Key Points
-
Proteases function as molecular switches in signalling circuits on the cell surface and in the extracellular milieu. In light of the many proteases that are encoded by the genome, and the even larger number of bioactive substrates, it is crucial to identify which substrates individual enzymes cleave and which proteases cleave a particular substrate.
-
Four general approaches are discussed that are commonly used to link proteases and relevant substrates: biochemistry, cell biology, proteomics and animal models (with a focus on mouse models).
-
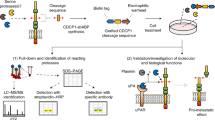
Biochemical studies use purified enzymes and substrates, and provide valuable information on which peptide and proteins a protease can cleave, the substrate's cleavage sites, and the inhibitor profile for small molecules and naturally occurring protease inhibitors.
-
Cell-biological assays help determine the function of enzymes in the context of an intact cell. Gain- and loss-of-function experiments can link enzymes and substrates and can help build hypotheses about enzyme function. Regulation of enzymes by activators and inhibitors of signal transduction as well as by transcriptional activation can be evaluated.
-
Degradomics studies of proteolysis use liquid chromatography or gel-based approaches for the mass spectrometric analysis of proteolysis. Degradomics enables the identification of hundreds or thousands of proteins in complex proteomes that have been moulded by proteolysis. Through isotope tagging, changes in the abundance levels of multiple peptides of a sample enables the identification of cleaved native substrates in cell-based systems to define the substrate degradome of a protease.
-
Mouse models allow an analysis of a protease's function in the context of an intact organism and help establish its expression pattern and relevance in development, adult homeostasis and disease models. Loss-of-function models help evaluate the contribution of enzymes to development and disease in vivo, whereas gain-of-function models yield insights into the consequences of dysregulated enzyme activity.
-
The combined application of these different approaches provides insights that exceed the sum of the individual approaches, and help resolve questions that arise from individual approaches. For example, they can help resolve the question: which of several candidate enzymes are relevant for processing a substrate in different cells and tissues or under different circumstances during development and in disease?
Abstract
Proteases function as molecular switches in signalling circuits at the cell surface and in the extracellular milieu. In light of the many proteases that are encoded by the genome, and the even larger number of bioactive substrates, it is crucial to identify which proteases cleave a particular substrate and which substrates individual proteases cleave. Elucidating the substrate degradomes of proteases will help us to understand the function of proteases in development and disease and to validate proteases as drug targets.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Macfarlane, R. G. An enzyme cascade in the blood clotting mechanism, and its function as a biochemical amplifier. Nature 202, 498–499 (1964).
Davie, E. W. & Ratnoff, O. D. Waterfall sequence for intrinsic blood clotting. Science 145, 1310–1312 (1964).
Davie, E. W. & Neurath, H. Identification of a peptide released during autocatalytic activation of trypsinogen. J. Biol. Chem. 212, 515–529 (1955).
Black, R. et al. A metalloprotease disintegrin that releases tumour-necrosis factor-α from cells. Nature 385, 729–733 (1997).
Moss, M. L. et al. Cloning of a disintegrin metalloproteinase that processes precursor tumour-necrosis factor-α. Nature 385, 733–736 (1997).
Peschon, J. J. et al. An essential role for ectodomain shedding in mammalian development. Science 282, 1281–1284 (1998). An excellent example of linking an enzyme, ADAM17, to several substrates, including TGFα, through the analysis of Adam17 -knockout mice and cell-based assays in cells lacking ADAM17, derived from these mice.
McQuibban, G. A. et al. Inflammation dampened by gelatinase A cleavage of monocyte chemoattractant protein-3. Science 289, 1202–1206 (2000).
Parks, W. C., Wilson, C. L. & López-Boado. Matrix metalloproteinases as modulators of inflammation and innate immunity. Nature Rev. Immunol. 4, 617–629 (2004).
Hintermann, E. & Quaranta, V. Epithelial cell motility on laminin-5: regulation by matrix assembly, proteolysis, integrins and erbB receptors. Matrix Biol. 23, 75–85 (2004).
O'Reilly, M. S., Holmgren, L., Chen, C. & Folkman, J. Angiostatin induces and sustains dormancy of human primary tumors in mice. Nature Med. 2, 689–692 (1996).
Bergers, G., Javaherian, K., Lo, K. M., Folkman, J. & Hanahan, D. Effects of angiogenesis inhibitors on multistage carcinogenesis in mice. Science 284, 808–812 (1999).
