Key Points
-
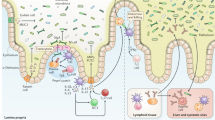
Germ-free mice have many immunological defects in the intestine, including smaller mesenteric lymph nodes and Peyer's patches, decreased numbers of interleukin-17 (IL-17)-producing T helper 17 (TH17) cells and defects in regulatory T (TReg) cells. In addition, germ-free mice have impaired immune responses to certain pathogens, including Shigella and Listeria species. These findings indicate that the intestinal microbiota might influence both pro- and anti-inflammatory responses.
-
Intestinal homeostasis depends on maintaining a proper balance between pro- and anti-inflammatory pathways that are mediated by TH17 and TReg cells, respectively. Improper regulation of inflammatory pathways can lead to diseases such as inflammatory bowel disease (IBD). IBD results from the breakdown in immune tolerance to gut bacteria. Indeed, spontaneous disease does not occur in many mouse models of experimental colitis, including IL-2- and IL-10-deficient mice, when raised under germ-free conditions.
-
Susceptibility to IBD is influenced by many factors, including genetic and dietary factors. However, recent studies have shown that the intestinal microbiota could be an important factor in driving disease.
-
Similar to observations made in animal models, changes in the microbiota of humans have been implicated in disease. Numerous studies have revealed a significant alteration in the microbiota of patients with IBD compared with healthy individuals, although whether changes in the intestinal microbiota are the cause or effect of disease still needs to be determined.
-
Several species of bacteria that peacefully reside in the intestine have been shown to have a protective role during IBD. These bacteria are referred to as probiotic bacteria and include species such as Lactobacillus casei and Bifidobacteria longum. The mechanisms by which these bacteria protect from disease are thought to involve modulation of TReg cell responses. A product of B. fragilis, polysaccharide A, elicits IL-10 production by CD4+ T cells and mediates protection from IBD, demonstrating that symbiotic intestinal bacteria have developed strategies to influence the host immune system.
-
Does harbouring certain strains of bacteria predispose an individual to disease or protect from it? As symbiotic bacteria seem to have evolved mechanisms to promote protection from potentially pathogenic bacteria in the microbiota, disease may result from the absence of these symbiotic organisms and their beneficial molecules. Therefore, dysbiosis (a shift in the composition of the intestinal microbiota) could be an underlying factor in the development of IBD.
Abstract
Immunological dysregulation is the cause of many non-infectious human diseases such as autoimmunity, allergy and cancer. The gastrointestinal tract is the primary site of interaction between the host immune system and microorganisms, both symbiotic and pathogenic. In this Review we discuss findings indicating that developmental aspects of the adaptive immune system are influenced by bacterial colonization of the gut. We also highlight the molecular pathways that mediate host–symbiont interactions that regulate proper immune function. Finally, we present recent evidence to support that disturbances in the bacterial microbiota result in dysregulation of adaptive immune cells, and this may underlie disorders such as inflammatory bowel disease. This raises the possibility that the mammalian immune system, which seems to be designed to control microorganisms, is in fact controlled by microorganisms.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Ley, R. E., Peterson, D. A. & Gordon, J. I. Ecological and evolutionary forces shaping microbial diversity in the human intestine. Cell 124, 837–848 (2006).
Dethlefsen, L., McFall-Ngai, M. & Relman, D. A. An ecological and evolutionary perspective on human–microbe mutualism and disease. Nature 449, 811–818 (2007).
Hooper, L. V. Bacterial contributions to mammalian gut development. Trends Microbiol. 12, 129–134 (2004).
Mazmanian, S. K. & Kasper, D. L. The love–hate relationship between bacterial polysaccharides and the host immune system. Nature Rev. Immunol. 6, 849–858 (2006).
Peterson, D. A., Frank, D. N., Pace, N. R. & Gordon, J. I. Metagenomic approaches for defining the pathogenesis of inflammatory bowel diseases. Cell Host Microbe 3, 417–427 (2008).
Frank, D. N. & Pace, N. R. Gastrointestinal microbiology enters the metagenomics era. Curr. Opin. Gastroenterol. 24, 4–10 (2008).
