Key Points
-
HIV controllers are a small group of HIV-1-infected patients who maintain undetectable or minimal levels of HIV-1 replication in the absence of antiretroviral therapy.
-
Many elite controllers harbour replication-competent viruses, and several studies have consistently failed to identify specific viral sequence abnormalities in these patients, which suggests that the controller phenotype is primarily achieved by host factors.
-
Genome-wide association studies have shown that genetic variations associated with HIV-1 immune control are exclusively located in the human HLA class I locus. Such genetic variations are most frequently detectable in the HLA-B binding pocket, but can also be found in selected areas encoding non-pocket elements of HLA-B or HLA-C.
-
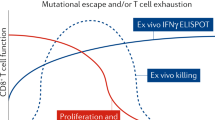
Highly-potent HIV-1-specific CD8+ T cells seem to be the backbone of antiviral immune defence in many — but not all — elite controllers. These cells are highly effective at restricting HIV-1 replication in in vitro inhibition assays and they have functional and phenotypic characteristics that distinguish them from HIV-1-specific T cells from individuals with progressive HIV-1 infection.
-
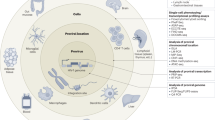
Innate immune defence mechanisms are likely to modulate immune activity in HIV-1 controllers. In particular, dendritic cells, γδ T cells and the cell-intrinsic restriction of viral replication seem to support antiviral immune activities in HIV-1 controllers.
-
Elite controllers provide living evidence that the human immune system is able to effectively control HIV-1 replication. A better understanding of the mechanisms involved in HIV-1 immune control in these patients might enable the design of clinical strategies to improve the immune response to HIV-1 in broader patient populations.
Abstract
Untreated HIV-1 infection typically progresses to AIDS within 10 years, but less than 1% of infected individuals remain healthy and have normal CD4+ T cell counts and undetectable viral loads; some individuals have remained this way for 35 years and counting. Through a combination of large population studies of cohorts of these 'HIV-1 controllers' and detailed studies of individual patients, a heterogeneous picture has emerged regarding the basis for this remarkable resistance to AIDS progression. In this Review, we highlight the host genetic factors, the viral genetic factors and the immunological factors that are associated with the controller phenotype, we discuss emerging methodological approaches that could facilitate a better understanding of spontaneous HIV-1 immune control in the future, and we delineate implications for a 'functional cure' of HIV-1 infection.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Virgin, H. W., Wherry, E. J. & Ahmed, R. Redefining chronic viral infection. Cell 138, 30–50 (2009).
Ahmed, R., Salmi, A., Butler, L. D., Chiller, J. M. & Oldstone, M. B. Selection of genetic variants of lymphocytic choriomeningitis virus in spleens of persistently infected mice. Role in suppression of cytotoxic T lymphocyte response and viral persistence. J. Exp. Med. 160, 521–540 (1984).
Ahmed, R. et al. Molecular basis of organ-specific selection of viral variants during chronic infection. J. Virol. 65, 4242–4247 (1991).
World Health Organization. Progress report 2011: Global HIV/AIDS response. World Health Organization [online], (2011).
Deeks, S. G. & Walker, B. D. Human immunodeficiency virus controllers: mechanisms of durable virus control in the absence of antiretroviral therapy. Immunity 27, 406–416 (2007).
Pereyra, F. et al. The major genetic determinants of HIV-1 control affect HLA class I peptide presentation. Science 330, 1551–1557 (2010). This paper describes a large GWAS in more than 2,000 HIV-1 controllers and progressors. The main finding was that genetic determinants of HIV-1 immune control are exclusively located in the HLA class I locus.
Mariani, R. et al. High frequency of defective nef alleles in a long-term survivor with nonprogressive human immunodeficiency virus type 1 infection. J. Virol. 70, 7752–7764 (1996).
Michael, N. L. et al. Defective accessory genes in a human immunodeficiency virus type 1-infected long-term survivor lacking recoverable virus. J. Virol. 69, 4228–4236 (1995).
Alexander, L. et al. Unusual polymorphisms in human immunodeficiency virus type 1 associated with nonprogressive infection. J. Virol. 74, 4361–4376 (2000).
