Key Points
-
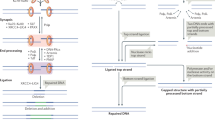
Mammalian non-homologous DNA end joining (NHEJ) is the primary pathway for the repair of DNA double-strand breaks (DSBs) throughout the cell cycle, including during S and G2 phases.
-
NHEJ relies on the Ku protein to thread onto each broken DNA end. Ku recruits the enzymes and complexes that are needed to trim (nucleases) or to fill in (polymerases) the ends to make them optimally ligatable by the DNA ligase IV complex.
-
The configuration of the DNA ends determines which of several subpathways of NHEJ is able to join the ends. Because NHEJ is flexible and iterative, any of these subpathways can be used but some pathways are more efficient than others for certain DNA ends.
-
When NHEJ is absent owing to a lack of Ku or the DNA ligase complex, alternative end joining (a-EJ) can join the ends using microhomology (usually >4 bp) and there is often some evidence of templated insertions of substantial length (>10 nucleotides). DNA polymerase θ (Pol θ) is of key importance for a-EJ.
-
The single-strand annealing (SSA) pathway requires further end resection by exonuclease 1 (EXO1), Bloom syndrome RecQ-like helicase (BLM) or DNA replication helicase/nuclease 2 (DNA2) to generate the long 3′ single-strand DNA (ssDNA) tails (>20 nucleotides) that are bound by replication protein A (RPA) to prevent the formation of DNA secondary structures. The 3′ ssDNA tails are annealed by RAD52.
Abstract
DNA double-strand breaks (DSBs) are the most dangerous type of DNA damage because they can result in the loss of large chromosomal regions. In all mammalian cells, DSBs that occur throughout the cell cycle are repaired predominantly by the non-homologous DNA end joining (NHEJ) pathway. Defects in NHEJ result in sensitivity to ionizing radiation and the ablation of lymphocytes. The NHEJ pathway utilizes proteins that recognize, resect, polymerize and ligate the DNA ends in a flexible manner. This flexibility permits NHEJ to function on a wide range of DNA-end configurations, with the resulting repaired DNA junctions often containing mutations. In this Review, we discuss the most recent findings regarding the relative involvement of the different NHEJ proteins in the repair of various DNA-end configurations. We also discuss the shunting of DNA-end repair to the auxiliary pathways of alternative end joining (a-EJ) or single-strand annealing (SSA) and the relevance of these different pathways to human disease.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Lieber, M. R. The mechanism of double-strand DNA break repair by the nonhomologous DNA end-joining pathway. Annu. Rev. Biochem. 79, 181–211 (2010).
Lieber, M. R. & Karanjawala, Z. E. Ageing, repetitive genomes and DNA damage. Nat. Rev. Mol. Cell Biol. 5, 69–75 (2004).
Martin, G. M., Smith, A. C., Ketterer, D. J., Ogburn, C. E. & Disteche, C. M. Increased chromosomal aberrations in first metaphases of cells isolated from the kidneys of aged mice. Isr. J. Med. Sci. 21, 296–301 (1985).
Meek, K., Dang, V. & Lees-Miller, S. P. DNA-PK: the means to justify the ends? Adv. Immunol. 99, 33–58 (2008).
Chang, H. H. Y. et al. Different DNA end configurations dictate which NHEJ components are most important for joining efficiency. J. Biol. Chem. 291, 24377–24389 (2016). This paper describes a biochemical reconstitution of NHEJ using a direct gel assay (without PCR) and all of the major NHEJ components.
Goodarzi, A. A. et al. DNA-PK autophosphorylation facilitates Artemis endonuclease activity. EMBO J. 25, 3880–3889 (2006).
Gu, J. et al. DNA-PKcs regulates a single-stranded DNA endonuclease activity of Artemis. DNA Repair (Amst.) 9, 429–437 (2010).
Chang, H. H., Watanabe, G. & Lieber, M. R. Unifying the DNA end-processing roles of the artemis nuclease: Ku-dependent Artemis resection at blunt DNA ends. J. Biol. Chem. 290, 24036–24050 (2015).
Chang, H. H. & Lieber, M. R. Structure-specific nuclease activities of Artemis and the Artemis: DNA-PKcs complex. Nucleic Acids Res. 44, 4991–4997 (2016).
Dominski, Z. Nucleases of the metallo-β-lactamase family and their role in DNA and RNA metabolism. Crit. Rev. Biochem. Mol. Biol. 42, 67–93 (2007).
Li, S. et al. Evidence that the DNA endonuclease ARTEMIS also has intrinsic 5′-exonuclease activity. J. Biol. Chem. 289, 7825–7834 (2014).
