Key Points
-
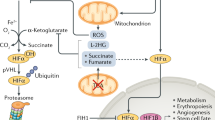
Hypoxia-inducible factor (HIF) is an α/β heterodimeric DNA-binding complex that directs an extensive transcriptional response to hypoxia. The activity of HIF is induced in hypoxic cells through the stabilization and activation of its α-subunit. HIFα subunits are regulated by a newly discovered signalling mechanism — that is, the oxygen-dependent enzymatic hydroxylation of specific amino-acid residues.
-
The hydroxylation of conserved prolyl residues in two independent degradation domains in the central region of HIFα promotes interactions with the von Hippel–Lindau ubiquitylation complex, which targets HIFα for degradation by the ubiquitin–proteasome pathway. Hydroxylation at a conserved asparaginyl residue in the HIFα carboxy-terminal activation domain blocks interaction with the p300 transcriptional co-activator.
-
To date, three HIF prolyl hydroxylases, known as prolyl hydroxylase domain (PHD)1–3, and one asparaginyl hydroxylase, known as factor inhibiting HIF (FIH), have been defined. These enzymes all belong to the non-haem, Fe2+-dependent, 2-oxoglutarate-dependent-oxygenase superfamily.
-
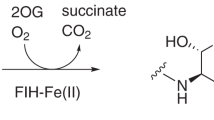
These enzymes possess a common 'jelly-roll' (double-stranded β-helix) core and coordinate the catalytic Fe2+ using a two-histidine, one-carboxylate 'facial triad'. During catalysis, the splitting of molecular oxygen occurs with one oxygen atom being incorporated into the HIF prolyl or asparaginyl residue and the other being incorporated into succinate during the oxidative decarboxylation of 2-oxoglutarate.
-
The absolute requirement of the HIF hydroxylases for molecular oxygen conveys oxygen sensitivity. Additional cofactor and co-substrate requirements for Fe2+, the citric-acid-cycle intermediate 2-oxoglutarate and ascorbate might help these enzymes generate the flexibility that is required for an oxygen-sensing function in complex multicellular animals.
Abstract
The transcription factor HIF (hypoxia-inducible factor) has a central role in oxygen homeostasis in animals ranging from nematode worms to man. Recent studies have shown that this factor is regulated by an unprecedented signalling mechanism that involves post-translational hydroxylation. This hydroxylation is catalysed by a set of non-haem, Fe2+-dependent enzymes that belong to the 2-oxoglutarate-dependent-oxygenase superfamily. The absolute requirement of these enzymes for molecular oxygen has provided new insights into the way cells sense oxygen.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Semenza, G. L. & Wang, G. L. A nuclear factor induced by hypoxia via de novo protein synthesis binds to the human erythropoietin gene enhancer at a site required for transcriptional activation. Mol. Cell. Biol. 12, 5447–5454 (1992).
Maxwell, P. H., Pugh, C. W. & Ratcliffe, P. J. Inducible operation of the erythropoietin 3′ enhancer in multiple cell lines: evidence for a widespread oxygen sensing mechanism. Proc. Natl Acad. Sci. USA 90, 2423–2427 (1993).
Wang, G. L., Jiang, B. -H., Rue, E. A. & Semenza, G. L. Hypoxia-inducible factor 1 is a basic-helix-loop-helix-PAS heterodimer regulated by cellular O2 tension. Proc. Natl Acad. Sci. USA 92, 5510–5514 (1995).
Semenza, G. L. Hypoxia-inducible factor 1: oxygen homeostasis and disease pathophysiology. Trends Mol. Med. 7, 345–350 (2001).
Wenger, R. H. Cellular adaptation to hypoxia: O2-sensing protein hydroxylases, hypoxia-inducible transcription factors, and O2-regulated gene expression. FASEB J. 16, 1151–1162 (2002).
Pugh, C. W. & Ratcliffe, P. J. Regulation of angiogenesis by hypoxia: role of the HIF system. Nature Med. 9, 677–684 (2003).