Miura, T. et al. Genetic characterization of human immunodeficiency virus type 1 in elite controllers: lack of gross genetic defects or common amino acid changes. J. Virol. 82, 8422–8430 (2008).
Julg, B. et al. Infrequent recovery of HIV from but robust exogenous infection of activated CD4+ T cells in HIV elite controllers. Clin. Infect. Dis. 51, 233–238 (2010).
Lamine, A. et al. Replication-competent HIV strains infect HIV controllers despite undetectable viremia (ANRS EP36 study). AIDS 21, 1043–1045 (2007).
Blankson, J. N. et al. Isolation and characterization of replication-competent human immunodeficiency virus type 1 from a subset of elite suppressors. J. Virol. 81, 2508–2518 (2007).
Bailey, J. R. et al. Transmission of human immunodeficiency virus type 1 from a patient who developed AIDS to an elite suppressor. J. Virol. 82, 7395–7410 (2008).
Buckheit, R. W. et al. Host factors dictate control of viral replication in two HIV-1 controller/chronic progressor transmission pairs. Nature Commun. 3, 716 (2012).
Bailey, J. R., Williams, T. M., Siliciano, R. F. & Blankson, J. N. Maintenance of viral suppression in HIV-1-infected HLA-B*57+ elite suppressors despite CTL escape mutations. J. Exp. Med. 203, 1357–1369 (2006).
Bailey, J. R., Brennan, T. P., O'Connell, K. A., Siliciano, R. F. & Blankson, J. N. Evidence of CD8+ T-cell-mediated selective pressure on human immunodeficiency virus type 1 nef in HLA-B*57+ elite suppressors. J. Virol. 83, 88–97 (2009).
Miura, T. et al. HLA-associated viral mutations are common in human immunodeficiency virus type 1 elite controllers. J. Virol. 83, 3407–3412 (2009).
Miura, T. et al. HLA-associated alterations in replication capacity of chimeric NL4-3 viruses carrying gag-protease from elite controllers of human immunodeficiency virus type 1. J. Virol. 83, 140–149 (2009).
Brumme, Z. L. et al. Reduced replication capacity of NL4-3 recombinant viruses encoding reverse transcriptase-integrase sequences from HIV-1 elite controllers. J. Acquir. Immune Def. Syndr. 56, 100–108 (2011).
Miura, T. et al. Impaired replication capacity of acute/early viruses in persons who become HIV controllers. J. Virol. 84, 7581–7591 (2010).
Kaslow, R. A. et al. Influence of combinations of human major histocompatibility complex genes on the course of HIV-1 infection. Nature Med. 2, 405–411 (1996).
Carrington, M. et al. HLA and HIV-1: heterozygote advantage and B*35-Cw*04 disadvantage. Science 283, 1748–1752 (1999).
Pereyra, F. et al. Genetic and immunologic heterogeneity among persons who control HIV infection in the absence of therapy. J. Infect. Dis. 197, 563–571 (2008).
Migueles, S. A. et al. HLA B*5701 is highly associated with restriction of virus replication in a subgroup of HIV-infected long term nonprogressors. Proc. Natl Acad. Sci. USA 97, 2709–2714 (2000).
Fellay, J. et al. A whole-genome association study of major determinants for host control of HIV-1. Science 317, 944–947 (2007).
Pelak, K. et al. Host determinants of HIV-1 control in African Americans. J. Infect. Dis. 201, 1141–1149 (2010).
Limou, S. et al. Genomewide association study of an AIDS-nonprogression cohort emphasizes the role played by HLA genes (ANRS Genomewide Association Study 02). J. Infect. Dis. 199, 419–426 (2009).
Sobieszczyk, M. E., Lingappa, J. R. & McElrath, M. J. Host genetic polymorphisms associated with innate immune factors and HIV-1. Curr. Opin. HIV AIDS 6, 427–434 (2011).
Chatterjee, K. Host genetic factors in susceptibility to HIV-1 infection and progression to AIDS. J. Genet. 89, 109–116 (2010).
Kosmrlj, A. et al. Effects of thymic selection of the T-cell repertoire on HLA class I-associated control of HIV infection. Nature 465, 350–354 (2010).