Moshous, D. et al. Artemis, a novel DNA double-strand break repair/V(D)J recombination protein, is mutated in human severe combined immune deficiency. Cell 105, 177–186 (2001). This paper reports that Artemis is mutated in human SCID.
Ma, Y., Pannicke, U., Schwarz, K. & Lieber, M. R. Hairpin opening and overhang processing by an Artemis/DNA-dependent protein kinase complex in nonhomologous end joining and V(D)J recombination. Cell 108, 781–794 (2002). This paper reports the discovery that Artemis is a ss–ds endonuclease (and a 5′ exonuclease).
Niewolik, D. et al. DNA-PKcs dependence of artemis endonucleolytic activity: differences between hairpins and 5′ or 3′ overhangs. J. Biol. Chem. 281, 33900–33909 (2006).
Niewolik, D., Peter, I., Butscher, C. & Schwarz, K. Autoinhibition of the nuclease ARTEMIS is mediated by a physical interaction between its catalytic and C-terminal domains. J. Biol. Chem. 292, 3351–3365 (2017).
Malu, S. et al. Artemis C-terminal region facilitates V(D)J recombination through its interactions with DNA Ligase IV and DNA-PKcs. J. Exp. Med. 209, 955–963 (2012).
De Ioannes, P., Malu, S., Cortes, P. & Aggarwal, A. K. Structural basis of DNA ligase IV–Artemis interaction in nonhomologous end-joining. Cell Rep. 2, 1505–1512 (2012).
Riballo, E. et al. A pathway of double-strand break rejoining dependent upon ATM, Artemis, and proteins locating to γ-H2AX foci. Mol. Cell 16, 715–724 (2004).
Kurosawa, A. et al. The requirement of Artemis in double-strand break repair depends on the type of DNA damage. DNA Cell Biol. 27, 55–61 (2008).
Kanno, S. et al. A novel human AP endonuclease with conserved zinc-finger-like motifs involved in DNA strand break responses. EMBO J. 26, 2094–2103 (2007).
Li, S. et al. Polynucleotide kinase and aprataxin-like forkhead-associated protein (PALF) acts as both a single-stranded DNA endonuclease and a single-stranded DNA 3′ exonuclease and can participate in DNA end joining in a biochemical system. J. Biol. Chem. 286, 36368–36377 (2011).
Grundy, G. J. et al. APLF promotes the assembly and activity of non-homologous end joining protein complexes. EMBO J. 32, 112–125 (2013).
Pannunzio, N. R., Li, S., Watanabe, G. & Lieber, M. R. Nonhomologous end joining often uses microhomology: implications for alternative end joining. DNA Repair (Amst.) 17, 74–80 (2014).
Bebenek, K., Pedersen, L. C. & Kunkel, T. A. Structure–function studies of DNA polymerase lambda. Biochemistry 53, 2781–2792 (2014).
Moon, A. F. et al. Sustained active site rigidity during synthesis by human DNA polymerase μ. Nat. Struct. Mol. Biol. 21, 253–260 (2014).
Ma, Y. et al. A biochemically defined system for mammalian nonhomologous DNA end joining. Mol. Cell 16, 701–713 (2004).
Bertocci, B., DeSmet, A., Weill, J.-C. & Reynaud, C. A. Non-overlapping functions of polX family DNA polymerases, pol m, pol l, and TdT, during immunoglobulin V(D)J recombination in vivo. Immunity 25, 31–41 (2006).
Pryor, J. M. et al. Essential role for polymerase specialization in cellular nonhomologous end joining. Proc. Natl Acad. Sci. USA 112, E4537–E4545 (2015).
NickMcElhinny, S. A. & Ramsden, D. A. Polymerase μ is a DNA-directed DNA/RNA polymerase. Mol. Cell. Biol. 23, 2309–2315 (2003).
NickMcElhinny, S. A. et al. A gradient of template dependence defines distinct biological roles for family X polymerases in nonhomologous end joining. Mol. Cell 19, 357–366 (2005).
Gu, J. et al. XRCC4:DNA ligase IV can ligate incompatible DNA ends and can ligate across gaps. EMBO J. 26, 1010–1023 (2007).
Lieber, M. R. The polymerases for V(D)J recombination. Immunity 25, 7–9 (2006).
Daley, J. M., Laan, R. L. V., Suresh, A. & Wilson, T. E. DNA joint dependence of pol X family polymerase action in nonhomologous end joining. J. Biol. Chem. 280, 29030–29037 (2005).
Daley, J. M., Palmbos, P. L., Wu, D. & Wilson, T. E. Nonhomologous end joining in yeast. Ann. Rev. Genet. 39, 431–451 (2005).