Huang, L. E. & Bunn, H. F. Hypoxia-inducible factor and its biomedical relevance. J. Biol. Chem. 278, 19575–19578 (2003).
Semenza, G. L. Targeting HIF-1 for Cancer Therapy. Nature Rev. Cancer 3, 721–732 (2003).
Masson, N. & Ratcliffe, P. J. HIF prolyl and asparaginyl hydroxylases in the biological response to intracellular O2 levels. J. Cell Sci. 116, 3041–3049 (2003).
Maynard, M. A. et al. Multiple splice variants of the human HIF-3α locus are targets of the von Hippel–Lindau E3 ubiquitin ligase complex. J. Biol. Chem. 278, 11032–11040 (2003).
Makino, Y. et al. Inhibitory PAS domain protein is a negative regulator of hypoxia-inducible gene expression. Nature 414, 550–554 (2001).
Ivan, M. et al. HIFα targeted for VHL-mediated destruction by proline hydroxylation: implications for O2 sensing. Science 292, 464–468 (2001).
Jaakkola, P. et al. Targeting of HIF-α to the von Hippel–Lindau ubiquitylation complex by O2-regulated prolyl hydroxylation. Science 292, 468–472 (2001).
Yu, F., White, S. B., Zhao, Q. & Lee, F. S. HIF-1α binding to VHL is regulated by stimulus-sensitive proline hydroxylation. Proc. Natl Acad. Sci. USA 98, 9630–9635 (2001). References 12, 13 and 14 define prolyl hydroxylation as the key oxygen-dependent modification that promotes the interaction of HIFαsubunits with pVHL.
Masson, N., Willam, C., Maxwell, P. H., Pugh, C. W. & Ratcliffe, P. J. Independent function of two destruction domains in hypoxia-inducible factor-α chains activated by prolyl hydroxylation. EMBO J. 20, 5197–5206 (2001). This paper shows that HIFα subunits contain two prolyl hydroxylation sites, each of which can interact independently with pVHL.
Maxwell, P. H. et al. The tumour suppressor protein VHL targets hypoxia-inducible factors for oxygen-dependent proteolysis. Nature 399, 271–275 (1999).
Ohh, M. et al. Ubiquitination of hypoxia-inducible factor requires direct binding to the β-domain of the von Hippel–Lindau protein. Nature Cell Biol. 2, 423–427 (2000).
Hon, W. C. et al. Structural basis for the recognition of hydroxyproline in HIF-1α by pVHL. Nature 417, 975–978 (2002).
Min, J. -H. et al. Structure of an HIF-1α–pVHL complex: hydroxyproline recognition in signaling. Science 296, 1886–1889 (2002). References 18 and 19 describe X-ray crystal structures of hydroxylated HIF1α peptides bound to a VBC complex and show how hydrogen bonding to the hydroxylated peptide can discriminate between hydroxylated and non-hydroxylated HIFα subunits.
Epstein, A. C. R. et al. C. elegans EGL-9 and mammalian homologues define a family of dioxygenases that regulate HIF by prolyl hydroxylation. Cell 107, 43–54 (2001).
Bruick, R. K. & McKnight, S. L. A conserved family of prolyl-4-hydroxylases that modify HIF. Science 294, 1337–1340 (2001). References 20 and 21 define the HIFα prolyl hydroxylase (PHD) enzymes and demonstrate their conservation in C. elegans and D. melanogaster.
Lando, D., Peet, D. J., Whelan, D. A., Gorman, J. J. & Whitelaw, M. L. Asparagine hydroxylation of the HIF transactivation domain: a hypoxic switch. Science 295, 858–861 (2002). This paper defines asparaginyl hydroxylation as the key oxygen-dependent modification that regulates HIFα CAD activity by preventing its interaction with the CH-1 domain of p300.