Chessman, D. et al. Human leukocyte antigen class I-restricted activation of CD8+ T cells provides the immunogenetic basis of a systemic drug hypersensitivity. Immunity 28, 822–832 (2008).
O'Brien, S. J., Gao, X. & Carrington, M. HLA and AIDS: a cautionary tale. Trends Mol. Med. 7, 379–381 (2001).
Chen, H. et al. TCR clonotypes modulate the protective effect of HLA class I molecules in HIV-1 infection. Nature Immunol. 13, 691–700 (2012).
Prentice, H. A. et al. HLA-B*57 versus HLA-B*81 in HIV-1 infection: slow and steady wins the race? J. Virol. 87, 4043–4051 (2013).
McLaren, P. J. et al. Fine-mapping classical HLA variation associated with durable host control of HIV-1 infection in African Americans. Hum. Mol. Genet. 21, 4334–4347 (2012).
Thomas, R. et al. HLA-C cell surface expression and control of HIV/AIDS correlate with a variant upstream of HLA-C. Nature Genet. 41, 1290–1294 (2009).
Kulkarni, S. et al. Differential microRNA regulation of HLA-C expression and its association with HIV control. Nature 472, 495–498 (2011).
Goulder, P. J. & Walker, B. D. HIV and HLA class I: an evolving relationship. Immunity 37, 426–440 (2012).
Herbeck, J. T. et al. Multistage genomewide association study identifies a locus at 1q41 associated with rate of HIV-1 disease progression to clinical AIDS. J. Infect. Dis. 201, 618–626 (2010).
Martin, M. P. et al. Epistatic interaction between KIR3DS1 and HLA-B delays the progression to AIDS. Nature Genet. 31, 429–434 (2002).
Martin, M. P. et al. Innate partnership of HLA-B and KIR3DL1 subtypes against HIV-1. Nature Genet. 39, 733–740 (2007).
O'Connell, K. A., Han, Y., Williams, T. M., Siliciano, R. F. & Blankson, J. N. Role of natural killer cells in a cohort of elite suppressors: low frequency of the protective KIR3DS1 allele and limited inhibition of human immunodeficiency virus type 1 replication in vitro. J. Virol. 83, 5028–5034 (2009).
Huang, J. et al. HLA-B*35-Px-mediated acceleration of HIV-1 infection by increased inhibitory immunoregulatory impulses. J. Exp. Med. 206, 2959–2966 (2009).
Jones, D. C. et al. HLA class I allelic sequence and conformation regulate leukocyte Ig-like receptor binding. J. Immunol. 186, 2990–2997 (2011).
Williams, K. et al. Epigenetic regulation of telomerase expression in HIV-1-specific CD8+ T cells. AIDS 24, 1964–1966 (2010).
van Grevenynghe, J. et al. Transcription factor FOXO3a controls the persistence of memory CD4+ T cells during HIV infection. Nature Med. 14, 266–274 (2008).
Saez-Cirion, A. et al. HIV controllers exhibit potent CD8 T cell capacity to suppress HIV infection ex vivo and peculiar cytotoxic T lymphocyte activation phenotype. Proc. Natl Acad. Sci. USA 104, 6776–6781 (2007).
Saez-Cirion, A. et al. Heterogeneity in HIV suppression by CD8 T cells from HIV controllers: association with Gag-specific CD8 T cell responses. J. Immunol. 182, 7828–7837 (2009).
Quigley, M. et al. Transcriptional analysis of HIV-specific CD8+ T cells shows that PD-1 inhibits T cell function by upregulating BATF. Nature Med. 16, 1147–1151 (2010).
Hersperger, A. R. et al. Perforin expression directly ex vivo by HIV-specific CD8 T-cells is a correlate of HIV elite control. PLoS Pathog. 6, e1000917 (2010).
Migueles, S. A. et al. Lytic granule loading of CD8+ T cells is required for HIV-infected cell elimination associated with immune control. Immunity 29, 1009–1021 (2008). This study identifies the enhanced cytotoxic properties of HIV-1-specific CD8+ T cells as a functional characteristic that correlates with spontaneous HIV-1 immune control.
Hersperger, A. R. et al. Increased HIV-specific CD8+ T-cell cytotoxic potential in HIV elite controllers is associated with T-bet expression. Blood 117, 3799–3808 (2011).
