. Mol. Cell Biol. 8, 931–937 (2007).
Mizushima, N. Autophagy: process and function. Genes Dev. 21, 2861–2873 (2007).
Mizushima, N., Levine, B., Cuervo, A. M. & Klionsky, D. J. Autophagy fights disease through cellular self-digestion. Nature 451, 1069–1075 (2008).
Takeshige, K., Baba, M., Tsuboi, S., Noda, T. & Ohsumi, Y. Autophagy in yeast demonstrated with proteinase-deficient mutants and conditions for its induction. J. Cell Biol. 119, 301–311 (1992). Reports the discovery of autophagy in yeast.
Baba, M., Takeshige, K., Baba, N. & Ohsumi, Y. Ultrastructural analysis of the autophagic process in yeast: detection of autophagosomes and their characterization. J. Cell Biol. 124, 903–913 (1994).
Baba, M., Osumi, M. & Ohsumi, Y. Analysis of the membrane structures involved in autophagy in yeast by freeze-replica method. Cell Struct. Funct. 20, 465–471 (1995). References 6 and 7 report the detailed morphological characterization of yeast autophagy.
Tsukada, M. & Ohsumi, Y. Isolation and characterization of autophagy-defective mutants of Saccharomyces cerevisiae. FEBS Lett. 333, 169–174 (1993). First report on autophagy-defective yeast mutants.
Baba, M., Osumi, M., Scott, S. V., Klionsky, D. J. & Ohsumi, Y. Two distinct pathways for targeting proteins from the cytoplasm to the vacuole/lysosome. J. Cell Biol. 139, 1687–1695 (1997).
Harding, T. M., Hefner-Gravink, A., Thumm, M. & Klionsky, D. J. Genetic and phenotypic overlap between autophagy and the cytoplasm to vacuole protein targeting pathway. J. Biol. Chem. 271, 17621–17624 (1996).
Thumm, M. et al. Isolation of autophagocytosis mutants of Saccharomyces cerevisiae. FEBS Lett. 349, 275–280 (1994).
Yuan, W., Tuttle, D. L., Shi, Y. J., Ralph, G. S. & Dunn, W. A. Jr. Glucose-induced microautophagy in Pichia pastoris requires the α-subunit of phosphofructokinase. J. Cell Sci. 110, 1935–1945 (1997).
Sakai, Y., Koller, A., Rangell, L. K., Keller, G. A. & Subramani, S. Peroxisome degradation by microautophagy in Pichia pastoris: identification of specific steps and morphological intermediates. J. Cell Biol. 141, 625–636 (1998).
Mukaiyama, H. et al. Paz2 and 13 other PAZ gene products regulate vacuolar engulfment of peroxisomes during micropexophagy. Genes Cells 7, 75–90 (2002).
Titorenko, V. I., Keizer, I., Harder, W. & Veenhuis, M. Isolation and characterization of mutants impaired in the selective degradation of peroxisomes in the yeast Hansenula polymorpha. J. Bacteriol. 177, 357–363 (1995).
Klionsky, D. J. et al. A unified nomenclature for yeast autophagy-related genes. Dev. Cell 5, 539–545 (2003).
Kamada, Y. et al. Tor-mediated induction of autophagy via an Apg1 protein kinase complex. J. Cell Biol. 150, 1507–1513 (2000). Shows dephosphorylation of Atg13 in response to nutrient starvation, which allows Atg13 to interact with Atg1, leading to stimulation of the kinase activity of Atg1.
Kihara, A., Noda, T., Ishihara, N. & Ohsumi, Y. Two distinct Vps34 phosphatidylinositol 3-kinase complexes function in autophagy and carboxypeptidase Y sorting in Saccharomyces cerevisiae. J. Cell Biol. 152, 519–530 (2001). Identifies two distinct PtdIns 3-kinase complexes, one of which contains Atg14 as a specific component and is required for autophagy.
Mizushima, N. et al. A protein conjugation system essential for autophagy. Nature 395, 395–398 (1998).
Ichimura, Y. et al. A ubiquitin-like system mediates protein lipidation. Nature 408, 488–492 (2000). References 19 and 20 report the discovery of two ubiquitin-like protein conjugates required for autophagy, Atg12–Atg5 and Atg82013 PE, respectively.