Daley, J. M. & Wilson, T. E. Rejoining of DNA double-strand breaks as a function of overhang length. Mol. Cell. Biol. 25, 896–906 (2005).
Daley, J. M. & Wilson, T. E. Evidence that base stacking potential in annealed 3′ overhangs determines polymerase utilization in yeast nonhomologous end joining. DNA Repair (Amst.) 7, 67–76 (2007).
Grawunder, U. et al. Activity of DNA ligase IV stimulated by complex formation with XRCC4 protein in mammalian cells. Nature 388, 492–495 (1997).
Dai, Y. et al. Nonhomologous end joining and V(D)J recombination require an additional factor. Proc. Natl Acad. Sci. USA 100, 2462–2467 (2003).
Ahnesorg, P., Smith, P. & Jackson, S. P. XLF interacts with the XRCC4–DNA ligase IV complex to promote nonhomologous end-joining. Cell 124, 301–313 (2006).
Buck, D. et al. Cernunnos, a novel nonhomologous end-joining factor, is mutated in human immunodeficiency with microcephaly. Cell 124, 287–299 (2006). References 39 and 40 describe the discovery of Cernunnos (also known as NEJ1 and XLF).
Brouwer, I. et al. Sliding sleeves of XRCC4–XLF bridge DNA and connect fragments of broken DNA. Nature 535, 566–569 (2016).
Ochi, T. et al. DNA repair. PAXX, a paralog of XRCC4 and XLF, interacts with Ku to promote DNA double-strand break repair. Science 347, 185–188 (2015).
Xing, M. et al. Interactome analysis identifies a new paralogue of XRCC4 in non-homologous end joining DNA repair pathway. Nat. Commun. 6, 6233 (2015). References 42 and 43 describe the discovery of PAXX.
Roy, S. et al. XRCC4/XLF interaction is variably required for DNA repair and is not required for ligase IV stimulation. Mol. Cell. Biol. 35, 3017–3028 (2015).
Tadi, S. K. et al. PAXX is an accessory c-NHEJ factor that associates with Ku70 and has overlapping functions with XLF. Cell Rep. 17, 541–555 (2016).
Bernstein, N. K. et al. The molecular architecture of the mammalian DNA repair enzyme, polynucleotide kinase. Mol. Cell 17, 657–670 (2005).
Ahel, I. et al. The neurodegenerative disease protein aprataxin resolves abortive DNA ligation intermediates. Nature 443, 713–716 (2006).
Koch, C. A. et al. Xrcc4 physically links DNA end processing by polynucleotide kinase to DNA ligation by DNA ligase IV. EMBO J. 23, 3874–3885 (2004).
Inamdar, K. V. et al. Conversion of phosphoglycolate to phosphate termini on 3′ overhangs of DNA double strand breaks by the human tyrosyl-DNA phosphodiesterase hTdp1. J. Biol. Chem. 277, 27162–27168 (2002).
Chen, B. et al. GC/MS methods to quantify the 2-deoxypentos-4-ulose and 3′-phosphoglycolate pathways of 4′ oxidation of 2-deoxyribose in DNA: application to DNA damage produced by gamma radiation and bleomycin. Chem. Res. Toxicol. 20, 1701–1708 (2007).
Zhou, T. et al. Deficiency in 3′-phosphoglycolate processing in human cells with a hereditary mutation in tyrosyl-DNA phosphodiesterase (TDP1). Nucleic Acids Res. 33, 289–297 (2005).
Sinden, R. R. & Wells, R. D. DNA structure, mutations and human genetic diseases. Curr. Opin. Biotechnol. 3, 612–622 (1992).
Mimori, T. & Hardin, J. A. Mechanism of interaction between Ku protein and DNA. J. Biol. Chem. 261, 10375–10379 (1986).
Costantini, S., Woodbine, L., Andreoli, L., Jeggo, P. A. & Vindigni, A. Interaction of the Ku heterodimer with the DNA ligase IV/Xrcc4 complex and its regulation by DNA-PK. DNA Repair (Amst.) 6, 712–722 (2007).
Sibanda, B. L. et al. Crystal structure of an Xrcc4–DNA ligase IV complex. Nat. Struct. Biol. 8, 1015–1019 (2001).
Grawunder, U., Zimmer, D., Kulesza, P. & Lieber, M. R. Requirement for an interaction of XRCC4 with DNA ligase IV for wild-type V(D)J recombination and DNA double-strand break repair in vivo. J. Biol. Chem. 273, 24708–24714 (1998).
Grawunder, U., Zimmer, D. & Lieber, M. R. DNA ligase IV binds to XRCC4 via a motif located between rather than within its BRCT domains. Curr. Biol. 8, 873–876 (1998).