Migueles, S. A. et al. HIV-specific CD8+ T cell proliferation is coupled to perforin expression and is maintained in nonprogressors. Nature Immunol. 3, 1061–1068 (2002).
Zimmerli, S. C. et al. HIV-1-specific IFN-α/IL-2-secreting CD8 T cells support CD4-independent proliferation of HIV-1-specific CD8 T cells. Proc. Natl Acad. Sci. USA 102, 7239–7244 (2005).
Betts, M. R. et al. HIV nonprogressors preferentially maintain highly functional HIV-specific CD8+ T cells. Blood 107, 4781–4789 (2006).
Ndhlovu, Z. M. et al. Elite controllers with low to absent effector CD8+ T cell responses maintain highly functional, broadly directed central memory responses. J. Virol. 86, 6959–6969 (2012).
Ferre, A. L. et al. Mucosal immune responses to HIV-1 in elite controllers: a potential correlate of immune control. Blood 113, 3978–3989 (2009). This paper analyses the phenotypic and functional characteristics of cellular immune responses to HIV-1 in rectal mucosal tissues and shows a selective enrichment of highly functional HIV-1-specific CD8+ T cells in intestinal mucosal tissues from HIV-1 controllers.
Elahi, S. et al. Protective HIV-specific CD8+ T cells evade Treg cell suppression. Nature Med. 17, 989–995 (2011). This work proposes that HLA-B*57-restricted HIV-1-specific CD8+ T cells are less susceptible to the inhibitory effects of T Reg cells than are CD8+ T cells restricted by other HLA alleles. This effect might contribute to the delayed HIV-1 disease progression that is frequently observed in HLA-B*57-expressing individuals.
Dahirel, V. et al. Coordinate linkage of HIV evolution reveals regions of immunological vulnerability. Proc. Natl Acad. Sci. USA 108, 11530–11535 (2011).
Kiepiela, P. et al. CD8+ T-cell responses to different HIV proteins have discordant associations with viral load. Nature Med. 13, 46–53 (2007).
Zuniga, R. et al. Relative dominance of Gag p24-specific cytotoxic T lymphocytes is associated with human immunodeficiency virus control. J. Virol. 80, 3122–3125 (2006).
Edwards, B. H. et al. Magnitude of functional CD8+ T-cell responses to the gag protein of human immunodeficiency virus type 1 correlates inversely with viral load in plasma. J. Virol. 76, 2298–2305 (2002).
Novitsky, V. A. et al. Interactive association of proviral load and IFN-γ-secreting T cell responses in HIV-1C infection. Virology 349, 142–155 (2006).
Mudd, P. A. et al. Vaccine-induced CD8+ T cells control AIDS virus replication. Nature 491, 129–133 (2012).
Emu, B. et al. HLA class I-restricted T-cell responses may contribute to the control of human immunodeficiency virus infection, but such responses are not always necessary for long-term virus control. J. Virol. 82, 5398–5407 (2008).
Brennan, C. A. et al. Early HLA-B*57-restricted CD8+ T lymphocyte responses predict HIV-1 disease progression. J. Virol. 86, 10505–10516 (2012).
Janbazian, L. et al. Clonotype and repertoire changes drive the functional improvement of HIV-specific CD8 T cell populations under conditions of limited antigenic stimulation. J. Immunol. 188, 1156–1167 (2012).
Cubas, R. A. et al. Inadequate T follicular cell help impairs B cell immunity during HIV infection. Nature Med. 19, 494–499 (2013).
Vingert, B. et al. HIV controller CD4+ T cells respond to minimal amounts of Gag antigen due to high TCR avidity. PLoS Pathog. 6, e1000780 (2010).
Chevalier, M. F. et al. HIV-1-specific interleukin-21+ CD4+ T cell responses contribute to durable viral control through the modulation of HIV-specific CD8+ T cell function. J. Virol. 85, 733–741 (2011).
Lichterfeld, M. et al. Loss of HIV-1-specific CD8+ T cell proliferation after acute HIV-1 infection and restoration by vaccine-induced HIV-1-specific CD4+ T cells. J. Exp. Med. 200, 701–712 (2004).