Shintani, T., Suzuki, K., Kamada, Y., Noda, T. & Ohsumi, Y. Apg2p functions in autophagosome formation on the perivacuolar structure. J. Biol. Chem. 276, 30452–30460 (2001).
Wang, C. W. et al. Apg2 is a novel protein required for the cytoplasm to vacuole targeting, autophagy, and pexophagy pathways. J. Biol. Chem. 276, 30442–30451 (2001).
Noda, T. et al. Apg9p/Cvt7p is an integral membrane protein required for transport vesicle formation in the Cvt and autophagy pathways. J. Cell Biol. 148, 465–480 (2000).
Barth, H., Meiling-Wesse, K., Epple, U. D. & Thumm, M. Autophagy and the cytoplasm to vacuole targeting pathway both require Aut10p. FEBS Lett. 508, 23–28 (2001).
Kirisako, T. et al. Formation process of autophagosome is traced with Apg8/Aut7p in yeast. J. Cell Biol. 147, 435–446 (1999). Reports the identification of Atg8 as a marker protein for the autophagosome and the isolation membrane.
Suzuki, K. et al. The pre-autophagosomal structure organized by concerted functions of APG genes is essential for autophagosome formation. EMBO J. 20, 5971–5981 (2001). Reports the identification of the PAS.
Suzuki, K., Kubota, Y., Sekito, T. & Ohsumi, Y. Hierarchy of Atg proteins in pre-autophagosomal structure organization. Genes Cells 12, 209–218 (2007). Reveals that Atg proteins organize the PAS in a hierarchical manner.
Cao, Y., Cheong, H., Song, H. & Klionsky, D. J. In vivo reconstitution of autophagy in Saccharomyces cerevisiae. J. Cell Biol. 182, 703–713 (2008).
Noda, T. & Ohsumi, Y. Tor, a phosphatidylinositol kinase homologue, controls autophagy in yeast. J. Biol. Chem. 273, 3963–3966 (1998).
Wullschleger, S., Loewith, R. & Hall, M. N. TOR signaling in growth and metabolism. Cell 124, 471–484 (2006).
Yorimitsu, T., Zaman, S., Broach, J. R. & Klionsky, D. J. Protein kinase A and Sch9 cooperatively regulate induction of autophagy in Saccharomyces cerevisiae. Mol. Biol. Cell 18, 4180–4189 (2007).
Budovskaya, Y. V., Stephan, J. S., Reggiori, F., Klionsky, D. J. & Herman, P. K. The Ras/cAMP-dependent protein kinase signaling pathway regulates an early step of the autophagy process in Saccharomyces cerevisiae. J. Biol. Chem. 279, 20663–20671 (2004).
Matsuura, A., Tsukada, M., Wada, Y. & Ohsumi, Y. Apg1p, a novel protein kinase required for the autophagic process in Saccharomyces cerevisiae. Gene 192, 245–250 (1997).
Funakoshi, T., Matsuura, A., Noda, T. & Ohsumi, Y. Analyses of APG13 gene involved in autophagy in yeast, Saccharomyces cerevisiae. Gene 192, 207–213 (1997).
Kabeya, Y. et al. Atg17 functions in cooperation with Atg1 and Atg13 in yeast autophagy. Mol. Biol. Cell 16, 2544–2553 (2005).
Kawamata, T. et al. Characterization of a novel autophagy-specific gene, ATG29. Biochem. Biophys. Res. Commun. 338, 1884–1889 (2005).
Kabeya, Y., Kawamata, T., Suzuki, K. & Ohsumi, Y. Cis1/Atg31 is required for autophagosome formation in Saccharomyces cerevisiae. Biochem. Biophys. Res. Commun. 356, 405–410 (2007).
Kawamata, T., Kamada, Y., Kabeya, Y., Sekito, T. & Ohsumi, Y. Organization of the pre-autophagosomal structure responsible for autophagosome formation. Mol. Biol. Cell 19, 2039–2050 (2008).
Cheong, H., Nair, U., Geng, J. & Klionsky, D. J. The Atg1 kinase complex is involved in the regulation of protein recruitment to initiate sequestering vesicle formation for nonspecific autophagy in Saccharomyces cerevisiae. Mol. Biol. Cell 19, 668–681 (2008). References 38 and 39 show that the Atg1 kinase and its regulators collaboratively function to organize, or reorganize, the PAS in response to nutrient starvation.