Overall, C. M. Dilating the degradome: matrix metalloproteinase-2 cuts to the heart of the matter. Biochem. J. 383, e5–e7 (2004).
Overall, C. M. & Kleifeld, O. Validating MMPs as drug targets and anti-targets for cancer therapy. Nature Rev. Cancer 6, 227–239 (2006).
Turk, B. Targeting proteases: successes, failures and future prospects. Nature Rev. Drug Discov. 5, 785–799 (2006).
López-Otin, C. & Overall, C. M. Protease degradomics: a new challenge for proteomics. Nature Rev. Mol. Cell Biol. 3, 509–519 (2002).
Rosendahl, M. S. et al. Identification and characterization of a pro-tumor necrosis factor-α-processing enzyme from the ADAM family of zinc metalloproteases. J. Biol. Chem. 272, 24588–24593 (1997).
Zheng, Y., Saftig, P., Hartmann, D. & Blobel, C. Evaluation of the contribution of different ADAMs to TNFα shedding and of the function of the TNFα ectodomain in ensuring selective stimulated shedding by the TNFα convertase (TACE/ADAM17). J. Biol. Chem. 279, 42898–42906 (2004).
Haro, H. et al. Matrix metalloproteinase-7-dependent release of tumor necrosis factor-α in a model of herniated disc resorption. J. Clin. Invest. 105, 143–150 (2000).
Tam, E. M., Morrison, C. J., Wu, Y. I., Stack, M. S. & Overall, C. M. Membrane protease proteomics: isotope-coded affinity tag MS identification of undescribed MT1-matrix metalloproteinase substrates. Proc. Natl Acad. Sci. USA 101, 6917–6922 (2004). A key paper describing the use of isotope mass tags and liquid-chromatography-based mass-spectrometric identification of cleaved products of native substrates in the cellular context.
Zhang, K. et al. Metalloproteinase cleavage of the chemokine SDF-1α induces neuronal apoptosis in HIV encephalitis. Nature Neurosci. 6, 1064–1071 (2003).
Shilling, O. & Overall, C. M. Proteomic discovery of protease substrates. Curr. Opin. Chem. Biol. 11, 1–10 (2007).
Roghani, M. et al. Metalloprotease-disintegrin MDC9: intracellular maturation and catalytic activity. J. Biol. Chem. 274, 3531–3540 (1999).
Loechel, F., Gilpin, B. J., Engvall, E., Albrechtsen, R. & Wewer, U. M. Human ADAM 12 (meltrin α) is an active metalloprotease. J. Biol. Chem. 273, 16993–16997 (1998).
Zou, J. et al. Catalytic activity of human ADAM33. J. Biol. Chem. 279, 9818–9830 (2004).
Smith, M. M., Shi, L. & Navre, M. Rapid identification of highly active and selective substrates for stromelysin and matrilysin using bacteriophage peptide display libraries. J. Biol. Chem. 270, 6440–6449 (1995).
Matthews, D. J. & Wells, J. A. Substrate phage: selection of protease substrates by monovalent phage display. Science 260, 1113–1117 (1993). A classic paper on the development of phage display to screen for protease-cleavage sites.
Rosse, G. et al. Rapid identification of substrates for novel proteases using a combinatorial peptide library. J. Comb. Chem. 2, 461–466 (2000).
Turk, B. E. & Cantley, L. C. Using peptide libraries to identify optimal cleavage motifs for proteolytic enzymes. Methods 32, 398–405 (2004).
Cunningham, B. C., Henner, D. J. & Wells, J. A. Engineering human prolactin to bind to the human growth hormone receptor. Science 247, 1461–1465 (1990).
Zhu, L. et al. The role of dipeptidyl peptidase IV in the cleavage of glucagon family peptides: in vivo metabolism of pituitary adenylate cyclase activating polypeptide-(1–38). J. Biol. Chem. 278, 22418–22423 (2003).
Thornberry, N. A. et al. A combinatorial approach defines specificities of members of the caspase family and granzyme B. Functional relationships established for key mediators of apoptosis. J. Biol. Chem. 272, 17907–17911 (1997). Describes the development of a now widely used technique for characterizing the active sites of proteases and for defining cleavage-site specificities.
Salisbury, C. M., Maly, D. J. & Ellman, J. A. Peptide microarrays for the determination of protease substrate specificity. J. Am. Chem. Soc. 124, 14868–14870 (2002).
Gao, X. et al. High density peptide microarrays. In situ synthesis and applications. Mol. Divers. 8, 177–187 (2004).