NickMcElhinny, S. A., Snowden, C. M., McCarville, J. & Ramsden, D. A. Ku recruits the XRCC4–ligase IV complex to DNA ends. Mol. Cell. Biol. 20, 2996–3003 (2000).
Herrmann, G., Lindahl, T. & Schar, P. S. cerevisiae LIF1: a function involved in DNA double-strand break repair related to mammalian XRCC4. EMBO J. 17, 4188–4198 (1998).
Westmoreland, J. W., Summers, J. A., Holland, C. L., Resnick, M. A. & Lewis, L. K. Blunt-ended DNA double-strand breaks induced by endonucleases PvuII and EcoRV are poor substrates for repair in Saccharomyces cerevisiae. DNA Repair (Amst.) 9, 617–626 (2010).
Sibanda, B. L., Chirgadze, D. Y. & Blundell, T. L. Crystal structure of DNA-PKcs reveals a large open-ring cradle comprised of HEAT repeats. Nature 463, 118–121 (2010). This paper describes the crystal structure of DNA-PKcs at 6.6 Å.
Sibanda, B. L., Chirgadze, D. Y., Ascher, D. B. & Blundell, T. L. DNA-PKcs structure suggests an allosteric mechanism modulating DNA double-strand break repair. Science 355, 520–524 (2017). This paper describes the crystal structure of DNA-PKcs at 4.3 Å. Although no DNA is present, the 20 kDa portion of the C terminus of Ku80 is present.
West, R. B., Yaneva, M. & Lieber, M. R. Productive and nonproductive complexes of Ku and DNA-PK at DNA termini. Mol. Cell. Biol. 18, 5908–5920 (1998).
Spagnolo, L., Rivera-Calzada, A., Pearl, L. H. & Llorca, O. Three-dimensional structure of the human DNA-PKcs/Ku70/Ku80 complex assembled on DNA and its implications for DNA DSB repair. Mol. Cell 22, 511–519 (2006).
Ma, Y. et al. The DNA-PKcs phosphorylation sites of human artemis. J. Biol. Chem. 280, 33839–33846 (2005).
Lu, H. et al. A biochemically defined system for coding joint formation in human V(D)J recombination. Mol. Cell 31, 485–497 (2008).
Blommers, M. J. et al. Effects of base sequence on the loop folding in DNA hairpins. Biochemistry 28, 7491–7498 (1989).
Povirk, L. F. Processing of damaged DNA ends for double-strand break repair in mammalian cells. ISRN Mol. Biol. 2012, 345805 (2012).
Henner, W. D., Grunberg, S. M. & Haseltine, W. A. Enzyme action at 3′ termini of ionizing radiation-induced DNA strand breaks. J. Biol. Chem. 258, 15198–15205 (1983).
Henner, W. D., Rodriguez, L. O., Hecht, S. M. & Haseltine, W. A. γ ray induced deoxyribonucleic acid strand breaks. 3′ glycolate termini. J. Biol. Chem. 258, 711–713 (1983).
Valerie, K. & Povirk, L. F. Regulation and mechanisms of mammalian double-strand break repair. Oncogene 22, 5792–5812 (2003).
Povirk, L. F., Zhou, T., Zhou, R., Cowan, M. J. & Yannone, S. M. Processing of 3′-phosphoglycolate-terminated DNA double strand breaks by Artemis nuclease. J. Biol. Chem. 282, 3547–3558 (2007).
Yannone, S. M. et al. Coordinate 5′ and 3′ endonucleolytic trimming of terminally blocked blunt DNA double-strand break ends by Artemis nuclease and DNA-dependent protein kinase. Nucleic Acids Res. 36, 3354–3365 (2008).
Ochi, T., Gu, X. & Blundell, T. L. Structure of the catalytic region of DNA ligase IV in complex with an Artemis fragment sheds light on double-strand break repair. Structure 21, 672–679 (2013). References 16, 17 and 74 describe the interaction of Artemis with DNA ligase IV.
Moon, A. F. et al. Structural insight into the substrate specificity of DNA polymerase mu. Nat. Struc. Mol. Biol. 14, 45–53 (2007).
Junop, M. S. et al. Crystal structure of the XRCC4 DNA repair protein and implications for end joining. EMBO J. 19, 5962–5970 (2000).
Andres, S. N., Modesti, M., Tsai, C. J., Chu, G. & Junop, M. S. Crystal structure of human XLF: a twist in nonhomologous DNA end-joining. Mol. Cell 28, 1093–1101 (2007).
Tsai, C. J., Kim, S. A. & Chu, G. Cernunnos/XLF promotes the ligation of mismatched and noncohesive DNA ends. Proc. Natl Acad. Sci. USA 104, 7851–7856 (2007).