Cheong, H. et al. Atg17 regulates the magnitude of the autophagic response. Mol. Biol. Cell 16, 3438–3453 (2005).
Sekito, T., Kawamata, T., Ichikawa, R., Suzuki, K. & Ohsumi, Y. Atg17 recruits Atg9 to organize the pre-autophagosomal structure. Genes Cells 14, 525–538 (2009).
Klionsky, D. J. The molecular machinery of autophagy: unanswered questions. J. Cell Sci. 118, 7–18 (2005).
Hara, T. et al. FIP200, a ULK-interacting protein, is required for autophagosome formation in mammalian cells. J. Cell Biol. 181, 497–510 (2008).
Chan, E. Y., Longatti, A., McKnight, N. C. & Tooze, S. A. Kinase-inactivated ULK proteins inhibit autophagy via their conserved C-terminal domains using an Atg13-independent mechanism. Mol. Cell. Biol. 29, 157–171 (2009).
Hosokawa, N. et al. Nutrient-dependent mTORC1 association with the ULK1–Atg13–FIP200 complex required for autophagy. Mol. Biol. Cell 20, 1981–1991 (2009).
Chang, Y. Y. & Neufeld, T. P. An Atg1/Atg13 complex with multiple roles in TOR-mediated autophagy regulation. Mol. Biol. Cell 20, 2004–2014 (2009).
Jung . et al. ULK–Atg13–FIP200 complexes mediate mTOR signaling to the autophagy machinery. Mol. Biol. Cell 20, 1992–2003 (2009).
Ganley . et al. ULK1˙ATG13˙FIP200 complex mediates mTOR signaling and is essential for autophagy. J. Biol. Chem. 284, 12297–12305.
Petiot, A., Ogier-Denis, E., Blommaart, E. F., Meijer, A. J. & Codogno, P. Distinct classes of phosphatidylinositol 3′-kinases are involved in signaling pathways that control macroautophagy in HT-29 cells. J. Biol. Chem. 275, 992–998 (2000).
Obara, K., Sekito, T., Niimi, K. & Ohsumi, Y. The Atg18–Atg2 complex is recruited to autophagic membranes via phosphatidylinositol 3-phosphate and exerts an essential function. J. Biol. Chem. 283, 23972–23980 (2008).
Schu, P. V. et al. Phosphatidylinositol 3-kinase encoded by yeast VPS34 gene essential for protein sorting. Science 260, 88–91 (1993).
Obara, K., Sekito, T. & Ohsumi, Y. Assortment of phosphatidylinositol 3-kinase complexes—Atg14p directs association of complex I to the pre-autophagosomal structure in Saccharomyces cerevisiae. Mol. Biol. Cell 17, 1527–1539 (2006).
Dove, S. K. et al. Svp1p defines a family of phosphatidylinositol 3,5-bisphosphate effectors. EMBO J. 23, 1922–1933 (2004).
Stromhaug, P. E., Reggiori, F., Guan, J., Wang, C. W. & Klionsky, D. J. Atg21 is a phosphoinositide binding protein required for efficient lipidation and localization of Atg8 during uptake of aminopeptidase I by selective autophagy. Mol. Biol. Cell 15, 3553–3566 (2004).
Efe, J. A., Botelho, R. J. & Emr, S. D. Atg18 regulates organelle morphology and Fab1 kinase activity independent of its membrane recruitment by phosphatidylinositol 3,5-bisphosphate. Mol. Biol. Cell 18, 4232–4244 (2007).
Obara, K., Noda, T., Niimi, K. & Ohsumi, Y. Transport of phosphatidylinositol 3-phosphate into the vacuole via autophagic membranes in S. cerevisiae. Genes Cells, 13, 537–547 (2008).
Reggiori, F., Tucker, K. A., Stromhaug, P. E. & Klionsky, D. J. The Atg1–Atg13 complex regulates Atg9 and Atg23 retrieval transport from the pre-autophagosomal structure. Dev. Cell 6, 79–90 (2004).
Young, A. R. et al. Starvation and ULK1-dependent cycling of mammalian Atg9 between the TGN and endosomes. J. Cell Sci. 119, 3888–3900 (2006).
Xie, Z. & Klionsky, D. J. Autophagosome formation: core machinery and adaptations. Nature Cell Biol. 9, 1102–1109 (2007).