Marnett, A. B., Nomura, A. M., Shimba, N., Ortiz de Montellano, P. R. & Craik, C. S. Communication between the active sites and dimer interface of a herpesvirus protease revealed by a transition-state inhibitor. Proc. Natl Acad. Sci. USA 101, 6870–6875 (2004).
Greenbaum, D. C. et al. Small molecule affinity fingerprinting. A tool for enzyme family subclassification, target identification, and inhibitor design. Chem. Biol. 9, 1085–1094 (2002). A comprehensive analysis of the development and use of activity-based probes that launched recent in vivo imaging studies, inhibitor design and active-site characterization.
Barrios, A. M. & Craik, C. S. Scanning the prime-site substrate specificity of proteolytic enzymes: a novel assay based on ligand-enhanced lanthanide ion fluorescence. Bioorg. Med. Chem. Lett. 12, 3619–3623 (2002).
Turk, B. E., Huang, L. L., Piro, E. T. & Cantley, L. C. Determination of protease cleavage site motifs using mixture-based oriented peptide libraries. Nature Biotechnol. 19, 661–667 (2001).
Becherer, J. D. & Blobel, C. P. Biochemical properties and functions of membrane-anchored metalloprotease-disintegrin proteins (ADAMs). Curr. Top. Dev. Biol. 54, 101–123 (2003).
Gomis-Ruth, F. X. Structural aspects of the metzincin clan of metalloendopeptidases. Mol. Biotechnol. 24, 157–202 (2003).
Overall, C. M. Molecular determinants of metalloproteinase substrate specificity: matrix metalloproteinase substrate binding domains, modules, and exosites. Mol. Biotechnol. 22, 51–86 (2002).
Tortorella, M. et al. The thrombospondin motif of aggrecanase-1 (ADAMTS-4) is critical for aggrecan substrate recognition and cleavage. J. Biol. Chem. 275, 25791–25797 (2000).
Fields, G. B. A model for interstitial collagen catabolism by mammalian collagenases. J. Theor. Biol. 153, 585–602 (1991).
Boyd, S. E., Pike, R. N., Rudy, G. B., Whisstock, J. C. & Garcia de la Banda, M. PoPS: a computational tool for modeling and predicting protease specificity. J. Bioinform. Comput. Biol. 3, 551–585 (2005).
Berman, J. et al. Rapid optimization of enzyme substrates using defined substrate mixtures. J. Biol. Chem. 267, 1434–1437 (1992).
Overall, C. M., McQuibban, G. A. & Clark-Lewis, I. Discovery of chemokine substrates for matrix metalloproteinases by exosite scanning: a new tool for degradomics. Biol. Chem. 383, 1059–1066 (2002).
Boldt, H. B. et al. The Lin12-notch repeats of pregnancy-associated plasma protein-A bind calcium and determine its proteolytic specificity. J. Biol. Chem. 279, 38525–38531 (2004).
Torres-Collado, A. X., Kisiel, W., Iruela-Arispe, M. L. & Rodriguez-Manzaneque, J. C. ADAMTS1 interacts with, cleaves, and modifies the extracellular location of the matrix inhibitor tissue factor pathway inhibitor-2. J. Biol. Chem. 281, 17827–17837 (2006).
Overall, C. M. et al. Protease degradomics: mass spectrometry discovery of protease substrates and the CLIP-CHIP, a dedicated DNA microarray of all human proteases and inhibitors. Biol. Chem. 385, 493–504 (2004).
Wild-Bode, C., Fellerer, K., Kugler, J., Haass, C. & Capell, A. A basolateral sorting signal directs ADAM10 to adherens junctions and is required for its function in cell migration. J. Biol. Chem. 281, 23824–23829 (2006).
Loechel, F., Overgaard, M. T., Oxvig, C., Albrechtsen, R. & Wewer, U. M. Regulation of human ADAM 12 protease by the prodomain. Evidence for a functional cysteine switch. J. Biol. Chem. 274, 13427–13433 (1999).
Pei, D. & Weiss, S. J. Furin-dependent intracellular activation of the human stromelysin-3 zymogen. Nature 375, 244–247 (1995).
Tchougounova, E. et al. A key role for mast cell chymase in the activation of pro-matrix metalloprotease-9 and pro-matrix metalloprotease-2. J. Biol. Chem. 280, 9291–9296 (2005).
