Key Points
-
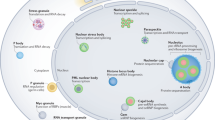
PML is a tumour suppressor protein that typically localizes to and is necessary for the formation of nuclear macromolecular structures called promyelocytic leukaemia nuclear bodies (PML-NBs).
-
PML-NBs are discrete nuclear foci of 0.2–1.0 μm in diameter that are present in most mammalian cell nuclei. They typically number 1–30 bodies per nucleus, depending on the cell type, cell-cycle phase and differentiation stage.
-
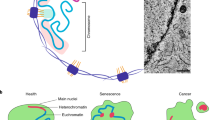
PML-NBs are dynamic structures that undergo significant changes in number, size and position during cell-cycle progression and in response to cellular stresses such as DNA damage and induction of senescence.
-
PML-NBs can be found near other nuclear organelles, such as Cajal bodies, and associate with genomic regions that are transcriptionally active. In addition, a specific association has been shown with specific chromosomal loci.
-
Many proteins transiently and constitutively localize to PML-NBs. As a result, PML-NBs have been implicated in the regulation of diverse cellular functions including induction of apoptosis and senescence, inhibition of proliferation, maintenance of genomic stability and antiviral responses.
-
Recent data suggest that PML-NBs are heterogeneous structures and that different PML-NBs may regulate specific cellular functions according to their protein composition, their position in the nucleus and their mobility.
Abstract
The promyelocytic leukaemia (PML) tumour suppressor protein epitomizes the PML-nuclear body (PML-NB) and is crucially required for the proper assembly of this macromolecular nuclear structure. Unlike other, more specialized subnuclear structures such as Cajal and Polycomb group bodies, PML-NBs are functionally promiscuous and have been implicated in the regulation of diverse cellular functions. PML-NBs are dynamic structures that favour the sequestration and release of proteins, mediate their post-translational modifications and promote specific nuclear events in response to various cellular stresses. Recent data suggest that PML-NBs may be heterogeneous in composition, mobility and function.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Melnick, A. & Licht, J. D. Deconstructing a disease: RARα, its fusion partners, and their roles in the pathogenesis of acute promyelocytic leukemia. Blood 93, 3167–3215 (1999). This review on the molecular pathogenesis of APL is a classic. Although some aspects are perhaps outdated, it still provides a broad and detailed description of the functions of the fusion partners of RARα.
Ishov, A. M. et al. PML is critical for ND10 formation and recruits the PML-interacting protein daxx to this nuclear structure when modified by SUMO-1. J. Cell Biol. 147, 221–234 (1999).
Salomoni, P. & Pandolfi, P. P. The role of PML in tumor suppression. Cell 108, 165–170 (2002).
Dellaire, G. & Bazett-Jones, D. P. PML nuclear bodies: dynamic sensors of DNA damage and cellular stress. Bioessays 26, 963–977 (2004). An important review on the structure of PML-NBs and their properties and dynamics in cells exposed to DNA damage.
Trotman, L. C. et al. Identification of a tumour suppressor network opposing nuclear Akt function. Nature 441, 523–527 (2006).
Bernardi, R. et al. PML inhibits HIF-1α translation and neoangiogenesis through repression of mTOR. Nature 442, 779–785 (2006).
Stuurman, N. et al. A monoclonal antibody recognizing nuclear matrix-associated nuclear bodies. J. Cell Sci. 101, 773–784 (1992).
Boisvert, F. M., Hendzel, M. J. & Bazett-Jones, D. P. Promyelocytic leukemia (PML) nuclear bodies are protein structures that do not accumulate RNA. J. Cell Biol. 148, 283–292 (2000).
Eskiw, C. H., Dellaire, G. & Bazett-Jones, D. P. Chromatin contributes to structural integrity of promyelocytic leukemia bodies through a SUMO-1-independent mechanism. J. Biol. Chem. 279, 9577–9585 (2004). The authors of this report used a combination of correlative fluorescence microscopy and electron spectroscopic imaging to demonstrate that the integrity of PML-NBs is maintained through contacts with surrounding chromatin fibres.