Liu, X., Shao, Z., Jiang, W., Lee, B. J. & Zha, S. PAXX promotes KU accumulation at DNA breaks and is essential for end-joining in XLF-deficient mice. Nat. Commun. 8, 13816 (2017).
Balmus, G. et al. Synthetic lethality between PAXX and XLF in mammalian development. Genes Dev. 30, 2152–2157 (2016).
Lescale, C. et al. Specific roles of XRCC4 paralogs PAXX and XLF during V(D)J recombination. Cell Rep. 16, 2967–2979 (2016).
Kumar, V., Alt, F. W. & Frock, R. L. PAXX and XLF DNA repair factors are functionally redundant in joining DNA breaks in a G1-arrested progenitor B-cell line. Proc. Natl Acad. Sci. USA 113, 10619–10624 (2016).
Saito, S., Kurosawa, A. & Adachi, N. Mutations in XRCC4 cause primordial dwarfism without causing immunodeficiency. J. Hum. Genet. 61, 679–685 (2016).
Gauss, G. H. & Lieber, M. R. Mechanistic constraints on diversity in human V(D)J recombination. Mol. Cell. Biol. 16, 258–269 (1996).
Bhargava, R., Onyango, D. O. & Stark, J. M. Regulation of single-strand annealing and its role in genome maintenance. Trends Genet. 32, 566–575 (2016).
Lisby, M. & Rothstein, R. Cell biology of mitotic recombination. Cold Spring Harb. Perspect. Biol. 7, a016535 (2015).
Haber, J. E. Mating-type gene switching in Saccharomyces cerevisiae. Annu. Rev. Genet. 32, 561–599 (1998).
Moynahan, M. E. & Jasin, M. Mitotic homologous recombination maintains genomic stability and suppresses tumorigenesis. Nat. Rev. Mol. Cell Biol. 11, 196–207 (2010).
Mimitou, E. P. & Symington, L. S. Ku prevents Exo1 and Sgs1-dependent resection of DNA ends in the absence of a functional MRX complex or Sae2. EMBO J. 29, 3358–3369 (2010).
Escribano-Diaz, C. et al. A cell cycle-dependent regulatory circuit composed of 53BP1-RIF1 and BRCA1-CtIP controls DNA repair pathway choice. Mol. Cell 49, 872–883 (2013).
Daley, J. M., Niu, H., Miller, A. S. & Sung, P. Biochemical mechanism of DSB end resection and its regulation. DNA Repair (Amst.) 32, 66–74 (2015).
Xie, A. et al. Distinct roles of chromatin-associated proteins MDC1 and 53BP1 in mammalian double-strand break repair. Mol. Cell 28, 1045–1057 (2007).
Koole, W. et al. A polymerase theta-dependent repair pathway suppresses extensive genomic instability at endogenous G4 DNA sites. Nat. Commun. 5, 3216 (2014).
Wood, R. D. & Doublie, S. DNA polymerase θ (POLQ), double-strand break repair, and cancer. DNA Repair (Amst.) 44, 22–32 (2016). This paper comprehensively summarizes the current information about Pol θ.
Wyatt, D. W. et al. Essential roles for polymerase theta-mediated end joining in the repair of chromosome breaks. Mol. Cell 63, 662–673 (2016).
Chan, S. H., Yu, A. M. & McVey, M. Dual roles for DNA polymerase θ in alternative end-joining repair of double-strand breaks in Drosophila. PLoS Genet. 6, e1001005 (2010).
Kent, T., Mateos-Gomez, P. A., Sfeir, A. & Pomerantz, R. T. Polymerase θ is a robust terminal transferase that oscillates between three different mechanisms during end-joining. eLife 5, e13740 (2016).
Mateos-Gomez, P. A. et al. Mammalian polymerase θ promotes alternative NHEJ and suppresses recombination. Nature 518, 254–257 (2015).
Yousefzadeh, M. J. et al. Mechanism of suppression of chromosomal instability by DNA polymerase POLQ. PLoS Genet. 10, e1004654 (2014).
Chen, X. et al. Human DNA ligases I, III, and IV-purification and new specific assays for these enzymes. Methods Enzymol. 409, 39–52 (2006).
Han, L. & Yu, K. Altered kinetics of nonhomologous end joining and class switch recombination in ligase IV-deficient B cells. J. Exp. Med. 205, 2745–2753 (2008). This paper describes the effect of DNA ligase IV knockout on immunoglobulin class switch recombination in a murine B cell line.