Lundequist, A., Tchougounova, E., Abrink, M. & Pejler, G. Cooperation between mast cell carboxypeptidase A and the chymase mouse mast cell protease 4 in the formation and degradation of angiotensin II. J. Biol. Chem. 279, 32339–32344 (2004).
Sahin, U. et al. Distinct roles for ADAM10 and ADAM17 in ectodomain shedding of six EGFR-ligands. J. Cell Biol. 164, 769–779 (2004). Mouse embryonic fibroblasts from different Adam -knockout mice were used in cell-based assays to identify which ADAM is required for processing of individual epidermal-growth-factor-receptor ligands.
Horiuchi, K. et al. Substrate selectivity and regulation of EGF-receptor ligand sheddases by phorbol esters and calcium influx. Mol. Biol. Cell 18, 176–188 (2007).
Hartmann, D. et al. The disintegrin/metalloprotease ADAM 10 is essential for Notch signalling but not for α-secretase activity in fibroblasts. Hum. Mol. Genet. 11, 2615–2624 (2002).
Hinkle, C. L. et al. Selective roles for tumor necrosis factor α-converting enzyme/ADAM17 in the shedding of the epidermal growth factor receptor ligand family: the juxtamembrane stalk determines cleavage efficiency. J. Biol. Chem. 279, 24179–24188 (2004).
Billinghurst, R. C. et al. Enhanced cleavage of type II collagen by collagenases in osteoarthritic articular cartilage. J. Clin. Invest. 99, 1534–1545 (1997).
Gschwind, A., Hart, S., Fischer, O. M. & Ullrich, A. TACE cleavage of proamphiregulin regulates GPCR-induced proliferation and motility of cancer cells. EMBO J. 22, 2411–2421 (2003).
Butler, G. S. & Overall, C. M. Proteomic validation of protease drug targets: Pharmacoproteomics of matrix metalloproteinase inhibitor drugs using isotope-coded affinity tag labelling and tandem mass spectrometry. Curr. Pharm. Des. 13, 263–270 (2007).
Nagano, O. et al. Cell-matrix interaction via CD44 is independently regulated by different metalloproteinases activated in response to extracellular Ca2+ influx and PKC activation. J. Cell Biol. 165, 893–902 (2004).
Prenzel, N. et al. EGF receptor transactivation by G-protein-coupled receptors requires metalloproteinase cleavage of proHB-EGF. Nature 402, 884–888 (1999).
Fan, H. & Derynck, R. Ectodomain shedding of TGF-α and other transmembrane proteins is induced by receptor tyrosine kinase activation and MAP kinase signaling cascades. EMBO J. 18, 6962–6972 (1999).
Hwang, I. K., Park, S. M., Kim, S. Y. & Lee, S. T. A proteomic approach to identify substrates of matrix metalloproteinase-14 in human plasma. Biochim. Biophys. Acta 1702, 79–87 (2004).
Lee, A. Y. et al. Identification of caspase-3 degradome by two-dimensional gel electrophoresis and matrix-assisted laser desorption/ionization–time of flight analysis. Proteomics 4, 3429–3436 (2004).
Zhou, X. W., Blackman, M. J., Howell, S. A. & Carruthers, V. B. Proteomic analysis of cleavage events reveals a dynamic two-step mechanism for proteolysis of a key parasite adhesive complex. Mol. Cell. Proteomics 3, 565–576 (2004).
Bredemeyer, A. J. et al. A proteomic approach for the discovery of protease substrates. Proc. Natl Acad. Sci. USA 101, 11785–11790 (2004). Describes the successful application of DIGE to identify substrates for granzyme A and B.
Bech-Serra, J. J. et al. Proteomic identification of desmoglein 2 and activated leukocyte cell adhesion molecule as substrates of ADAM17 and ADAM10 by difference gel electrophoresis. Mol. Cell. Biol. 26, 5086–5095 (2006). DIGE is used to identify substrates for ADAM10 and ADAM17, and the role of these ADAMs in cleaving the newly identified substrates (desmoglein-2 and activated leukocyte cell-adhesion molecule (ALCAM)) was corroborated with cells from ADAM10- or ADAM17-deficient mice.
Washburn, M. P., Wolters, D. & Yates, J. R. 3rd. Large-scale analysis of the yeast proteome by multidimensional protein identification technology. Nature Biotechnol. 19, 242–247 (2001).
Ahram, M., Adkins, J. N., Auberry, D. L., Wunschel, D. S. & Springer, D. L. A proteomic approach to characterize protein shedding. Proteomics 5, 123–131 (2005).