McNeill, L. A. et al. Hypoxia-inducible factor asparaginyl hydroxylase (FIH-1) catalyses hydroxylation at the β-carbon of asparagine-803. J. Biochem. 367, 571–575 (2002).
Freedman, S. J. et al. Structural basis for recruitment of CBP/p300 by hypoxia-inducible factor-1α. Proc. Natl Acad. Sci. USA 99, 5367–5372 (2002).
Dames, S. A., Martinez-Yamout, M., Guzman, R. N. D., Dyson, H. J. & Wright, P. E. Structural basis for Hif-1α/CBP recognition in the cellular hypoxic response. Proc. Natl Acad. Sci. USA 99, 5271–5276 (2002). References 24 and 25 describe the NMR solution structure of HIF1α CAD complexed to the CH-1 domain of p300 and predict how hydroxylation of Asn803 in the HIF1α CAD might disrupt the interaction.
Lando, D. et al. FIH-1 is an asparaginyl hydroxylase enzyme that regulates the transcriptional activity of hypoxia-inducible factor. Genes Dev. 16, 1466–1471 (2002).
Hewitson, K. S. et al. Hypoxia inducible factor (HIF) asparagine hydroxylase is identical to Factor Inhibiting HIF (FIH) and is related to the cupin structural family. J. Biol. Chem. 277, 26351–26355 (2002). References 26 and 27 define FIH as a HIFα asparaginyl hydroxylase.
Mahon, P. C., Hirota, K. & Semenza, G. L. FIH-1: a novel protein that interacts with HIF-1α and VHL to mediate repression of HIF-1 transcriptional activity. Genes Dev. 15, 2675–2686 (2001).
Lavista-Llanos, S. et al. Control of the hypoxic response in Drosophila melanogaster by the basic helix-loop-helix PAS protein Similar. Mol. Cell. Biol. 22, 6842–6853 (2002).
Berra, E. et al. HIF prolyl-hydroxylase 2 is the key oxygen sensor setting low steady-state levels of HIF-1α in normoxia. EMBO J. 22, 4082–4090 (2003). This paper provides genetic evidence for the crucial importance of PHD2 in setting the oxygenated levels of HIF1α
Schofield, C. J. & Zhang, Z. Structural and mechanistic studies on 2-oxoglutarate-dependent oxygenases and related enzymes. Curr. Opin. Struct. Biol. 9, 722–731 (1999).
Hegg, E. L. & Que, L. The 2-His-1-carboxylate facial triad — an emerging structural motif in mononuclear non-heme iron(II) enzymes. Eur. J. Biochem. 250, 625–629 (1997).
Aravind, L. & Koonin, E. V. The DNA-repair protein AlkB, EGL-9, and leprecan define new families of 2-oxoglutarate- and iron-dependent dioxygenases. Genome Biol. 2, RESEARCH0007 (2001).
Valegard, K. et al. Structure of a cephalosporin synthase. Nature 394, 805–809 (1998).
Lee, H. -J. et al. Kinetic and crystallographic studies on deacetoxycephalosporin C synthase (DAOCS). J. Mol. Biol. 308, 937–948 (2001).
Roach, P. L. et al. Crystal-structure of isopenicillin N-synthase is the first from a new structural family of enzymes. Nature 375, 700–704 (1995).
Myllyharju, J. & Kivirikko, K. I. Characterization of the iron- and 2-oxoglutarate-binding sites of human prolyl 4-hydroxylase. EMBO J. 16, 1173–1180 (1997).
Bugg, T. D. H. Oxygenases: mechanisms and structural motifs for O2 activation. Curr. Opin. Chem. Biol. 5, 550–555 (2001).
Dunwell, J. M., Culham, A., Carter, C. E., Sosa-Aguirre, C. R. & Goodenough, P. W. Evolution of functional diversity in the cupin superfamily. Trends Biochem. Sci. 26, 740–746 (2001).