Masani, S., Han, L., Meek, K. & Yu, K. Redundant function of DNA ligase 1 and 3 in alternative end-joining during immunoglobulin class switch recombination. Proc. Natl Acad. Sci. USA 113, 1261–1266 (2016).
Sfeir, A. & Symington, L. S. Microhomology-mediated end joining: a back-up survival mechanism or dedicated pathway? Trends Biochem. Sci. 40, 701–714 (2015).
Makharashvili, N. et al. Catalytic and noncatalytic roles of the CtIP endonuclease in double-strand break end resection. Mol. Cell 54, 1022–1033 (2014).
Anand, R., Ranjha, L., Cannavo, E. & Cejka, P. Phosphorylated CtIP Functions as a co-factor of the MRE11-RAD50-NBS1 endonuclease in DNA end resection. Mol. Cell 64, 940–950 (2016).
Deshpande, R. A., Lee, J. H., Arora, S. & Paull, T. T. Nbs1 converts the human Mre11/Rad50 nuclease complex into an endo/exonuclease machine specific for protein-DNA adducts. Mol. Cell 64, 593–606 (2016).
Robert, I., Dantzer, F. & Reina-San-Martin, B. Parp1 facilitates alternative NHEJ, whereas Parp2 suppresses IgH/c-myc translocations during immunoglobulin class switch recombination. J. Exp. Med. 206, 1047–1056 (2009).
Lange, S. S., Takata, K. & Wood, R. D. DNA polymerases and cancer. Nat. Rev. Cancer 11, 96–110 (2011).
Yu, A. M. & McVey, M. Synthesis-dependent microhomology-mediated end joining accounts for multiple types of repair junctions. Nucleic Acids Res. 38, 5706–5717 (2010).
Lieber, M. R., Hesse, J. E., Mizuuchi, K. & Gellert, M. Lymphoid V(D)J recombination: nucleotide insertion at signal joints as well as coding joints. Proc. Natl Acad. Sci. USA 85, 8588–8592 (1988).
Lieber, M. R. Mechanisms of human lymphoid chromosomal translocations. Nat. Rev. Cancer 16, 387–398 (2016).
Murga Penas, E. M. et al. The t(14;18)(q32;q21)/IGH-MALT1 translocation in MALT lymphomas contains templated nucleotide insertions and a major breakpoint region similar to follicular and mantle cell lymphoma. Blood 115, 2214–2219 (2010).
Jaeger, U. et al. Follicular lymphomas BCL-2/IgH junctions contain templated nucleotide insertions: novel insights into the mechanism of t(14;18) translocation. Blood 95, 3520–3529 (2000).
Welzel, N. et al. Templated nucleotide addition and immunoglobulin JH-gene utilization in t(11;14) junctions: implications for the mechanism of translocation and the origin of mantle cell lymphoma. Cancer Res. 61, 1629–1636 (2001). References 113 and 114 were the first to describe template insertions in lymphoid chromosomal translocations.
Pan-Hammarstrom, Q. et al. Impact of DNA ligase IV on nonhomologous end joining pathways during class switch recombination in human cells. J. Exp. Med. 201, 189–194 (2005).
Ceccaldi, R., Rondinelli, B. & D'Andrea, A. D. Repair pathway choices and consequences at the double-strand break. Trends Cell Biol. 26, 52–64 (2016).
Bhargava, R., Carson, C. R., Lee, G. & Stark, J. M. Contribution of canonical nonhomologous end joining to chromosomal rearrangements is enhanced by ATM kinase deficiency. Proc. Natl Acad. Sci. USA 114, 728–733 (2017).
Symington, L. S. Mechanism and regulation of DNA end resection in eukaryotes. Crit. Rev. Biochem. Mol. Biol. 51, 195–212 (2016).
Mimitou, E. P. & Symington, L. S. Sae2, Exo1 and Sgs1 collaborate in DNA double-strand break processing. Nature 455, 770–774 (2008).
Zhu, Z., Chung, W. H., Shim, E. Y., Lee, S. E. & Ira, G. Sgs1 helicase and two nucleases Dna2 and Exo1 resect DNA double-strand break ends. Cell 134, 981–994 (2008). References 119 and 120 describe resection mechanisms for a-EJ and SSA in yeast.
Sung, P., Krejci, L., Van Komen, S. & Sehorn, M. G. Rad51 recombinase and recombination mediators. J. Biol. Chem. 278, 42729–42732 (2003).
Paques, F. & Haber, J. E. Two pathways for removal of nonhomologous DNA ends during double-strand break repair in S. cerevisiae. Mol. Cell. Biol. 17, 6765–6771 (1997).
Pannunzio, N. R., Manthey, G. M. & Bailis, A. M. RAD59 is required for efficient repair of simultaneous double-strand breaks resulting in translocations in Saccharomyces cerevisiae. DNA Repair (Amst.) 7, 788–800 (2008).