Guo, L. et al. A proteomic approach for the identification of cell-surface proteins shed by metalloproteases. Mol. Cell. Proteomics 1, 30–36 (2002).
Dean, R. A. & Overall, C. M. Proteomic discovery of metalloproteinase substrates in the cellular context by iTRAQ™ labeling reveals a diverse MMP-2 substrate degradome. Mol. Cell. Proteomics (in the press).
Xia, W. & Wolfe, M. S. Intramembrane proteolysis by presenilin and presenilin-like proteases. J. Cell Sci. 116, 2839–2844 (2003).
Ji, C., Guo, N. & Li, L. Differential dimethyl labeling of N-termini of peptides after guanidination for proteome analysis. J. Proteome Res. 4, 2099–2108 (2005). Clever use of chemical guanidination of Lys side chains to mask these from the free N terminus of proteins, potentially including cleaved substrates, which can be adopted for degradomics analysis of substrate cleavage.
Van Damme, P. et al. Caspase-specific and nonspecific in vivo protein processing during Fas-induced apoptosis. Nature Methods 2, 771–777 (2005). The first comprehensive and working method for N-terminone analysis of proteolysis.
McDonald, L., Robertson, D. H., Hurst, J. L. & Beynon, R. J. Positional proteomics: selective recovery and analysis of N-terminal proteolytic peptides. Nature Methods 2, 955–957 (2005).
Schulz-Knappe, P. et al. Peptidomics: the comprehensive analysis of peptides in complex biological mixtures. Comb. Chem. High Throughput Screen. 4, 207–217 (2001).
Adermann, K., John, H., Standker, L. & Forssmann, W. G. Exploiting natural peptide diversity: novel research tools and drug leads. Curr. Opin. Biotechnol. 15, 599–606 (2004).
Villanueva, J. et al. Serum peptide profiling by magnetic particle-assisted, automated sample processing and MALDI-TOF mass spectrometry. Anal. Chem. 76, 1560–1570 (2004).
Pan, H. et al. The role of prohormone convertase-2 in hypothalamic neuropeptide processing: a quantitative neuropeptidomic study. J. Neurochem. 98, 1763–1777 (2006). Peptidomic analysis of protease-knockout and wild-type mice focusing on neuropeptide products that result from prohormone convertase-2 activity.
Egeblad, M., & Werb, Z. New functions for the matrix metalloproteinases in cancer progression. Nature Rev. Cancer 2, 161–174 (2002).
Gocheva, V. et al. Distinct roles for cysteine cathepsin genes in multistage tumorigenesis. Genes Dev. 20, 543–556 (2006).
Joyce, J. A. et al. Cathepsin cysteine proteases are effectors of invasive growth and angiogenesis during multistage tumorigenesis. Cancer Cell 5, 443–453 (2004).
Lynch, C. C. et al. MMP-7 promotes prostate cancer-induced osteolysis via the solubilization of RANKL. Cancer Cell 7, 485–496 (2005). Microarray analysis of a rodent prostate cancer model, which metastasizes to bone, showed upregulation of MMP7 in osteoclasts at the tumour–bone interface, and demonstrated a requirement for MMP7 in receptor activator of NF-κB ligand (RANKL).
Willem, M. et al. Control of peripheral nerve myelination by the β-secretase BACE1. Science 314, 664–666 (2006). Expression analysis revealed high amounts of BACE in peripheral nerves during myelination, and BACE-deficient mice were found to display hypomyelination and defects in processing of neuregulin, an ErbB-ligand that is crucial for myelination.
Acuff, H. B. et al. Analysis of host- and tumor-derived proteinases using a custom dual species microarray reveals a protective role for stromal matrix metalloproteinase-12 in non-small cell lung cancer. Cancer Res. 66, 7968–7975 (2006).
Peduto, L. et al. ADAM12 is highly expressed in carcinoma-associated stroma and is required for mouse prostate tumor progression. Oncogene 25, 5462–5466 (2006).
Horiuchi, K. et al. Potential role for ADAM15 in pathological neovascularization in mice. Mol. Cell. Biol. 23, 5614–5624 (2003).
Saghatelian, A., Jessani, N., Joseph, A., Humphrey, M. & Cravatt, B. F. Activity-based probes for the proteomic profiling of metalloproteases. Proc. Natl Acad. Sci. USA 101, 10000–10005 (2004).