Ching, R. W., Dellaire, G., Eskiw, C. H. & Bazett-Jones, D. P. PML bodies: a meeting place for genomic loci? J. Cell Sci. 118, 847–854 (2005).
Borden, K. L. Pondering the promyelocytic leukemia protein (PML) puzzle: possible functions for PML nuclear bodies. Mol. Cell. Biol. 22, 5259–5269 (2002).
Grande, M. A. et al. PML-containing nuclear bodies: their spatial distribution in relation to other nuclear components. J. Cell. Biochem. 63, 280–291 (1996).
Wang, J. et al. Promyelocytic leukemia nuclear bodies associate with transcriptionally active genomic regions. J. Cell Biol. 164, 515–526 (2004).
Shiels, C. et al. PML bodies associate specifically with the MHC gene cluster in interphase nuclei. J. Cell Sci. 114, 3705–3716 (2001).
Kumar, P. P. et al. Functional interaction between PML and SATB1 regulates chromatin-loop architecture and transcription of the MHC class I locus. Nature Cell Biol. 9, 45–56 (2007).
Sun, Y., Durrin, L. K. & Krontiris, T. G. Specific interaction of PML bodies with the TP53 locus in Jurkat interphase nuclei. Genomics 82, 250–252 (2003).
Aoto, T., Saitoh, N., Ichimura, T., Niwa, H. & Nakao, M. Nuclear and chromatin reorganization in the MHC-Oct3/4 locus at developmental phases of embryonic stem cell differentiation. Dev. Biol. 298, 354–367 (2006).
Chang, S. K. et al. Proto-oncogene PML enhances antigen presentation by MHC class I molecules in human lung cancer cells. Mol. Cells 14, 130–135 (2002).
Bruno, S. et al. The PML gene is not involved in the regulation of MHC class I expression in human cell lines. Blood 101, 3514–3519 (2003).
Larghero, J. et al. Alteration of the PML proto-oncogene in leukemic cells does not abrogate expression of MHC class I antigens. Leukemia 13, 1295–1296 (1999).
Zhang, H. et al. Concordant down-regulation of proto-oncogene PML and major histocompatibility antigen HLA class I expression in high-grade prostate cancer. Cancer Immun. 3, 2 (2003).
Cabrera, C. M., Jimenez, P., Concha, A., Garrido, F. & Ruiz-Cabello, F. Promyelocytic leukemia (PML) nuclear bodies are disorganized in colorectal tumors with total loss of major histocompatibility complex class I expression and LMP7 downregulation. Tissue Antigens 63, 446–452 (2004).
Gurrieri, C. et al. Loss of the tumor suppressor PML in human cancers of multiple histologic origins. J. Natl Cancer Inst. 96, 269–279 (2004). A systematic analysis of PML expression in various tumours of different histological origin allowed the authors to affirm that PML and PML-NBs are lost in some tumours and that, in some cases, this correlates with disease progression.
Koken, M. H. et al. The PML growth-suppressor has an altered expression in human oncogenesis. Oncogene 10, 1315–1324 (1995).
Gambacorta, M. et al. Heterogeneous nuclear expression of the promyelocytic leukemia (PML) protein in normal and neoplastic human tissues. Am. J. Pathol. 149, 2023–2035 (1996).
Flenghi, L. et al. Characterization of a new monoclonal antibody (PG-M3) directed against the aminoterminal portion of the PML gene product: immunocytochemical evidence for high expression of PML proteins on activated macrophages, endothelial cells, and epithelia. Blood 85, 1871–1880 (1995).
Cho, Y., Lee, I., Maul, G. G. & Yu, E. A novel nuclear substructure, ND10: distribution in normal and neoplastic human tissues. Int. J. Mol. Med. 1, 717–724 (1998).
Terris, B. et al. PML nuclear bodies are general targets for inflammation and cell proliferation. Cancer Res. 55, 1590–1597 (1995).