Tilton, J. C. et al. Changes in paracrine interleukin-2 requirement, CCR7 expression, frequency, and cytokine secretion of human immunodeficiency virus-specific CD4+ T cells are a consequence of antigen load. J. Virol. 81, 2713–2725 (2007).
Soghoian, D. Z. et al. HIV-specific cytolytic CD4 T cell responses during acute HIV infection predict disease outcome. Sci. Transl. Med. 4, 123ra25 (2012).
Ferre, A. L. et al. HIV controllers with HLA-DRB1*13 and HLA-DQB1*06 alleles have strong, polyfunctional mucosal CD4+ T-cell responses. J. Virol. 84, 11020–11029 (2010).
Kaufmann, D. E. et al. Comprehensive analysis of human immunodeficiency virus type 1-specific CD4 responses reveals marked immunodominance of gag and nef and the presence of broadly recognized peptides. J. Virol. 78, 4463–4477 (2004).
Ranasinghe, S. et al. Association of HLA-DRB1-restricted CD4 T cell responses with HIV immune control. Nature Med. (in the press).
Doria-Rose, N. A. et al. Breadth of human immunodeficiency virus-specific neutralizing activity in sera: clustering analysis and association with clinical variables. J. Virol. 84, 1631–1636 (2010).
Lambotte, O. et al. Heterogeneous neutralizing antibody and antibody-dependent cell cytotoxicity responses in HIV-1 elite controllers. AIDS 23, 897–906 (2009).
Bailey, J. R. et al. Neutralizing antibodies do not mediate suppression of human immunodeficiency virus type 1 in elite suppressors or selection of plasma virus variants in patients on highly active antiretroviral therapy. J. Virol. 80, 4758–4770 (2006).
Deeks, S. G. et al. Neutralizing antibody responses against autologous and heterologous viruses in acute versus chronic human immunodeficiency virus (HIV) infection: evidence for a constraint on the ability of HIV to completely evade neutralizing antibody responses. J. Virol. 80, 6155–6164 (2006).
Sather, D. N. et al. Factors associated with the development of cross-reactive neutralizing antibodies during human immunodeficiency virus type 1 infection. J. Virol. 83, 757–769 (2009).
Euler, Z. et al. Cross-reactive neutralizing humoral immunity does not protect from HIV type 1 disease progression. J. Infect. Dis. 201, 1045–1053 (2010).
Mahalanabis, M. et al. Continuous viral escape and selection by autologous neutralizing antibodies in drug-naive human immunodeficiency virus controllers. J. Virol. 83, 662–672 (2009).
Scheid, J. F. et al. Broad diversity of neutralizing antibodies isolated from memory B cells in HIV-infected individuals. Nature 458, 636–640 (2009).
Hessell, A. J. et al. Fc receptor but not complement binding is important in antibody protection against HIV. Nature 449, 101–104 (2007).
Brocca-Cofano, E. et al. Vaccine-elicited SIV and HIV envelope-specific IgA and IgG memory B cells in rhesus macaque peripheral blood correlate with functional antibody responses and reduced viremia. Vaccine 29, 3310–3319 (2011).
Smalls-Mantey, A. et al. Antibody-dependent cellular cytotoxicity against primary HIV-infected CD4+ T cells is directly associated with the magnitude of surface IgG binding. J. Virol. 86, 8672–8680 (2012).
Huang, J. et al. Leukocyte immunoglobulin-like receptors maintain unique antigen-presenting properties of circulating myeloid dendritic cells in HIV-1-infected elite controllers. J. Virol. 84, 9463–9471 (2010).
Barblu, L. et al. Plasmacytoid dendritic cells from HIV controllers produce IFN-α and differentiate into functional killer pDC under HIV activation. J. Infect. Dis. 206, 790–801 (2012).
Machmach, K. et al. Plasmacytoid dendritic cells reduce HIV production in elite controllers. J. Virol. 86, 4245–4252 (2012).
Altfeld, M., Fadda, L., Frleta, D. & Bhardwaj, N. DCs and NK cells: critical effectors in the immune response to HIV-1. Nature Rev. Immunol. 11, 176–186 (2011).
Alter, G. et al. HIV-1 adaptation to NK-cell-mediated immune pressure. Nature 476, 96–100 (2011).