Stanton, H. et al. ADAMTS5 is the major aggrecanase in mouse cartilage in vivo and in vitro. Nature 434, 648–652 (2005).
Glasson, S. S. et al. Deletion of active ADAMTS5 prevents cartilage degradation in a murine model of osteoarthritis. Nature 434, 644–648 (2005). In references 90 and 91, Adamts4 - and Adamts5 - knockout mice were used to test which of these is the relevant aggrecanase in vitro and in a mouse model for osteo arthritis. ADAMTS5 emerged as the principal enzyme.
Yamazaki, S. et al. Mice with defects in HB-EGF ectodomain shedding show severe developmental abnormalities. J. Cell Biol. 163, 469–475 (2003).
Ruuls, S. R. et al. Membrane-bound TNF supports secondary lymphoid organ structure but is subservient to secreted TNF in driving autoimmune inflammation. Immunity 15, 533–543 (2001).
Ge, G. & Greenspan, D. BMP1 controls TGFβ1 activation via cleavage of latent TGFβ-binding protein. J. Cell Biol. 175, 111–120 (2006).
Alfalah, M. et al. A point mutation in the juxtamembrane stalk of human angiotensin I-converting enzyme invokes the action of a distinct secretase. J. Biol. Chem. 276, 21105–21109 (2001).
Jackson, L. F. et al. Defective valvulogenesis in HB-EGF and TACE-null mice is associated with aberrant BMP signaling. EMBO J. 22, 2704–2716 (2003).
Sternlicht, M. D. et al. Mammary ductal morphogenesis requires paracrine activation of stromal EGFR via ADAM17-dependent shedding of epithelial amphiregulin. Development 132, 3923–3933 (2005).
de Visser, K. E., Korets, L. V. & Coussens, L. M. De novo carcinogenesis promoted by chronic inflammation is B lymphocyte dependent. Cancer Cell 7, 411–423 (2005).
Lammich, S. et al. Constitutive and regulated α-secretase cleavage of Alzheimer's amyloid precursor protein by a disintegrin metalloprotease. Proc. Natl Acad. Sci. USA 96, 3922–3927 (1999).
Schlöndorff, J. S., Lum, L. & Blobel, C. P. Biochemical and pharmacological criteria define two shedding activities for TRANCE/OPGL that are distinct from the TNFα convertase (TACE). J. Biol. Chem. 276, 14665–14674 (2001).
Hikita, A. et al. Negative regulation of osteoclastogenesis by ectodomain shedding of receptor activator of NF-κB ligand. J. Biol. Chem. 281, 36846–36855 (2006).
Sternlicht, M. D. et al. The stromal proteinase MMP3/stromelysin-1 promotes mammary carcinogenesis. Cell 98, 137–146 (1999).
Wen, C., Metzstein, M. M. & Greenwald, I. SUP-17, a Caenorhabditis elegans ADAM protein related to Drosophila KUZBANIAN, and its role in LIN-12/NOTCH signaling. Development 124, 4759–4767 (1997).
Pan, D. & Rubin, J. KUZBANIAN controls proteolytic processing of NOTCH and mediates lateral inhibition during Drosophila and vertebrate neurogenesis. Cell 90, 271–280 (1997).
Li, Q., Park, P. W., Wilson, C. L. & Parks, W. C. Matrilysin shedding of syndecan-1 regulates chemokine mobilization and transepithelial efflux of neutrophils in acute lung injury. Cell 111, 635–646 (2002). An elegant analysis of the role of matrilysin (also known as MMP7) in shedding syndecan-1 with an attached CXC chemokine, and the requirement of this process in neutrophil efflux to sites of lung injury in mice.
Vassar, R. et al. β-secretase cleavage of Alzheimer's amyloid precursor protein by the transmembrane aspartic protease BACE. Science 286, 735–741 (1999).
Roberds, S. L. et al. BACE knockout mice are healthy despite lacking the primary β-secretase activity in brain: implications for Alzheimer's disease therapeutics. Hum. Mol. Genet. 10, 1317–1324 (2001).
Buxbaum, J. D. et al. Evidence that tumor necrosis factor α converting enzyme is involved in regulated α-secretase cleavage of the Alzheimer amyloid protein precursor. J. Biol. Chem. 273, 27765–27767 (1998).
Weskamp, G. et al. ADAM10 is a principal 'sheddase' of the low-affinity immunoglobulin E receptor CD23. Nature Immunology 7, 1393–1398 (2006). Gain- and loss-of-function studies with mouse cells, in vivo shedding studies in mice and use of a selective pharmacological inhibitor on mouse and human B cells identified ADAM10 as the major sheddase for CD23, a target for the treatment of allergic disease and rheumatoid arthritis.