Nisole, S., Stoye, J. P. & Saib, A. TRIM family proteins: retroviral restriction and antiviral defence. Nature Rev. Microbiol. 3, 799–808 (2005).
Everett, R. D. & Chelbi-Alix, M. K. PML and PML nuclear bodies: Implications in antiviral defence. Biochimie 89, 819–830 (2007).
Choi, Y. H., Bernardi, R., Pandolfi, P. P. & Benveniste, E. N. The promyelocytic leukemia protein functions as a negative regulator of IFN-γ signaling. Proc. Natl Acad. Sci. USA 103, 18715–18720 (2006).
Stadler, M. et al. Transcriptional induction of the PML growth suppressor gene by interferons is mediated through an ISRE and a GAS element. Oncogene 11, 2565–2573 (1995).
Dror, N. et al. Interferon regulatory factor-8 is indispensable for the expression of promyelocytic leukemia and the formation of nuclear bodies in myeloid cells. J. Biol. Chem. 282, 5633–5640 (2007). This paper shows that a loss of expression of IRF8 in human CML correlates with a loss of expression of PML. This is also the first report to demonstrate that loss of expression of PML can be detected in leukaemias in addition to solid tumours.
Jiang, W. Q. & Ringertz, N. Altered distribution of the promyelocytic leukemia-associated protein is associated with cellular senescence. Cell Growth Differ. 8, 513–522 (1997).
Pearson, M. et al. PML regulates p53 acetylation and premature senescence induced by oncogenic Ras. Nature 406, 207–210 (2000).
Ferbeyre, G. et al. PML is induced by oncogenic ras and promotes premature senescence. Genes Dev. 14, 2015–2027 (2000). References 35 and 36 are the first reports that implicate PML-NBs in the regulation of cellular senescence. They show that PML-NBs increase in size and number following the induction of senescence and that PML regulates p53 acetylation and activation.
de Stanchina, E. et al. PML is a direct p53 target that modulates p53 effector functions. Mol. Cell 13, 523–535 (2004).
Chang, K. S., Fan, Y. H., Andreeff, M., Liu, J. & Mu, Z. M. The PML gene encodes a phosphoprotein associated with the nuclear matrix. Blood 85, 3646–3653 (1995).
Hayakawa, F. & Privalsky, M. L. Phosphorylation of PML by mitogen-activated protein kinases plays a key role in arsenic trioxide-mediated apoptosis. Cancer Cell 5, 389–401 (2004).
Bernardi, R. et al. PML regulates p53 stability by sequestering Mdm2 to the nucleolus. Nature Cell Biol. 6, 665–672 (2004).
Yang, S., Kuo, C., Bisi, J. E. & Kim, M. K. PML-dependent apoptosis after DNA damage is regulated by the checkpoint kinase hCds1/Chk2. Nature Cell Biol. 4, 865–870 (2002).
Scaglioni, P. P. et al. A CK2-dependent mechanism for degradation of the PML tumor suppressor. Cell 126, 269–283 (2006).
Duprez, E. et al. SUMO-1 modification of the acute promyelocytic leukaemia protein PML: implications for nuclear localisation. J. Cell Sci. 112, 381–393 (1999).
Zhong, S. et al. Role of SUMO-1-modified PML in nuclear body formation. Blood 95, 2748–2752 (2000).
Best, J. L. et al. SUMO-1 protease-1 regulates gene transcription through PML. Mol. Cell 10, 843–855 (2002).
Nefkens, I. et al. Heat shock and Cd2+ exposure regulate PML and Daxx release from ND10 by independent mechanisms that modify the induction of heat-shock proteins 70 and 25 differently. J. Cell Sci. 116, 513–524 (2003).
Kamitani, T., Nguyen, H. P., Kito, K., Fukuda-Kamitani, T. & Yeh, E. T. Covalent modification of PML by the sentrin family of ubiquitin-like proteins. J. Biol. Chem. 273, 3117–3120 (1998).