Barker, E., Martinson, J., Brooks, C., Landay, A. & Deeks, S. Dysfunctional natural killer cells, in vivo, are governed by HIV viremia regardless of whether the infected individual is on antiretroviral therapy. AIDS 21, 2363–2365 (2007).
Vieillard, V., Fausther-Bovendo, H., Samri, A. & Debre, P. Specific phenotypic and functional features of natural killer cells from HIV-infected long-term nonprogressors and HIV controllers. J. Acquir. Immune Def. Syndr. 53, 564–573 (2010).
Riedel, D. J. et al. Natural viral suppressors of HIV-1 have a unique capacity to maintain γδ T cells. AIDS 23, 1955–1964 (2009).
O'Connell, K. A., Rabi, S. A., Siliciano, R. F. & Blankson, J. N. CD4+ T cells from elite suppressors are more susceptible to HIV-1 but produce fewer virions than cells from chronic progressors. Proc. Natl Acad. Sci. USA 108, E689–E698 (2011).
Chen, H. et al. CD4+ T cells from elite controllers resist HIV-1 infection by selective upregulation of p21. J. Clin. Invest. 121, 1549–1560 (2011).
Saez-Cirion, A. et al. Restriction of HIV-1 replication in macrophages and CD4+ T cells from HIV controllers. Blood 118, 955–964 (2011). References 98 and 99 show a reduced susceptibility of CD4+ T cells from elite controllers to HIV-1 infection through the inhibition of early viral replication steps.
Elahi, S., Niki, T., Hirashima, M. & Horton, H. Galectin-9 binding to Tim-3 renders activated human CD4+ T cells less susceptible to HIV-1 infection. Blood 119, 4192–4204 (2012).
Goldfeld, A. E., Birch-Limberger, K., Schooley, R. T. & Walker, B. D. HIV-1 infection does not induce tumor necrosis factor-α or interferon-β gene transcription. J. Acquir. Immune Def. Syndr. 4, 41–47 (1991).
Manel, N. et al. A cryptic sensor for HIV-1 activates antiviral innate immunity in dendritic cells. Nature 467, 214–217 (2010).
Yan, N., Regalado-Magdos, A. D., Stiggelbout, B., Lee-Kirsch, M. A. & Lieberman, J. The cytosolic exonuclease TREX1 inhibits the innate immune response to human immunodeficiency virus type 1. Nature Immunol. 11, 1005–1013 (2010).
Card, C. M. et al. Decreased immune activation in resistance to HIV-1 infection is associated with an elevated frequency of CD4+CD25+FOXP3+ regulatory T cells. J. Infect. Dis. 199, 1318–1322 (2009).
Hunt, P. W. et al. Relationship between T cell activation and CD4+ T cell count in HIV-seropositive individuals with undetectable plasma HIV RNA levels in the absence of therapy. J. Infect. Dis. 197, 126–133 (2008).
Kamya, P. et al. T cell activation does not drive CD4 decline in longitudinally followed HIV-infected elite controllers. AIDS Res. Ther. 8, 20 (2011).
Hatano, H. et al. Evidence for persistent low-level viremia in individuals who control human immunodeficiency virus in the absence of antiretroviral therapy. J. Virol. 83, 329–335 (2009).
Pereyra, F. et al. Persistent low-level viremia in HIV-1 elite controllers and relationship to immunologic parameters. J. Infect. Dis. 200, 984–990 (2009).
Mens, H. et al. HIV-1 continues to replicate and evolve in patients with natural control of HIV infection. J. Virol. 84, 12971–12981 (2010).
Brandt, L. et al. Low level of regulatory T cells and maintenance of balance between regulatory T cells and TH17 cells in HIV-1-infected elite controllers. J. Acquir. Immune Def. Syndr. 57, 101–108 (2011).
Hunt, P. W. et al. A low T regulatory cell response may contribute to both viral control and generalized immune activation in HIV controllers. PLoS ONE 6, e15924 (2011).
Favre, D. et al. Tryptophan catabolism by indoleamine 2,3-dioxygenase 1 alters the balance of TH17 to regulatory T cells in HIV disease. Sci. Transl. Med. 2, 32ra36 (2010).
Vigneault, F. et al. Transcriptional profiling of CD4 T cells identifies distinct subgroups of HIV-1 elite controllers. J. Virol. 85, 3015–3019 (2011).