Dann, C. E. I., Bruick, R. K. & Deisenhofer, J. Structure of factor-inhibiting hypoxia-inducible factor 1: an asparaginyl hydroxylase involved in the hypoxic response pathway. Proc. Natl Acad. Sci. USA 99, 15351–15356 (2002).
Elkins, J. M. et al. Structure of factor-inhibiting hypoxia-inducible factor (HIF) reveals mechanism of oxidative modification of HIF-1α. J. Biol. Chem. 278, 1802–1806 (2003).
Lee, C., Kim, S. J., Jeong, D. G., Lee, S. I. & Ryu, S. E. Structure of human FIH-1 reveals a unique active site pocket and interaction sites for HIF-1 and von Hippel–Lindau. J. Biol. Chem. 278, 7558–7563 (2003). References 40, 41 and 42 describe X-ray crystal structures of the HIFα asparaginyl hydroxylase FIH. Reference 41 also describes the structure of the FIH–Fe2+–2-oxoglutarate–HIFα-CAD complex, which highlights an unusual two-site interaction.
Lee, H. -J. et al. Alteration of the co-substrate selectivity of deacetoxycephalosporin C synthase. J. Biol. Chem. 276, 18290–18295 (2001).
Clissold, P. M. & Ponting, C. P. JmjC: cupin metalloenzyme-like domains in jumonji, hairless and phospholipase A2β. Trends Biochem. Sci. 26, 7–9 (2001).
Linke, S. et al. Substrate requirements of the oxygen-sensing asparaginyl hydroxylase factor inhibiting HIF. J. Biol. Chem. 279, 14391–14397 (2004).
Hirsila, M., Koivunen, P., Gunzler, V., Kivirikko, K. I. & Myllyharju, J. Characterization of the human prolyl 4-hydroxylases that modify the hypoxia-inducible factor HIF. J. Biol. Chem. 278, 30772–30780 (2003).
Huang, L. E., Pete, E. A., Schau, M., Milligan, J. & Gu, J. Leu-574 of HIF-1α is essential for the von Hippel–Lindau (VHL)-mediated degradation pathway. J. Biol. Chem. 277, 41750–41755 (2002).
Zhang, Z. et al. Crystal structure of a clavaminate synthase–Fe(II)–2-oxoglutarate–substrate–NO complex: evidence for metal centred rearrangements. FEBS Lett. 517, 7–12 (2002).
Wu, M., Moon, H. S., Begley, T. P., Myllyharju, J. & Kivirikko, K. I. Mechanism-based inactivation of the human prolyl-4-hydroxylase by 5-oxaproline-containing peptides: evidence for a prolyl radical intermediate. J. Am. Chem. Soc. 121, 587–588 (1999).
Rohde, J. -U. et al. Crystallographic and spectroscopic characterisation of a nonheme Fe(IV)=O complex. Science 299, 1037–1039 (2003).
Price, J. C., Barr, E. W., Tirupati, B., Bollinger, J. M. & Krebs, C. The first direct characterization of a high-valent iron intermediate in the reaction of an α-ketoglutarate-dependent dioxygenase: a high-spin Fe(IV) complex in taurine/α-ketoglutarate dioxygenase (TauD) from Escherichia coli. Biochemistry 42, 7497–7508 (2003).
Pavel, E. G. et al. Circular dichroism and magnetic circular dichroism spectroscopic studies of the non-heme ferrous active site in clavaminate synthase and its interaction with α-ketoglutarate cosubstrate. J. Am. Chem. Soc. 120, 743–753 (1998).
Zhou, J. et al. Spectroscopic studies of substrate interactions with clavaminate synthase 2, a multifunctional α-KG-dependent non-heme iron enzyme: correlation with mechanisms and reactivities. J. Am. Chem. Soc. 123, 7388–7398 (2001).
Ryle, M. J. et al. O2 and α-ketoglutarate-dependent tyrosyl formation in TauD, an α-keto acid-dependent non-heme iron dioxygenase. Biochemistry 42, 1854–1862 (2003).