Ayaydin, F. & Dasso, M. Distinct in vivo dynamics of vertebrate SUMO paralogues. Mol. Biol. Cell 15, 5208–5218 (2004).
Fu, C. et al. Stabilization of PML nuclear localization by conjugation and oligomerization of SUMO-3. Oncogene 24, 5401–5413 (2005).
Mukhopadhyay, D. et al. SUSP1 antagonizes formation of highly SUMO2/3-conjugated species. J. Cell Biol. 174, 939–949 (2006).
Nacerddine, K. et al. The SUMO pathway is essential for nuclear integrity and chromosome segregation in mice. Dev. Cell 9, 769–779 (2005). Describes the generation of mice that are deficient for the SUMO E2-conjugating enzyme UBC9. Loss of UBC9 leads to early embryonic lethality and major defects in chromosome condensation and segregation. Moreover, PML-NBs are disrupted along with other nuclear structures.
Shen, T. H., Lin, H. K., Scaglioni, P. P., Yung, T. M. & Pandolfi, P. P. The mechanisms of PML-nuclear body formation. Mol. Cell 24, 331–339 (2006). PML is shown to have a SUMO-binding domain that is necessary for PML-NB formation. A model for the nucleation of PML-NBs is presented.
Saitoh, N. et al. In situ SUMOylation analysis reveals a modulatory role of RanBP2 in the nuclear rim and PML bodies. Exp. Cell Res. 312, 1418–1430 (2006).
Quimby, B. B., Yong-Gonzalez, V., Anan, T., Strunnikov, A. V. & Dasso, M. The promyelocytic leukemia protein stimulates SUMO conjugation in yeast. Oncogene 25, 2999–3005 (2006). So far, this is the only report to show that the expression of PML in yeast can promote sumoylation activity in vivo and in vitro and that this activity depends on the RING domain of PML.
Seeler, J. S. & Dejean, A. Nuclear and unclear functions of SUMO. Nature Rev. Mol. Cell Biol. 4, 690–699 (2003).
Sternsdorf, T., Jensen, K. & Will, H. Evidence for covalent modification of the nuclear dot-associated proteins PML and Sp100 by PIC1/SUMO-1. J. Cell Biol. 139, 1621–1634 (1997).
Dellaire, G., Eskiw, C. H., Dehghani, H., Ching, R. W. & Bazett-Jones, D. P. Mitotic accumulations of PML protein contribute to the re-establishment of PML nuclear bodies in G1. J. Cell Sci. 119, 1034–1042 (2006).
Heun, P. SUMOrganization of the nucleus. Curr. Opin. Cell Biol. 19, 350–355 (2007).
Lin, D. Y. et al. Role of SUMO-interacting motif in Daxx SUMO modification, subnuclear localization, and repression of sumoylated transcription factors. Mol. Cell 24, 341–354 (2006).
Muratani, M. et al. Metabolic-energy-dependent movement of PML bodies within the mammalian cell nucleus. Nature Cell Biol. 4, 106–110 (2002).
Eskiw, C. H., Dellaire, G., Mymryk, J. & Bazett-Jones, D. P. Size, position and dynamic behavior of PML nuclear bodies following cell stress as a paradigm for supramolecular trafficking and assembly. J. Cell Sci. 116, 4455–4466 (2003).
Wiesmeijer, K., Molenaar, C., Bekeer, I. M., Tanke, H. J. & Dirks, R. W. Mobile foci of Sp100 do not contain PML: PML bodies are immobile but PML and Sp100 proteins are not. J. Struct. Biol. 140, 180–188 (2002).
Lallemand-Breitenbach, V. et al. Role of promyelocytic leukemia (PML) sumolation in nuclear body formation, 11S proteasome recruitment, and As2O3-induced PML or PML/retinoic acid receptor a degradation. J. Exp. Med. 193, 1361–1371 (2001).
Anton, L. C. et al. Intracellular localization of proteasomal degradation of a viral antigen. J. Cell Biol. 146, 113–124 (1999).