Puente, X. S., Sanchez, L. M., Overall, C. M. & Lopez-Otin, C. Human and mouse proteases: a comparative genomic approach. Nature Rev. Genet. 4, 544–558 (2003). An excellent review on the identification and classification of all proteases in the human genome as well as diseases of proteolysis.
McDermott, M. F. et al. Germline mutations in the extracellular domains of the 55 kDa TNF receptor, TNFR1, define a family of dominantly inherited autoinflammatory syndromes. Cell 97, 133–144 (1999).
Selkoe, D. J. The cell biology of β-amyloid precursor protein and presenilin in Alzheimer's disease. Trends Cell Biol. 8, 447–453 (1998).
Milla, M. E. et al. Specific sequence elements are required for the expression of functional tumor necrosis factor-α-converting enzyme (TACE). J. Biol. Chem. 274, 30563–30570 (1999).
Grams, F. et al. X-ray structures of human neutrophil collagenase complexed with peptide hydroxamate and peptide thiol inhibitors. Implications for substrate binding and rational drug design. Eur. J. Biochem. 228, 830–841 (1995).
Zhao, Y. G., Wei, P. & Sang, Q. X. Inhibitory antibodies against endopeptidase activity of human adamalysin 19. Biochem. Biophys. Res. Commun. 289, 288–294 (2001).
Mort, J. S. & Roughley, P. J. Production of antibodies against degradative neoepitopes in aggrecan. Methods Mol. Med. 100, 237–250 (2004).
Parkin, E. T. et al. Structure-activity relationship of hydroxamate-based inhibitors on the secretases that cleave the amyloid precursor protein, angiotensin converting enzyme, CD23, and pro-tumor necrosis factor-α. Biochemistry 41, 4972–4981 (2002).
Murphy, G. et al. Role of TIMPs (tissue inhibitors of metalloproteinases) in pericellular proteolysis: the specificity is in the detail. Biochem. Soc. Symp., 65–80 (2003).
Gonzales, P. E. et al. Inhibition of the TNFα converting enzyme (TACE) by its Pro domain. J. Biol. Chem. 279, 31638–31645 (2004).
McQuibban, G. A. et al. Matrix metalloproteinase processing of monocyte chemoattractant proteins generates CC chemokine receptor antagonists with anti-inflammatory properties in vivo. Blood 100, 1160–1167 (2002).
Blank, M. & Blind, M. Aptamers as tools for target validation. Curr. Opin. Chem. Biol. 9, 336–342 (2005).
Akashi, H., Matsumoto, S. & Taira, K. Gene discovery by ribozyme and siRNA libraries. Nature Rev. Mol. Cell Biol. 6, 413–422 (2005).
Acknowledgements
C.M.O. is supported by a Canada Research Chair in Metalloproteinase Proteomics and Systems Biology, with research grants from the Canadian Institutes of Health Research, the National Cancer Institute of Canada (with funds raised by the Canadian Cancer Association), and the Canadian Breast Cancer Research Alliance special program grant on metastasis, as well as with a Centre Grant from the Michael Smith Research Foundation. C.P.B. is funded by grants from the National Institutes of Health, from the National Institute of General Medical Sciences and from the Eye Institute, and by a sponsored Research Agreement from Novartis, Basel, Switzerland.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Related links
Related links
FURTHER INFORMATION
Christopher M. Overall's homepage
Human, Mouse and Rat Degradomes
International Proteolysis Society
Glossary
- Extracellular matrix
-
A complex extracellular network of structural proteins, including collagens, glycoproteins and proteoglycans, that supports cell adhesion and migration, and that transmits information through interactions with cell receptors.
- Processing
-
Proteolysis that is distinct from degradation in that it represents a highly specific and efficient, yet limited, activity. Nonetheless, cleaving a protein at only one or two sites can result in a specific change of protein function.
- Ectodomain
-
The extracellular portion of a plasma-membrane protein. In secretory vesicles, the topologically equivalent compartment is the lumen.
- Chemokines
-
A large group of cytokines that elicit chemotactic responses from leukocytes and some other cells that express specific G-protein-coupled chemokine receptors.
- Matrix metalloproteinases
-
(MMPs). A family of 23 metzincin proteases in humans capable of degrading extracellular matrix proteins and of processing many bioactive molecules.