Mendoza, D. et al. Comprehensive analysis of unique cases with extraordinary control over HIV replication. Blood 119, 4645–4655 (2012).
Peng, X. et al. Virus-host interactions: from systems biology to translational research. Curr. Opin. Microbiol. 12, 432–438 (2009).
Aderem, A. et al. A systems biology approach to infectious disease research: innovating the pathogen–host research paradigm. MBio 2, e00325–e00310 (2011).
Potti, A. et al. A genomic strategy to refine prognosis in early-stage non-small-cell lung cancer. N. Engl. J. Med. 355, 570–580 (2006).
Chaussabel, D. et al. A modular analysis framework for blood genomics studies: application to systemic lupus erythematosus. Immunity 29, 150–164 (2008).
Martinez-Llordella, M. et al. Using transcriptional profiling to develop a diagnostic test of operational tolerance in liver transplant recipients. J. Clin. Invest. 118, 2845–2857 (2008).
Querec, T. D. et al. Systems biology approach predicts immunogenicity of the yellow fever vaccine in humans. Nature Immunol. 10, 116–125 (2009).
Nakaya, H. I., Li, S. & Pulendran, B. Systems vaccinology: learning to compute the behavior of vaccine induced immunity. Wiley Interdiscip. Rev. Syst. Biol. Med. 4, 193–205 (2011).
Nakaya, H. I. et al. Systems biology of vaccination for seasonal influenza in humans. Nature Immunol. 12, 786–795 (2011).
Richman, D. D. et al. The challenge of finding a cure for HIV infection. Science 323, 1304–1307 (2009).
Deeks, S. G. et al. Towards an HIV cure: a global scientific strategy. Nature Rev. Immunol. 12, 607–614 (2012).
Lifson, A. R. et al. Long-term human immunodeficiency virus infection in asymptomatic homosexual and bisexual men with normal CD4+ lymphocyte counts: immunologic and virologic characteristics. J. Infect. Dis. 163, 959–965 (1991).
Sauce, D. et al. HIV disease progression despite suppression of viral replication is associated with exhaustion of lymphopoiesis. Blood 117, 5142–5151 (2011).
Yang, Y. et al. CD4 T-cell regeneration in HIV-1 elite controllers. AIDS 26, 701–706 (2012).
Pereyra, F. et al. Increased coronary atherosclerosis and immune activation in HIV-1 elite controllers. AIDS 26, 2409–2412 (2012).
Hatano, H. et al. Prospective ART of asymptomatic HIV-1 controllers. Abstract 75LB. 20th Conference on Retroviruses and Opportunistic Infections [online], (2013).
Graf, E. H. et al. Elite suppressors harbor low levels of integrated HIV DNA and high levels of 2-LTR circular HIV DNA compared to HIV+ patients on and off HAART. PLoS Pathog. 7, e1001300 (2011).
Shan, L. et al. Stimulation of HIV-1-specific cytolytic T lymphocytes facilitates elimination of latent viral reservoir after virus reactivation. Immunity 36, 491–501 (2012). This paper shows that highly functional HIV-1-specific CD8+ T cells might contribute to the spontaneous control of HIV-1 replication. The study suggests that HIV-1-specific CTLs from HIV-1 controllers might also be able to effectively kill latent HIV-1-infected CD4+ T cells in which active HIV-1 replication is reactivated by pharmacological agents.
Loffredo, J. T. et al. Mamu-B*08-positive macaques control simian immunodeficiency virus replication. J. Virol. 81, 8827–8832 (2007).
Loffredo, J. T. et al. Patterns of CD8+ immunodominance may influence the ability of Mamu-B*08-positive macaques to naturally control simian immunodeficiency virus SIVmac239 replication. J. Virol. 82, 1723–1738 (2008).
Altfeld, M. et al. Influence of HLA-B57 on clinical presentation and viral control during acute HIV-1 infection. AIDS 17, 2581–2591 (2003).
Brumme, Z. L. et al. Marked epitope- and allele-specific differences in rates of mutation in human immunodeficiency type 1 (HIV-1) Gag, Pol, and Nef cytotoxic T-lymphocyte epitopes in acute/early HIV-1 infection. J. Virol. 82, 9216–9227 (2008).