Dino Rockel, T. & von Mikecz, A. Proteasome-dependent processing of nuclear proteins is correlated with their subnuclear localization. J. Struct. Biol. 140, 189–199 (2002).
Tsukamoto, T. et al. Visualization of gene activity in living cells. Nature Cell Biol. 2, 871–878 (2000).
Dellaire, G., Ching, R. W., Dehghani, H., Ren, Y. & Bazett-Jones, D. P. The number of PML nuclear bodies increases in early S phase by a fission mechanism. J. Cell Sci. 119, 1026–1033 (2006).
Zhong, S. et al. A role for PML and the nuclear body in genomic stability. Oncogene 18, 7941–7947 (1999).
Wang, X. W. et al. Functional interaction of p53 and BLM DNA helicase in apoptosis. J. Biol. Chem. 276, 32948–32955 (2001).
Bischof, O. et al. Regulation and localization of the Bloom syndrome protein in response to DNA damage. J. Cell Biol. 153, 367–380 (2001).
Everett, R. D., Lomonte, P., Sternsdorf, T., van Driel, R. & Orr, A. Cell cycle regulation of PML modification and ND10 composition. J. Cell Sci. 112, 4581–4588 (1999).
Maul, G. G., Yu, E., Ishov, A. M. & Epstein, A. L. Nuclear domain 10 (ND10) associated proteins are also present in nuclear bodies and redistribute to hundreds of nuclear sites after stress. J. Cell. Biochem. 59, 498–513 (1995).
Salomoni, P. et al. The promyelocytic leukemia protein PML regulates c-Jun function in response to DNA damage. Blood 105, 3686–3690 (2005).
Seker, H. et al. UV-C-induced DNA damage leads to p53-dependent nuclear trafficking of PML. Oncogene 22, 1620–1628 (2003).
Conlan, L. A., McNees, C. J. & Heierhorst, J. Proteasome-dependent dispersal of PML nuclear bodies in response to alkylating DNA damage. Oncogene 23, 307–310 (2004).
Carbone, R., Pearson, M., Minucci, S. & Pelicci, P. G. PML NBs associate with the hMre11 complex and p53 at sites of irradiation induced DNA damage. Oncogene 21, 1633–1640 (2002).
Dellaire, G. et al. Promyelocytic leukemia nuclear bodies behave as DNA damage sensors whose response to DNA double-strand breaks is regulated by NBS1 and the kinases ATM, Chk2, and ATR. J. Cell Biol. 175, 55–66 (2006).
Janderova-Rossmeislova, L. et al. PML protein association with specific nucleolar structures differs in normal, tumor and senescent human cells. J. Struct. Biol. 159, 56–70 (2007).
Kurki, S., Latonen, L. & Laiho, M. Cellular stress and DNA damage invoke temporally distinct Mdm2, p53 and PML complexes and damage-specific nuclear relocalization. J. Cell Sci. 116, 3917–3925 (2003).
Dellaire, G., Farrall, R. & Bickmore, W. A. The Nuclear Protein Database (NPD): sub-nuclear localisation and functional annotation of the nuclear proteome. Nucleic Acids Res. 31, 328–330 (2003).
Zhong, S., Salomoni, P. & Pandolfi, P. P. The transcriptional role of PML and the nuclear body. Nature Cell Biol. 2, E85–E90 (2000).
Condemine, W. et al. Characterization of endogenous human promyelocytic leukemia isoforms. Cancer Res. 66, 6192–6198 (2006). The first report in which the generation of specific antibodies against different isoforms of PML is described. All PML isoforms are shown to be expressed in vivo.
Lin, H. K., Bergmann, S. & Pandolfi, P. P. Cytoplasmic PML function in TGF-β signalling. Nature 431, 205–211 (2004).
Tashiro, S. et al. Repression of PML nuclear body-associated transcription by oxidative stress-activated Bach2. Mol. Cell. Biol. 24, 3473–3484 (2004).