Barlow, J. N., Zhang, Z. H., John, P., Baldwin, J. E. & Schofield, C. J. Inactivation of 1-aminocyclopropane-1-carboxylate oxidase involves oxidative modifications. Biochemistry 36, 3563–3569 (1997).
Kivirikko, K. I. & Myllyharju, J. Prolyl 4-hydroxylases and their protein disulfide isomerase subunit. Matrix Biol. 16, 357–368 (1998).
Wilmouth, R. C. et al. Structure and mechanism of anthocyanidin synthase from Arabidopsis thaliana. Structure 10, 93–103 (2002).
Myllyla, R., Majamaa, K., Gunzler, V., Hanauske-Abel, H. M. & Kivirikko, K. I. Ascorbate is consumed stoichiometrically in the uncoupled reactions catalyzed by prolyl 4-hydroxylase and lysyl hydroxylase. J. Biol. Chem. 259, 5403–5405 (1984).
Jiang, B. -H., Semenza, G. L., Bauer, C. & Marti, H. H. Hypoxia-inducible factor 1 levels vary exponentially over a physiologically relevant range of O2 tension. Am. J. Physiol. 271, C1172–C1180 (1996).
Koivunen, E., Hirsila, M., Gunzler, V., Kivirikko, K. I. & Myllyharju, J. Catalytic properties of the asparaginyl hydroxylase (FIH) in the oxygen sensing pathway are distinct from those of its prolyl-4-hydroxylases. J. Biol. Chem. 279, 9899–9904 (2004). Reference 60 analyses the kinetic properties of the HIF asparaginyl hydroxylase FIH and contrasts them with the properties of the HIF prolyl hydroxylases.
Kimura, H. et al. Hypoxia response element of the human vascular endothelial growth factor gene mediates transcriptional regulation by nitric oxide: control of hypoxia-inducible factor-1 activity by nitric oxide. Blood 95, 189–197 (2000).
Sandau, K. B., Zhou, J., Kietzmann, T. & Brune, B. Regulation of the hypoxia-inducible factor 1α by the inflammatory mediators nitric oxide and tumor necrosis factor-α in contrast to desferroxamine and phenylarsine oxide. J. Biol. Chem. 276, 39805–39811 (2001).
Metzen, E., Zhou, J., Jelkmann, W., Fandrey, J. & Brune, B. Nitric oxide impairs normoxic degradation of HIF-1α by inhibition of prolyl hydroxylases. Mol. Biol. Cell 14, 3470–3481 (2003).
Sogawa, K. et al. Inhibition of hypoxia-inducible factor 1 activity by nitric oxide donors in hypoxia. Proc. Natl Acad. Sci. USA 95, 7368–7373 (1998).
Huang, L. E., Willmore, W. G., Gu, J., Goldberg, M. A. & Bunn, H. F. Inhibition of hypoxia-inducible factor 1 activation by carbon monoxide and nitric oxide. J. Biol. Chem. 274, 9038–9044 (1999).
Wang, F. et al. HIF-1α-prolyl hydroxylase: molecular target of nitric oxide in the hypoxic signal transduction pathway. Biochem. Biophys. Res. Commun. 295, 657–662 (2002).
Hagen, T., Taylor, C. T., Lam, F. & Moncada, S. Redistribution of intracellular oxygen in hypoxia by nitric oxide: effect on HIF-1α. Science 302, 1975–1978 (2003).
Yuan, Y., Hilliard, G., Ferguson, T. & Millhorn, D. E. Cobalt inhibits the interaction between hypoxia inducible factor-α and von Hippel–Lindau protein by direct binding to hypoxia inducible factor-α. J. Biol. Chem. 278, 15911–15916 (2003).
Maxwell, P. H., Pugh, C. W. & Ratcliffe, P. J. Activation of the HIF pathway in cancer. Curr. Opin. Genet. Dev. 11, 293–299 (2001).