Valentine, L. E. et al. Infection with “escaped” virus variants impairs control of simian immunodeficiency virus SIVmac239 replication in Mamu-B*08-positive macaques. J. Virol. 83, 11514–11527 (2009).
Rosenberg, E. S. et al. Immune control of HIV-1 after early treatment of acute infection. Nature 407, 523–526 (2000).
Kaufmann, D. E. et al. Limited durability of viral control following treated acute HIV infection. PLoS Med. 1, e36 (2004).
Sáez-Cirión, A. et al. Post-treatment HIV-1 controllers with a long-term virological remission after the interruption of early initiated antiretroviral therapy ANRS VISCONTI Study. PLoS Pathog. 9, e1003211 (2013). This provocative study describes a series of HIV-1-infected patients who were able to spontaneously control HIV-1 replication after a long period of antiretroviral treatment that was started during primary HIV-1 infection. Interestingly, these 'post-treatment' controllers did not have high frequencies of HIV-1-specific CD8+ T cells and they did not have enriched expression of protective HLA class I alleles.
Lodi, S. et al. Immunovirologic control 24 months after interruption of antiretroviral therapy initiated close to HIV seroconversion. Arch. Intern. Med. 172, 1252–1255 (2012).
Acknowledgements
B.D.W. and X.G.Y. are supported by: the US National Institutes of Health grants AI098484, AI078799 and AI089339 (to X.G.Y.) as well as AI30914 and AI67073 (to B.D.W.); the Bill and Melinda Gates Foundation; the Mark and Lisa Schwartz Foundation; and the Phillip T. and Susan M. Ragon Foundation.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Related links
PowerPoint slides
Glossary
- CD4+ T cell counts
-
A normal CD4+ T cell count is 1,000 cells per microlitre (μl) of plasma, with a range of 600 to 1,400 cells per μl. The count falls during primary infection with HIV-1, then returns nearly to, or to lower than, normal levels. It then slowly falls, taking many years to reach the level of 200 cells per μl that characterizes the development of AIDS.
- HIV-1 controllers
-
HIV-1-infected patients who spontaneously maintain very low levels of viral replication in the absence of antiretroviral therapy.
- Elite controllers
-
HIV-1-infected patients who have undetectable levels of viral replication in the absence of antiretroviral therapy.
- Genome-wide association studies
-
(GWASs). Studies that assess upwards of one million single-nucleotide polymorphisms in the human genome for associations with disease outcomes. As such, they require very large numbers of individuals; the number of individuals that are required depends on the strength of the associations that are being investigated.
- Central memory T cells
-
Memory T cells that express L-selectin and CC-chemokine receptor 7 (CCR7) and that have the capacity to traffic from the blood to the secondary lymphoid organs. They have a nonpolarized differentiation state: they secrete interleukin-2 but not interferon-γ or interleukin-4. However, upon restimulation, they rapidly differentiate into cytokine-producing effector cells.
- Highly active antiretroviral therapy
-
(HAART). An aggressive combination therapy against HIV-1 infection that typically includes three or more protease and reverse- transcriptase inhibitors.
Rights and permissions
About this article
Cite this article
Walker, B., Yu, X. Unravelling the mechanisms of durable control of HIV-1. Nat Rev Immunol 13, 487–498 (2013). https://doi.org/10.1038/nri3478
Published:
Issue Date:
DOI: https://doi.org/10.1038/nri3478
This article is cited by
-
Reduced CCR5 expression among Uganda HIV controllers
Retrovirology (2023)
-
Virus-immune dynamics determined by prey-predator interaction network and epistasis in viral fitness landscape
Journal of Mathematical Biology (2023)
-
Global transcriptomic characterization of T cells in individuals with chronic HIV-1 infection
Cell Discovery (2022)
-
The RIO trial: rationale, design, and the role of community involvement in a randomised placebo-controlled trial of antiretroviral therapy plus dual long-acting HIV-specific broadly neutralising antibodies (bNAbs) in participants diagnosed with recent HIV infection—study protocol for a two-stage randomised phase II trial
Trials (2022)
-
Low CCR5 expression protects HIV-specific CD4+ T cells of elite controllers from viral entry
Nature Communications (2022)