He, C., Baba, M., Cao, Y. & Klionsky, D. J. Self-interaction is critical for Atg9 transport and function at the phagophore assembly site during autophagy. Mol. Biol. Cell 19, 5506–5516 (2008).
Mizushima, N. et al. Dissection of autophagosome formation using Apg5-deficient mouse embryonic stem cells. J. Cell Biol. 152, 657–668 (2001). Demonstration of Atg5 localization on the isolation membrane and real-time visualization of the process of autophagosome formation in living cells using GFP-tagged Atg5.
Kirisako, T. et al. The reversible modification regulates the membrane-binding state of Apg8/Aut7 essential for autophagy and the cytoplasm to vacuole targeting pathway. J. Cell Biol. 151, 263–276 (2000).
Kim, J., Huang, W. P. & Klionsky, D. J. Membrane recruitment of Aut7p in the autophagy and cytoplasm to vacuole targeting pathways requires Aut1p, Aut2p, and the autophagy conjugation complex. J. Cell Biol. 152, 51–64 (2001).
Nakatogawa, H., Ichimura, Y. & Ohsumi, Y. Atg8, a ubiquitin-like protein required for autophagosome formation, mediates membrane tethering and hemifusion. Cell 130, 165–178 (2007). Shows that Atg8–PE causes aggregation and hemifusion of liposomes in vitro and that these phenomena represent the function of Atg8 in autophagosome formation in vivo.
Kabeya, Y. et al. LC3, a mammalian homologue of yeast Apg8p, is localized in autophagosome membranes after processing. EMBO J. 19, 5720–5728 (2000).
Yoshimoto, K. et al. Processing of ATG8s, ubiquitin-like proteins, and their deconjugation by ATG4s are essential for plant autophagy. Plant Cell 16, 2967–2983 (2004).
Klionsky, D. J. et al. Guidelines for the use and interpretation of assays for monitoring autophagy in higher eukaryotes. Autophagy 4, 151–175 (2008).
Ichimura, Y. et al. In vivo and in vitro reconstitution of Atg8 conjugation essential for autophagy. J. Biol. Chem. 279, 40584–40592 (2004).
Xie, Z., Nair, U. & Klionsky, D. J. Atg8 controls phagophore expansion during autophagosome formation. Mol. Biol. Cell 19, 3290–3298 (2008).
Fujita, N. et al. An Atg4B mutant hampers the lipidation of LC3 paralogues and causes defects in autophagosome closure. Mol. Biol. Cell 19, 4651–4659 (2008).
Sou, Y. S. et al. The Atg8 conjugation system is indispensable for proper development of autophagic isolation membranes in mice. Mol. Biol. Cell 19, 4762–4775 (2008).
Mizushima, N., Noda, T. & Ohsumi, Y. Apg16p is required for the function of the Apg12p–Apg5p conjugate in the yeast autophagy pathway. EMBO J. 18, 3888–3896 (1999).
Kuma, A., Mizushima, N., Ishihara, N. & Ohsumi, Y. Formation of the approximately 350-kDa Apg12–Apg5˙Apg16 multimeric complex, mediated by Apg16 oligomerization, is essential for autophagy in yeast. J. Biol. Chem. 277, 18619–18625 (2002).
Mizushima, N. et al. Mouse Apg16L, a novel WD-repeat protein, targets to the autophagic isolation membrane with the Apg12–Apg5 conjugate. J. Cell Sci. 116, 1679–1688 (2003).
Geng, J., Baba, M., Nair, U. & Klionsky, D. J. Quantitative analysis of autophagy-related protein stoichiometry by fluorescence microscopy. J. Cell Biol. 182, 129–140 (2008).
Hanada, T. & Ohsumi, Y. Structure–function relationship of Atg12, a ubiquitin-like modifier essential for autophagy. Autophagy 1, 110–118 (2005).
Hanada, T. et al. The Atg12–Atg5 conjugate has a novel E3-like activity for protein lipidation in autophagy. J. Biol. Chem. 282, 37298–37302 (2007). Shows that the ubiquitin-like protein conjugate Atg12–Atg5 directly stimulates the conjugation reaction of the other ubiquitin-like protein Atg8 to PE.