Ira, G. et al. DNA end resection, homologous recombination and DNA damage checkpoint activation require CDK1. Nature 431, 1011–1017 (2004).
Jazayeri, A. et al. ATM- and cell cycle-dependent regulation of ATR in response to DNA double-strand breaks. Nat. Cell Biol. 8, 37–45 (2006).
Huertas, P. & Jackson, S. P. Human CtIP mediates cell cycle control of DNA end resection and double strand break repair. J. Biol. Chem. 284, 9558–9565 (2009).
Chen, L., Nievera, C. J., Lee, A. Y. & Wu, X. Cell cycle-dependent complex formation of BRCA1. CtIP. MRN is important for DNA double-strand break repair. J. Biol. Chem. 283, 7713–7720 (2008).
Cannavo, E. & Cejka, P. Sae2 promotes dsDNA endonuclease activity within Mre11–Rad50–Xrs2 to resect DNA breaks. Nature 514, 122–125 (2014).
Chen, X. et al. Cell cycle regulation of DNA double-strand break end resection by Cdk1-dependent Dna2 phosphorylation. Nat. Struct. Mol. Biol. 18, 1015–1019 (2011).
Tomimatsu, N. et al. Phosphorylation of EXO1 by CDKs 1 and 2 regulates DNA end resection and repair pathway choice. Nat. Commun. 5, 3561 (2014).
Zhou, Y. et al. Regulation of the DNA damage response by DNA-PKcs inhibitory phosphorylation of ATM. Mol. Cell 65, 91–104 (2017).
Beucher, A. et al. ATM and Artemis promote homologous recombination of radiation-induced DNA double-strand breaks in G2. EMBO J. 28, 3413–3427 (2009).
Biehs, R. et al. DNA double-strand break resection occurs during non-homologous end joining in G1 but is distinct from resection during homologous recombination. Mol. Cell 65, 671–684 (2017).
Falzon, M., Fewell, J. & Kuff, E. L. EBP-80, a transcription factor closely resembling the human autoantigen Ku, recognizes single- to double-strand transitions in DNA. J. Biol. Chem. 268, 10546–10552 (1993).
Ghezraoui, H. et al. Chromosomal translocations in human cells are generated by canonical nonhomologous end-joining. Mol. Cell 55, 829–842 (2014).
Boboila, C. et al. Alternative end-joining catalyzes robust IgH locus deletions and translocations in the combined absence of ligase 4 and Ku70. Proc. Natl Acad. Sci. USA 107, 3034–3039 (2010). This is the clearest example of a-EJ in mice with the absence of Ku70 and ligase IV.
Gostissa, M., Alt, F. W. & Chiarle, R. Mechanisms that promote and suppress chromosomal translocations in lymphocytes. Annu. Rev. Immunol. 29, 319–350 (2011).
Chiarle, R. et al. Genome-wide translocation sequencing reveals mechanisms of chromosome breaks and rearrangements in B cells. Cell 147, 107–119 (2011).
de Villartay, J. P. Congenital defects in V(D)J recombination. Br. Med. Bull. 114, 157–167 (2015).
Li, L. et al. A founder mutation in Artemis, an SNM1-like protein, causes SCID in Athabascan-speaking Native Americans. J. Immunol. 168, 6323–6329 (2002).
Volk, T. et al. DCLRE1C (ARTEMIS) mutations causing phenotypes ranging from atypical severe combined immunodeficiency to mere antibody deficiency. Hum. Mol. Genet. 24, 7361–7372 (2015).
Woodbine, L., Gennery, A. R. & Jeggo, P. A. The clinical impact of deficiency in DNA non-homologous end-joining. DNA Repair (Amst.) 16, 84–96 (2014).
de Villartay, J. P. When natural mutants do not fit our expectations: the intriguing case of patients with XRCC4 mutations revealed by whole-exome sequencing. EMBO Mol. Med. 7, 862–864 (2015).
I. Jspeert, H. et al. XLF deficiency results in reduced N-nucleotide addition during V(D)J recombination. Blood 128, 650–659 (2016).
Burg, M.v.d. et al. A DNA-PKcs mutation in a radiosensitive T-B− SCID patient inhibits Artemis activation and nonhomologous end-joining. J. Clin. Invest. 119, 91–98 (2009).
Woodbine, L. et al. PRKDC mutations in a SCID patient with profound neurological abnormalities. J. Clin. Invest. 123, 2969–2980 (2013).
Mathieu, A. L. et al. PRKDC mutations associated with immunodeficiency, granuloma, and autoimmune regulator-dependent autoimmunity. J. Allergy Clin. Immunol. 135, 1578–1588.e5 (2015).