Ley, R. E. et al. Evolution of mammals and their gut microbes. Science 320, 1647–1651 (2008).
Hooper, L. V. & Gordon, J. I. Commensal host–bacterial relationships in the gut. Science 292, 1115–1118 (2001).
Macpherson, A. J. & Harris, N. L. Interactions between commensal intestinal bacteria and the immune system. Nature Rev. Immunol. 4, 478–485 (2004).
Zaneveld, J. et al. Host–bacterial coevolution and the search for new drug targets. Curr. Opin. Chem. Biol. 12, 109–114 (2008).
Falk, P. G., Hooper, L. V., Midtvedt, T. & Gordon, J. I. Creating and maintaining the gastrointestinal ecosystem: what we know and need to know from gnotobiology. Microbiol. Mol. Biol. Rev. 62, 1157–1170 (1998).
Bouskra, D. et al. Lymphoid tissue genesis induced by commensals through NOD1 regulates intestinal homeostasis. Nature 456, 507–510 (2008). This study shows that peptidoglycans from Gram-negative bacteria induce the generation of ILFs through the recognition of NOD1. In the absence of ILF formation, marked changes in the composition of the microbiota occur.
Abrams, G. D., Bauer, H. & Sprinz, H. Influence of the normal flora on mucosal morphology and cellular renewal in the ileum. A comparison of germ-free and conventional mice. Lab. Invest. 12, 355–364 (1963).
Bry, L., Falk, P. G., Midtvedt, T. & Gordon, J. I. A model of host–microbial interactions in an open mammalian ecosystem. Science 273, 1380–1383 (1996).
Cash, H. L., Whitham, C. V., Behrendt, C. L. & Hooper, L. V. Symbiotic bacteria direct expression of an intestinal bactericidal lectin. Science 313, 1126–1130 (2006). These authors show that microbial colonization of germ-free mice induces the production of REG3γ, a secreted C-type lectin. REG3γ is shown to have antimicrobial activity by binding to peptidoglycans, suggesting that microbial species actively shape the intestinal environment to their advantage.
Sonnenburg, J. L., Chen, C. T. & Gordon, J. I. Genomic and metabolic studies of the impact of probiotics on a model gut symbiont and host. PLoS Biol. 4, e413 (2006). Using co-colonization of germ-free mice with B. thetaiotaomicron (a symbiont) and B. longum (a probiotic), this study shows that B. longum can increase the diversity of polysaccharides that can be degraded by B. thetaiotaomicron , demonstrating that distinct intestinal bacterial species can affect each other's function.
Sprinz, H. et al. The response of the germfree guinea pig to oral bacterial challenge with Escherichia coli and Shigella flexneri. Am. J. Pathol. 39, 681–695 (1961).
Maier, B. R. & Hentges, D. J. Experimental Shigella infections in laboratory animals. I. Antagonism by human normal flora components in gnotobiotic mice. Infect. Immun. 6, 168–173 (1972).
Zachar, Z. & Savage, D. C. Microbial interference and colonization of the murine gastrointestinal tract by Listeria monocytogenes. Infect. Immun. 23, 168–174 (1979).
Inagaki, H., Suzuki, T., Nomoto, K. & Yoshikai, Y. Increased susceptibility to primary infection with Listeria monocytogenes in germfree mice may be due to lack of accumulation of L-selectin+ CD44+ T cells in sites of inflammation. Infect. Immun. 64, 3280–3287 (1996).
Nardi, R. M., Silva, M. E., Vieira, E. C., Bambirra, E. A. & Nicoli, J. R. Intragastric infection of germfree and conventional mice with Salmonella typhimurium. Braz. J. Med. Biol. Res. 22, 1389–1392 (1989).
Stecher, B. et al. Salmonella enterica serovar typhimurium exploits inflammation to compete with the intestinal microbiota. PLoS Biol. 5, e244 (2007). These authors show that the intestinal pathogen S . Typhimurium uses inflammation to disturb the commensal microbiota to induce disease.