Fujioka, Y. et al. In vitro reconstitution of plant Atg8 and Atg12 conjugation systems essential for autophagy. J. Biol. Chem. 283, 1921–1928 (2008).
Sou, Y. S., Tanida, I., Komatsu, M., Ueno, T. & Kominami, E. Phosphatidylserine in addition to phosphatidylethanolamine is an in vitro target of the mammalian Atg8 modifiers, LC3, GABARAP, and GATE-16. J. Biol. Chem. 281, 3017–3024 (2006).
Oh-oka, K., Nakatogawa, H. & Ohsumi, Y. Physiological pH and acidic phospholipids contribute to substrate specificity in lipidation of Atg8. J. Biol. Chem. 283, 21847–21852 (2008).
Fujita, N. et al. The Atg16L complex specifies the site of LC3 lipidation for membrane biogenesis in autophagy. Mol. Biol. Cell 19, 2092–2100 (2008).
Kim, J., Scott, S. V., Oda, M. N. & Klionsky, D. J. Transport of a large oligomeric protein by the cytoplasm to vacuole protein targeting pathway. J. Cell Biol. 137, 609–618 (1997).
Scott, S. V., Guan, J., Hutchins, M. U., Kim, J. & Klionsky, D. J. Cvt19 is a receptor for the cytoplasm-to-vacuole targeting pathway. Mol. Cell 7, 1131–1141 (2001).
Shintani, T., Huang, W. P., Stromhaug, P. E. & Klionsky, D. J. Mechanism of cargo selection in the cytoplasm to vacuole targeting pathway. Dev. Cell 3, 825–837 (2002).
Chang, C. Y. & Huang, W. P. Atg19 mediates a dual interaction cargo sorting mechanism in selective autophagy. Mol. Biol. Cell 18, 919–929 (2007).
Noda, N. N. et al. Structural basis of target recognition by Atg8/LC3 during selective autophagy. Genes Cells 13, 1211–1218 (2008).
Bjorkoy, G. et al. p62/SQSTM1 forms protein aggregates degraded by autophagy and has a protective effect on huntingtin-induced cell death. J. Cell Biol. 171, 603–614 (2005).
Pankiv, S. et al. p62/SQSTM1 binds directly to Atg8/LC3 to facilitate degradation of ubiquitinated protein aggregates by autophagy. J. Biol. Chem. 282, 24131–24145 (2007).
Komatsu, M. et al. Homeostatic levels of p62 control cytoplasmic inclusion body formation in autophagy-deficient mice. Cell 131, 1149–1163 (2007).
Shvets, E., Fass, E., Scherz-Shouval, R. & Elazar, Z. The N-terminus and Phe52 residue of LC3 recruit p62/SQSTM1 into autophagosomes. J. Cell Sci. 121, 2685–2695 (2008).
Ichimura, Y. et al. Structural basis for sorting mechanism of p62 in selective autophagy. J. Biol. Chem. 283, 22847–22857 (2008). References 86–91 establish the molecular framework for p62-mediated degradation of disease-related protein inclusions through autophagy and its pathological significance.
Nice, D. C., Sato, T. K., Stromhaug, P. E., Emr, S. D. & Klionsky, D. J. Cooperative binding of the cytoplasm to vacuole targeting pathway proteins, Cvt13 and Cvt20, to phosphatidylinositol 3-phosphate at the pre-autophagosomal structure is required for selective autophagy. J. Biol. Chem. 277, 30198–30207 (2002).
Kim, J. et al. Cvt9/Gsa9 functions in sequestering selective cytosolic cargo destined for the vacuole. J. Cell Biol. 153, 381–396 (2001).
Shintani, T. & Klionsky, D. J. Cargo proteins facilitate the formation of transport vesicles in the cytoplasm to vacuole targeting pathway. J. Biol. Chem. 279, 29889–29894 (2004). Shows that PAS organization under nutrient-rich conditions, and thus Cvt vesicle formation, largely depends on the existence of the cargo molecule Ape1 in addition to the Cvt-specific proteins Atg11 and Atg19.
Yorimitsu, T. & Klionsky, D. J. Atg11 links cargo to the vesicle-forming machinery in the cytoplasm to vacuole targeting pathway. Mol. Biol. Cell 16, 1593–1605 (2005).
Farre, J. C., Manjithaya, R., Mathewson, R. D. & Subramani, S. PpAtg30 tags peroxisomes for turnover by selective autophagy. Dev. Cell 14, 365–376 (2008).