Kiesslich, A., von Mikecz, A. & Hemmerich, P. Cell cycle-dependent association of PML bodies with sites of active transcription in nuclei of mammalian cells. J. Struct. Biol. 140, 167–179 (2002).
Seeler, J. S., Marchio, A., Sitterlin, D., Transy, C. & Dejean, A. Interaction of SP100 with HP1 proteins: a link between the promyelocytic leukemia-associated nuclear bodies and the chromatin compartment. Proc. Natl Acad. Sci. USA 95, 7316–7321 (1998).
Everett, R. D. et al. A dynamic connection between centromeres and ND10 proteins. J. Cell Sci. 112, 3443–3454 (1999).
Luciani, J. J. et al. PML nuclear bodies are highly organised DNA-protein structures with a function in heterochromatin remodelling at the G2 phase. J. Cell Sci. 119, 2518–2531 (2006).
Zhang, R. et al. Formation of macroH2A-containing senescence-associated heterochromatin foci and senescence driven by ASF1a and HIRA. Dev. Cell 8, 19–30 (2005).
Ye, X. et al. Definition of pRB- and p53-dependent and independent steps in HIRA/ASF1a-mediated formation of senecence-associated heterochromatin foci (SAHF). Mol. Cell. Biol. 27, 2452–2465 (2007).
Bischof, O. et al. Deconstructing PML-induced premature senescence. EMBO J. 21, 3358–3369 (2002).
Beech, S. J., Lethbridge, K. J., Killick, N., McGlincy, N. & Leppard, K. N. Isoforms of the promyelocytic leukemia protein differ in their effects on ND10 organization. Exp. Cell Res. 307, 109–117 (2005).
Boe, S. O. et al. Promyelocytic leukemia nuclear bodies are predetermined processing sites for damaged DNA. J. Cell Sci. 119, 3284–3295 (2006).
Yeager, T. R. et al. Telomerase-negative immortalized human cells contain a novel type of promyelocytic leukemia (PML) body. Cancer Res. 59, 4175–4179 (1999).
Dunham, M. A., Neumann, A. A., Fasching, C. L. & Reddel, R. R. Telomere maintenance by recombination in human cells. Nature Genet. 26, 447–450 (2000).
Xu, Z. X., Timanova-Atanasova, A., Zhao, R. X. & Chang, K. S. PML colocalizes with and stabilizes the DNA damage response protein TopBP1. Mol. Cell. Biol. 23, 4247–4256 (2003).
Bernardi, R. & Pandolfi, P. P. Role of PML and the PML-nuclear body in the control of programmed cell death. Oncogene 22, 9048–9057 (2003).
Muntoni, A. & Reddel, R. R. The first molecular details of ALT in human tumor cells. Hum. Mol. Genet. 14, R191–R196 (2005).
Takahashi, Y., Lallemand-Breitenbach, V., Zhu, J. & de The, H. PML nuclear bodies and apoptosis. Oncogene 23, 2819–2824 (2004).
Yang, S. et al. Promyelocytic leukemia activates Chk2 by mediating Chk2 autophosphorylation. J. Biol. Chem. 281, 26645–26654 (2006).
Salomoni, P. & Khelifi, A. F. Daxx: death or survival protein? Trends Cell Biol. 16, 97–104 (2006).
Torii, S., Egan, D. A., Evans, R. A. & Reed, J. C. Human Daxx regulates Fas-induced apoptosis from nuclear PML oncogenic domains (PODs). EMBO J. 18, 6037–6049 (1999).
Zhong, S. et al. Promyelocytic leukemia protein (PML) and Daxx participate in a novel nuclear pathway for apoptosis. J. Exp. Med. 191, 631–640 (2000).
Chen, L. Y. & Chen, J. D. Daxx silencing sensitizes cells to multiple apoptotic pathways. Mol. Cell. Biol. 23, 7108–7121 (2003).