- Neoprotein
-
A protein with a new function generated from a protease-cleavage product that is functionally different from the parent protein.
- Degradome
-
The complete set of proteases that are expressed at a specific time by a cell, tissue or an organism. The degradome of a protease is its substrate repertoire.
- Anti-target
-
A molecule with essential roles in normal cell and tissue function, or life; the down modulation of an anti-target results in clinically unacceptable side effects, initiation of disease or deleteriously alters disease progression.
- Extracellular protease
-
An enzyme that has a catalytic domain in the extracellular compartment or in the lumen of a secretory compartment, both of which are topologically equivalent.
- Proteomics
-
Investigations and techniques for elucidating the proteome.
- Degradomics
-
All genomics, proteomics and systems biology investigations and techniques regarding the genetic, structural and functional identification and characterization of the proteases, inactive homologues, protease substrates and protease inhibitors that are present in an organism.
- Zymography
-
A method for determining protease substrates. Substrates are separated by non-denaturing gel electrophoresis and are incubated with a protease. Negative staining of the stained substrate gel reveals enzymatic activity because the protease has degraded the substrate in the gel.
- Activity-based probe
-
Mechanism-based inhibitor that has been modified by incorporating detection moieties, such as fluorophores, biotin and radioactive elements, to specifically target and visualize individual proteases or a family of proteases in complex samples.
- Non-prime residue
-
A residue in the substrate that is N-terminal to the proteolytic cleavage site is called a non-prime (P) residue, and in some proteases forms part of the recognition motif for substrate cleavage.
- Prime residue
-
A residue in the substrate that is C-terminal of the proteolytic cleavage site is called a prime (P′) residue and in some proteases forms part of the recognition motif for substrate cleavage.
- ADAM proteases
-
A disintegrin and metalloprotease (ADAM) proteases are multifunctional membrane proteins with crucial roles in ectodomain shedding of other membrane proteins, such as the ligands of the epidermal-growth-factor receptor.
- Tissue inhibitors of metalloproteinases (TIMPs)
-
A family of four specific inhibitors of matrix metalloproteinases and some ADAM proteases that are expressed by most mammalian cells.
- Aptamers
-
Aptamers are chemically synthesized (usually short) strands of oligonucleotides (DNA or RNA) that can adopt highly specific three-dimensional conformations. Aptamers are designed to have appropriate binding affinities and specificities towards certain target molecules.
- Exosite
-
A substrate-binding site that lies outside the active-site cleft of a protease. It is usually located on substrate-binding modules or domains and can function to accelerate the rate of substrate cleavage.
- Anti-neoepitope antibody
-
An antibody that specifically recognizes the free amino or carboxyl groups of the amino-acid residues from a cleaved scissile bond that forms the new N and C termini of the cleaved product.
- Proteome
-
The expressed set of proteins that are encoded by the genome and that are expressed by a particular cell or tissue.
- Peptide mapping
-
By proteomically comparing the abundance ratios of multiple peptides from a substrate with their location in the protein sequence, the domain that is proteolytically released can be predicted, as can the general location of the cleavage site.
- Terminopes
-
The N and C termini of a protein are chemically distinguished from the remainder of the intact molecule. Terminopes are generated following proteolytic cleavage. They might also be immunologically recognized as an antibody epitope, called a neoepitope.
- Singletons
-
In proteomics analyses of two peptide samples that are labelled with different isotopes, a singleton is a single ion peak that is detected in the mass spectrometry spectrum without its comparative isotopic counterpart owing to the absence of that parent protein or peptide in one sample. This might occur because of reduced expression or following the cleavage of an intact protein or in the generation of a unique N- or C-terminal peptide following cleavage.
- Driver lines
-
A system, such as the Cre–Lox system, for creating conditional gene deletions.
Rights and permissions
About this article
Cite this article
Overall, C., Blobel, C. In search of partners: linking extracellular proteases to substrates. Nat Rev Mol Cell Biol 8, 245–257 (2007). https://doi.org/10.1038/nrm2120
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrm2120
This article is cited by
-
Knowledge-transfer learning for prediction of matrix metalloprotease substrate-cleavage sites
Scientific Reports (2017)
-
The Transmembrane Serine Protease HAT-like 4 Is Important for Epidermal Barrier Function to Prevent Body Fluid Loss
Scientific Reports (2017)
-
Quantitative analysis of protease recognition by inhibitors in plasma using microscale thermophoresis
Scientific Reports (2016)