Carvalho, C. M. & Lupski, J. R. Mechanisms underlying structural variant formation in genomic disorders. Nat. Rev. Genet. 17, 224–238 (2016).
Abyzov, A. et al. Analysis of deletion breakpoints from 1,092 humans reveals details of mutation mechanisms. Nat. Commun. 6, 7256 (2015).
Stephens, P. J. et al. Massive genomic rearrangement acquired in a single catastrophic event during cancer development. Cell 144, 27–40 (2011).
Iles, N., Rulten, S., El-Khamisy, S. F. & Caldecott, K. W. APLF (C2orf13) is a novel human protein involved in the cellular response to chromosomal DNA strand breaks. Mol. Cell. Biol. 27, 3793–3803 (2007).
Eustermann, S. et al. Solution structures of the two PBZ domains from human APLF and their interaction with poly(ADP-ribose). Nat. Struct. Mol. Biol. 17, 241–243 (2010).
Li, G. Y. et al. Structure and identification of ADP-ribose recognition motifs of APLF and role in the DNA damage response. Proc. Natl Acad. Sci. USA 107, 9129–9134 (2010).
Mehrotra, P. V. et al. DNA repair factor APLF is a histone chaperone. Mol. Cell 41, 46–55 (2011).
Acknowledgements
The authors thank R. Mosteller for comments on the manuscript. Work in the authors' laboratory is supported by the US National Institutes of Health (NIH) (M.R.L.) and by the Japanese Ministry of Education, Culture, Sports, Science, and Technology (MEXT) (15H04323 to N.A.).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary information S1 (table)
Abundance of key proteins involved in nonhomologous DNA end joining (NHEJ), alternative end joining (a-EJ), single-strand annealing (SSA) and homologous recombination (HR). (PDF 399 kb)
Supplementary information S2 (figure)
The same DSB can be repaired in many different ways, depending on the order in which NHEJ proteins act. (PDF 200 kb)
Glossary
- V(D)J recombination
-
DNA recombination process during B or T lymphocyte activation in which the antigen receptors variable domain exons are assembled from sub-exonic segments called V, D and J to ultimately generate an immunoglobulin gene or T cell receptor, respectively.
- Immunoglobulin heavy chain class switch recombination
-
The DNA recombination process by which the immunoglobulin heavy chain isotype is changed from producing IgM to producing IgG, IgA or IgE.
- Microhomology
-
One or more base pairs of complementarity at the two DNA ends of a break.
- FAT domain
-
FRAP (FKBP12-rapamycin-associated protein), ATM (ataxia telangiectasia mutated), TRRAP (transformation/transcription domain-associated protein) domain. A structural domain found in phosphatidylinositol 3-kinase-like kinase family members.
- Pol X family polymerases
-
Subfamily of DNA polymerases; based on homology it includes Pol β, Pol μ, Pol λ and terminal deoxynucleotidyltransferase (TdT).
- BRCA1 C terminus
-
(BRCT). Protein domain of approximately 100 aa that binds to phosphoproteins that are often involved in the DNA damage response.
- DNA end breathing
-
Break of the hydrogen bonds between one or more base pairs in the anti-parallel strands of the DNA duplex break.
- Templated insertions
-
Nucleotide additions at a double-strand break repair junction that seem to be direct or inverted repeat copies derived from either strand of either of the two DNA ends.
- Chromothripsis
-
Shattering of chromosomal regions followed by random repair of the DNA fragments in some human neoplasms and inherited disorders.
Rights and permissions
About this article
Cite this article
Chang, H., Pannunzio, N., Adachi, N. et al. Non-homologous DNA end joining and alternative pathways to double-strand break repair. Nat Rev Mol Cell Biol 18, 495–506 (2017). https://doi.org/10.1038/nrm.2017.48
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrm.2017.48
This article is cited by
-
Improving CRISPR–Cas9 directed faithful transgene integration outcomes by reducing unwanted random DNA integration
Journal of Biomedical Science (2024)
-
GCN5 mediates DNA-PKcs crotonylation for DNA double-strand break repair and determining cancer radiosensitivity
British Journal of Cancer (2024)
-
Macroscopic inhibition of DNA damage repair pathways by targeting AP-2α with LEI110 eradicates hepatocellular carcinoma
Communications Biology (2024)
-
Dbf4-dependent kinase promotes cell cycle controlled resection of DNA double-strand breaks and repair by homologous recombination
Nature Communications (2024)
-
Precise genome-editing in human diseases: mechanisms, strategies and applications
Signal Transduction and Targeted Therapy (2024)