Croxton, R. et al. Daxx represses expression of a subset of antiapoptotic genes regulated by nuclear factor-kB. Cancer Res. 66, 9026–9035 (2006).
Meinecke, I. et al. Modification of nuclear PML protein by SUMO-1 regulates Fas-induced apoptosis in rheumatoid arthritis synovial fibroblasts. Proc. Natl Acad. Sci. USA 104, 5073–5078 (2007).
Milovic-Holm, K., Krieghoff, E., Jensen, K., Will, H. & Hofmann, T. G. FLASH links the CD95 signaling pathway to the cell nucleus and nuclear bodies. EMBO J. 26, 391–401 (2007).
Ferbeyre, G. PML a target of translocations in APL is a regulator of cellular senescence. Leukemia 16, 1918–1926 (2002).
Narita, M. et al. Rb-mediated heterochromatin formation and silencing of E2F target genes during cellular senescence. Cell 113, 703–716 (2003).
Bardos, J. I. & Ashcroft, M. Hypoxia-inducible factor-1 and oncogenic signalling. Bioessays 26, 262–269 (2004).
Wang, Z. G. et al. PML is essential for multiple apoptotic pathways. Nature Genet. 20, 266–272 (1998).
Jensen, K., Shiels, C. & Freemont, P. S. PML protein isoforms and the RBCC/TRIM motif. Oncogene 20, 7223–7233 (2001).
Yoshida, H. et al. PML-RARA inhibits PML IV enhancement of PU.1-induced C/EBPɛ expression in myeloid differentiation. Mol. Cell. Biol. 27, 5819–5834 (2007).
Fogal, V. et al. Regulation of p53 activity in nuclear bodies by a specific PML isoform. EMBO J. 19, 6185–6195 (2000).
Buschbeck, M. et al. PML4 induces differentiation by Myc destabilization. Oncogene 26, 3415–3422 (2006).
Nguyen, L. A. et al. Physical and functional link of the leukemia-associated factors AML1 and PML. Blood 105, 292–300 (2005).
Xu, Z. X., Zou, W. X., Lin, P. & Chang, K. S. A role for PML3 in centrosome duplication and genome stability. Mol. Cell 17, 721–732 (2005).
Jiang, W. Q., Zhong, Z. H., Henson, J. D. & Reddel, R. R. Identification of candidate alternative lengthening of telomeres genes by methionine restriction and RNA interference. Oncogene 26, 4635–4637 (2007).
Fasching, C. L., Bower, K. & Reddel, R. R. Telomerase-independent telomere length maintenance in the absence of alternative lengthening of telomeres-associated promyelocytic leukemia bodies. Cancer Res. 65, 2722–2729 (2005).
Henson, J. D. et al. A robust assay for alternative lengthening of telomeres in tumors shows the significance of alternative lengthening of telomeres in sarcomas and astrocytomas. Clin. Cancer Res. 11, 217–225 (2005).
Costa, A. et al. Telomere maintenance mechanisms in liposarcomas: association with histologic subtypes and disease progression. Cancer Res. 66, 8918–8924 (2006).
Potts, P. R. & Yu, H. The SMC5/6 complex maintains telomere length in ALT cancer cells through SUMOylation of telomere-binding proteins. Nature Struct. Mol. Biol. 14, 581–590 (2007).
Ishov, A. M., Vladimirova, O. V. & Maul, G. G. Heterochromatin and ND10 are cell-cycle regulated and phosphorylation-dependent alternate nuclear sites of the transcription repressor Daxx and SWI/SNF protein ATRX. J. Cell Sci. 117, 3807–3820 (2004).
Acknowledgements
We are grateful to L. DiSantis for critical reading of the manuscript and all members of the Pandolfi laboratory for insightful comments and discussion. This work was supported by the National Institutes of Health.
Author information
Authors and Affiliations
Corresponding author
Related links
Glossary
- Nuclear matrix
-
A network of nuclear proteins that provides a structural framework for organizing chromatin.