Semenza, G. L. Involvement of hypoxia-inducible factor 1 in human cancer. Intern. Med. 41, 79–83 (2002).
Chan, D. A., Sutphin, P. D., Denko, N. C. & Giaccia, A. J. Role of prolyl hydroxylation in oncogenically stabilized hypoxia-inducible factor-1α. J. Biol. Chem. 277, 40112–40117 (2002).
Knowles, H. J., Raval, R. R., Harris, A. L. & Ratcliffe, P. J. Effect of ascorbate on the activity of hypoxia inducible factor (HIF) in cancer cells. Cancer Res. 63, 1764–1768 (2003).
Le, N. T. V. & Richardson, D. R. The role of iron in cell cycle progression and the proliferation of neoplastic cells. Biochim. Biophys. Acta 1603, 31–46 (2002).
Falnes, P. O., Johansen, R. F. & Seeberg, E. AlkB-mediated oxidative demethylation reverses DNA damage in Escherichia coli. Nature 419, 178–182 (2002).
Trewick, S. C., Henshaw, T. F., Hausinger, R. P., Lindahl, T. & Sedgwick, B. Oxidative demethylation by Escherichia coli AlkB directly reverts DNA base damage. Nature 419, 174–178 (2002).
Baysal, B. E. et al. Mutations in SDHD, a mitochondrial complex II gene, in hereditary paraganglioma. Science 287, 848–851 (2000).
Baysal, B. E. Hereditary paraganglioma targets diverse paraganglia. J. Med. Genet. 39, 617–622 (2002).
Tomlinson, I. P. et al. Germline mutations in FH predispose to dominantly inherited uterine fibroids, skin leiomyomata and papillary renal cell cancer. Nature Genet. 30, 406–410 (2002).
Lieb, M. E., Menzies, K., Moschella, M. C., Ni, R. & Taubman, M. B. Mammalian EGLN genes have distinct patterns of mRNA expression and regulation. Biochem. Cell Biol. 80, 421–426 (2002).
Berra, E., Richard, D. E., Gothie, E. & Pouyssegur, J. HIF-1-dependent transcriptional activity is required for oxygen-mediated HIF-1α degradation. FEBS Lett. 491, 85–90 (2001).
del Peso, L. et al. The von Hippel–Lindau/hypoxia-inducible factor (HIF) pathway regulates the transcription of the HIF-proline hydroxylase genes in response to low oxygen. J. Biol. Chem. 278, 48690–48695 (2003).
Wax, S. D., Rosenfield, C. L. & Taubman, M. B. Identification of a novel growth factor-responsive gene in vascular smooth muscle cells. J. Biol. Chem. 269, 13041–13047 (1994).
Madden, S. L., Galella, E. A., Riley, D., Bertelsen, A. H. & Beaudry, G. A. Induction of cell growth regulatory genes by p53. Cancer Res. 56, 5384–5390 (1996).
Lipscomb, E. A., Sarmiere, P. D. & Freeman, R. S. SM-20 is a novel mitochondrial protein that causes caspase-dependent cell death in nerve growth factor-dependent neurons. J. Biol. Chem. 276, 5085–5092 (2001).
Seth, P. I. K., Porter, D. & Polyak, K. Novel estrogen and tamoxifen induced genes identified by SAGE (serial analysis of gene expression). Oncogene 21, 836–843 (2002).
Erez, N. et al. Expression of prolyl-hydroxylase-1 (PHD1/EGLN2) suppresses hypoxia inducible factor-1α activation and inhibits tumor growth. Cancer Res. 63, 8777–8783 (2003).
Gradin, K., Takasaki, C., Fujii-Kuriyama, Y. & Sogawa, K. The transcriptional activation function of the HIF-like factor requires phosphorylation at a conserved threonine. J. Biol. Chem. 277, 23508–23514 (2002).