Kanki, T. & Klionsky, D. J. Mitophagy in yeast occurs through a selective mechanism. J. Biol. Chem. 283, 32386–32393 (2008).
Otto, G. P., Wu, M. Y., Kazgan, N., Anderson, O. R. & Kessin, R. H. Dictyostelium macroautophagy mutants vary in the severity of their developmental defects. J. Biol. Chem. 279, 15621–15629 (2004).
Fujiki, Y. et al. An Arabidopsis homolog of yeast ATG6/VPS30 is essential for pollen germination. Plant Physiol. 143, 1132–1139 (2007).
Ravikumar, B. et al. Inhibition of mTOR induces autophagy and reduces toxicity of polyglutamine expansions in fly and mouse models of Huntington disease. Nature Genet. 36, 585–595 (2004).
Kageyama, T., Suzuki, K. & Ohsumi, Y. Lap3 is a selective target of autophagy in yeast, Saccharomyces cerevisiae. Biochem. Biophys. Res. Commun. 378, 551–557 (2009).
Onodera, J. & Ohsumi, Y. Ald6p is a preferred target for autophagy in yeast, Saccharomyces cerevisiae. J. Biol. Chem. 279, 16071–16076 (2004).
Kraft, C., Deplazes, A., Sohrmann, M. & Peter, M. Mature ribosomes are selectively degraded upon starvation by an autophagy pathway requiring the Ubp3p/Bre5p ubiquitin protease. Nature Cell Biol. 10, 602–610 (2008). References 102 and 103 report that the cytoplasmic protein Ald6 and ribosomal proteins, respectively, are preferentially degraded in yeast by mechanisms that seem to be distinct from those previously suggested for other selective types of autophagy.
Author information
Authors and Affiliations
Corresponding author
Glossary
- Macroautophagy
-
The sequestration of cytosolic components in autophagosomes and their subsequent degradation when autophagosomes fuse with lysosomes.
- Autophagosome
-
A double membrane-bound vesicle that is formed during autophagy. The vesicle sequesters materials to be degraded and delivers them to the lysosome in mammals or the vacuole in yeasts and plants.
- Ubiquitin–26S proteasome system
-
The system that degrades selected proteins that are first marked with ubiquitin chains and then degraded by the multi-catalytic proteinase complex, the proteasome.
- Autophagic body
-
The inner membrane-bound structure of the autophagosome that is released into the vacuolar lumen by fusion of the autophagosomal outer membrane with the vacuolar membrane.
- Vacuolar protein sorting
-
A pathway that mediates the selective transport of a subset of proteins from the late Golgi compartment to the vacuole through the endosome.
- E1
-
An enzyme that activates ubiquitin and ubiquitin-like proteins using ATP and transfers them to E2 enzymes.
- E2
-
An enzyme that receives ubiquitin and ubiquitin-like proteins from E1 enzymes and conjugates them to target molecules.
- Deconjugation enzyme
-
An enzyme that cleaves the isopeptide bond (the amide bond between autophagyrelated protein 8 (Atg8) and phosphatidylethanolamine) formed in ubiquitin and ubiquitin-like protein conjugates.
- Hemifusion
-
Fusion between outer leaflets of membranes while inner leaflets remain intact. This is regarded as a common intermediate state in biological membrane fusion events.
- E3
-
An enzyme that stimulates the conjugation reaction by E2 enzymes and is also involved in the selection of target molecules.
Rights and permissions
About this article
Cite this article
Nakatogawa, H., Suzuki, K., Kamada, Y. et al. Dynamics and diversity in autophagy mechanisms: lessons from yeast. Nat Rev Mol Cell Biol 10, 458–467 (2009). https://doi.org/10.1038/nrm2708
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrm2708
This article is cited by
-
The role of autophagy in viral infections
Journal of Biomedical Science (2023)
-
Phase separation in fungi
Nature Microbiology (2023)
-
The plant unique ESCRT component FREE1 regulates autophagosome closure
Nature Communications (2023)
-
Deficiency of cancer/testis antigen gene CT55 causes male infertility in humans and mice
Cell Death & Differentiation (2023)
-
A non-canonical role of ATG8 in Golgi recovery from heat stress in plants
Nature Plants (2023)