- Cajal body
-
A spherical organelle found in the nucleus of proliferative or transcriptionally active cells. It is involved in the biogenesis and trafficking of RNA–protein complexes that regulate RNA processing, and is named after Santiago Ramon y Cajal who described the organelle in 1903.
- Splicing speckle
-
A storage site of mRNA splicing factors.
- Immunofluorescence in situ hybridization
-
A technique that combines immunocytochemistry with fluorescence in situ hybridization to visualize the association between proteins and DNA.
- Major histocompatibility complex
-
(MHC). A family of cellular antigens that allow the immune system to distinguish self from non-self, encoded by a large genomic region in higher vertebrates.
- Acute promyelocytic leukaemia
-
(APL). A subtype of acute myeloid leukaemia that is characterized by the accumulation of immature granulocytes called promyelocytes.
- Interferon
-
A soluble glycoprotein (cytokine) that is produced by cells of the immune system and regulates antiviral and immunological responses by inducing the expression of a series of genes.
- Nuclear localization signal
-
(NLS). A short stretch of positively charged amino acids that mediates the transport of proteins into the nucleus.
- B-box
-
A zinc-binding protein domain defined by a series of conserved Cys and His residues.
- Coiled-coil domain
-
A protein structural domain that mediates subunit oligomerization. Coiled coils contain 2–5 α-helices that twist around each other to form a supercoil.
- Arsenic trioxide
-
A chemotherapeutic compound that is used to treat patients with acute promyelocytic leukaemia.
- E3 ubiquitin ligase
-
The third enzyme in a series — the first two are designated E1 and E2 — that is responsible for ubiquitylation of target proteins. E3 enzymes provide platforms for binding E2 enzymes and specific substrates, thereby coordinating ubiquitylation of the selected substrates.
- Sumoylation
-
An evolutionarily conserved post-translational modification whereby the small ubiquitin-like modifier (SUMO) protein becomes covalently conjugated to a subset of proteins by the concerted action of an E1 activating enzyme, an E2 conjugating enzyme and an E3 ligase.
- SUMO protease
-
An enzyme that catalyses the removal of SUMO moieties from proteins by cleaving isopeptide bonds.
- RING domain
-
A Cys-rich tandem zinc-finger domain of 40–60 amino acids that is often found in E3 ubiquitin ligases.
- SUMO E3 ligase
-
A protein that enhances the conjugation of SUMO to target proteins by the SUMO-conjugating UBC9 enzyme.
- Time-lapse microscopy
-
Microscopy in which the same cell is photographed at regular time intervals over several hours.
- Fluorescence recovery after photobleaching
-
(FRAP). A technique that measures the rate of recovery of fluorescence owing to the movement of a fluorescent marker into an area of the cell after that area has been rendered non-fluorescent by photobleaching.
- Centromeric region
-
The part of a chromosome that is attached to the spindle during nuclear division.
- Telomere
-
A specialized structure at the ends of a chromosome that contains repetitive DNA sequences. Telomere length declines with each cell division.
Rights and permissions
About this article
Cite this article
Bernardi, R., Pandolfi, P. Structure, dynamics and functions of promyelocytic leukaemia nuclear bodies. Nat Rev Mol Cell Biol 8, 1006–1016 (2007). https://doi.org/10.1038/nrm2277
Issue Date:
DOI: https://doi.org/10.1038/nrm2277
This article is cited by
-
HIRA vs. DAXX: the two axes shaping the histone H3.3 landscape
Experimental & Molecular Medicine (2024)
-
The immediate-early protein 1 of human herpesvirus 6B interacts with NBS1 and inhibits ATM signaling
EMBO Reports (2024)
-
The interactions between PML nuclear bodies and small and medium size DNA viruses
Virology Journal (2023)
-
PcTrim prevents early infection with white spot syndrome virus by inhibiting AP1-induced endocytosis
Cell Communication and Signaling (2023)
-
Loss of PML nuclear bodies in familial amyotrophic lateral sclerosis-frontotemporal dementia
Cell Death Discovery (2023)