Kuznetsova, A. V. et al. von Hippel–Lindau protein binds hyperphosphorylated large subunit of RNA polymerase II through a proline hydroxylation motif and targets it for ubiquitination. Proc. Natl Acad. Sci. USA 100, 2706–2711 (2003).
Hanson, E. S., Rawlins, M. L. & Leibold, E. A. Oxygen and iron regulation of iron regulatory protein 2. J. Biol. Chem. 278, 40337–40342 (2003).
Martin-Puig, S. et al. Role of iron (II)-2-oxoglutarate-dependent dioxygenases in the generation of hypoxia-induced phosphatidic acid through HIF1/2 and VHL-independent mechanisms. J. Biol. Chem. 279, 9504–9511 (2004).
Lavaissiere, L. et al. Overexpression of human aspartyl(asparaginyl) β-hydroxylase in hepatocellular carcinoma and cholangiocarcinoma. J. Clin. Invest. 98, 1313–1323 (1996).
Ivan, M. et al. Biochemical purification and pharmacological inhibition of a mammalian prolyl hydroxylase acting on hypoxia-inducible factor. Proc. Natl Acad. Sci. USA 99, 13459–13464 (2002).
Schlemminger, I. et al. Analogues of dealanylalahopcin are inhibitors of human HIF prolyl hydroxylases. Bioorg. Med. Chem. Lett. 13, 1451–1454 (2003).
Mole, D. R. et al. 2-oxoglutarate analogue inhibitors of HIF prolyl hydroxylase. Bioorg. Med. Chem. Lett. 13, 2677–2680 (2003).
Warnecke, C. et al. Activation of the hypoxia-inducible factor-pathway and stimulation of angiogenesis by application of prolyl hydroxylase inhibitors. FASEB J. 17, 1186–1188 (2003).
Zhang, Z. H. et al. Structural origins of the selectivity of the trifunctional oxygenase clavaminic acid synthase. Nature Struct. Biol. 7, 127–133 (2000).
Stapon, A., Li, R. F. & Townsend, C. A. Carbapenem biosynthesis: confirmation of stereochemical assignments and the role of CarC in the ring stereoinversion process from L-proline. J. Am. Chem. Soc. 125, 8486–8493 (2003).
Clifton, I. J. et al. Crystal structure of carbapenem synthase (CarC). J. Biol. Chem. 278, 20843–20850 (2003).
Kivirikko, K. I., Myllyla, R. & Pihlajaniemi, T. Protein hydroxylation: prolyl 4-hydroxylase, an enzyme with four cosubstrates and a multifunctional subunit. FASEB J. 3, 1609–1617 (1989).
Vaz, F. M. & Wanders, R. J. A. Carnitine biosynthesis in mammals. Biochem. J. 361, 417–429 (2002).
Jansen, G. A. et al. Human phytanoyl CoA hydroxylase: resolution of the gene structure and the molecular basis of Refsum's disease. Hum. Mol. Genet. 9, 1195–1200 (2000).
Aas, P. A. et al. Human and bacterial oxidative demethylases repair alkylation damage in both RNA and DNA. Nature 421, 859–863 (2003).
Welford, R. W. D., Schlemminger, I., McNeill, L. A., Hewitson, K. S. & Schofield, C. J. The selectivity and inhibition of AlkB. J. Biol. Chem. 278, 10157–10161 (2003).
Koong, A. C. et al. Candidate genes for the hypoxic tumor phenotype. Cancer Res. 60, 883–887 (2000).
Wykoff, C. C., Pugh, C. W., Maxwell, P. H., Harris, A. L. & Ratcliffe, P. J. Identification of novel hypoxia-dependent and independent target genes of the von Hippel–Lindau (VHL) tumor suppressor by mRNA differential expression profiling. Oncogene 19, 6297–6305 (2000).
Jiang, Y. et al. Gene expression profiling in a renal cell carcinoma cell line: dissecting VHL and hypoxia-dependent pathways. Mol. Cancer Res. 1, 453–462 (2003).
Fandrey, J. & Bunn, H. F. In vivo and in vitro regulation of the erythropoietin mRNA: measurement by competitive polymerase chain reaction. Blood 81, 617–623 (1993).
Rosenberger, C. et al. Expression of hypoxia-inducible factor-1α and -2α in hypoxic and ischemic rat kidneys. J. Am. Soc. Nephrol. 13, 1721–1732 (2002).
Acknowledgements
We thank all our colleagues for their dedicated contributions to the Oxford work in the area of hypoxic signalling, T. Murray-Rust for assistance with the figures, and the Wellcome Trust, the Biotechnology and Biological Sciences Research Council, the Medical Research Council and the European Union for funding our research.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors are founding scientists of ReOx Ltd., a company that aims to exploit the HIF hydroxylases as therapeutic targets.
Supplementary information
Glossary
- PAS FAMILY
-
(period circadian protein (PER), aryl-hydrocarbon receptor (AHR), aryl-hydrocarbon-receptor nuclear translocator (ARNT) and single-minded protein (SIM)). A group of interacting and structurally related basic helix–loop–helix transcription factors.
- ANGIOGENESIS
-
The growth and proliferation of new blood vessels from existing vasculature.
- ERYTHROPOIESIS
-
The formation of erythrocytes (red blood cells).
- UBIQUITIN–PROTEASOME PATHWAY
-
A system of selective, energy (ATP)-consuming protein degradation that involves the linking of ubiquitins to specific proteins and the subsequent targeting of these polyubiquitylated proteins to the 26S proteasome (a multi-catalytic protease).
- CUPIN SUPERFAMILY
-
The cupins are a diverse family of plant proteins, all of which contain at least one double-stranded β-helix or jelly-roll strucural motif. This motif is also present in all structurally characterized 2-oxoglutarate-dependent oxygenases including the HIF hydroxylases, and is characteristic of the jumonji transcription factors.
- TRIPLET-STATE MOLECULAR OXYGEN
-
Most 'natural' molecules exist in the singlet state — that is, they contain paired electrons. However, the most stable form of molecular oxygen (O2) is the triplet state, in which there are two unpaired electrons.
- K m
-
The Michaelis constant. A kinetic parameter for a specific substrate in an enzyme-catalysed reaction. Providing certain conditions are met, the Km for a substrate can equate to its binding constant, and the lower the value of Km, the tighter the substrate binds.
- pO2
-
The partial pressure that is exerted by molecular oxygen in a mixture of gases. It is also used to define the concentration of molecular oxygen in a solution or biological tissue that is at equilibrium with such a gas mixture.
- SYSTEMIC HYPOXIA
-
A reduction in the partial pressure of oxygen (pO2) throughout the organism.
- TISSUE ISCHAEMIA
-
Inadequate blood supply to a tissue, which causes other metabolic abnormalities in addition to hypoxia (for example, the defective delivery of substrates and removal of waste products).
Rights and permissions
About this article
Cite this article
Schofield, C., Ratcliffe, P. Oxygen sensing by HIF hydroxylases. Nat Rev Mol Cell Biol 5, 343–354 (2004). https://doi.org/10.1038/nrm1366
Issue Date:
DOI: https://doi.org/10.1038/nrm1366
This article is cited by
-
Hypoxia, oxidative stress, and the interplay of HIFs and NRF2 signaling in cancer
Experimental & Molecular Medicine (2024)
-
Role of pH-sensing receptors in colitis
Pflügers Archiv - European Journal of Physiology (2024)
-
A novel scenario in the therapeutic management of anemia of chronic kidney disease: placement and use of roxadustat
Journal of Nephrology (2024)
-
Glycolytic enzymes in non-glycolytic web: functional analysis of the key players
Cell Biochemistry and Biophysics (2024)
-
Acute hypoxia causes glycolysis and lipid metabolism alterations with HSPs’ persistent activation in Leiocassis longirostris
Ichthyological Research (2024)