Nardi, R. M. et al. Bacteriological and immunological aspects of conventional and germfree mice infected with Salmonella typhimurium. Rev. Latinoam. Microbiol. 33, 239–243 (1991).
Podolsky, D. K. The current future understanding of inflammatory bowel disease. Best Pract. Res. Clin. Gastroenterol. 16, 933–943 (2002).
Shanahan, F. Crohn's disease. Lancet 359, 62–69 (2002).
Targan, S. R. & Karp, L. C. Defects in mucosal immunity leading to ulcerative colitis. Immunol. Rev. 206, 296–305 (2005).
Bouma, G. & Strober, W. The immunological and genetic basis of inflammatory bowel disease. Nature Rev. Immunol. 3, 521–533 (2003).
Kullberg, M. C. et al. IL-23 plays a key role in Helicobacter hepaticus-induced T cell-dependent colitis. J. Exp. Med. 203, 2485–2494 (2006).
Hue, S. et al. Interleukin-23 drives innate and T cell-mediated intestinal inflammation. J. Exp. Med. 203, 2473–2483 (2006).
Duerr, R. H. et al. A genome-wide association study identifies IL23R as an inflammatory bowel disease gene. Science 314, 1461–1463 (2006).
Schmechel, S. et al. Linking genetic susceptibility to Crohn's disease with Th17 cell function: IL-22 serum levels are increased in Crohn's disease and correlate with disease activity and IL23R genotype status. Inflamm. Bowel Dis. 14, 204–212 (2008).
Kobayashi, T. et al. IL23 differentially regulates the Th1/Th17 balance in ulcerative colitis and Crohn's disease. Gut 57, 1682–1689 (2008).
Sartor, R. B. Microbial influences in inflammatory bowel diseases. Gastroenterology 134, 577–594 (2008).
Fontenot, J. D. & Rudensky, A. Y. A well adapted regulatory contrivance: regulatory T cell development and the forkhead family transcription factor Foxp3. Nature Immunol. 6, 331–337 (2005).
Vignali, D. A., Collison, L. W. & Workman, C. J. How regulatory T cells work. Nature Rev. Immunol. 8, 523–532 (2008).
Coombes, J. L. et al. A functionally specialized population of mucosal CD103+ DCs induces Foxp3+ regulatory T cells via a TGF-β and retinoic acid-dependent mechanism. J. Exp. Med. 204, 1757–1764 (2007).
Powrie, F. & Maloy, K. J. Immunology. Regulating the regulators. Science 299, 1030–1031 (2003).
Collison, L. W. et al. The inhibitory cytokine IL-35 contributes to regulatory T-cell function. Nature 450, 566–569 (2007).
Ivanov, II. et al. Specific microbiota direct the differentiation of IL-17-producing T-helper cells in the mucosa of the small intestine. Cell Host Microbe 4, 337–349 (2008).
Atarashi, K. et al. ATP drives lamina propria TH17 cell differentiation. Nature 455, 808–812 (2008).
Hall, J. A. et al. Commensal DNA limits regulatory T cell conversion and is a natural adjuvant of intestinal immune responses. Immunity 29, 637–649 (2008).
Rakoff-Nahoum, S., Paglino, J., Eslami-Varzaneh, F., Edberg, S. & Medzhitov, R. Recognition of commensal microflora by Toll-like receptors is required for intestinal homeostasis. Cell 118, 229–241 (2004). This is one of the first studies to suggest that recognition of TLR ligands from commensal bacteria by the host is important for the maintenance of intestinal homeostasis.
Ostman, S., Rask, C., Wold, A. E., Hultkrantz, S. & Telemo, E. Impaired regulatory T cell function in germ-free mice. Eur. J. Immunol. 36, 2336–2346 (2006).
Ishikawa, H. et al. Effect of intestinal microbiota on the induction of regulatory CD25+ CD4+ T cells. Clin. Exp. Immunol. 153, 127–135 (2008).
Strauch, U. G. et al. Influence of intestinal bacteria on induction of regulatory T cells: lessons from a transfer model of colitis. Gut 54, 1546–1552 (2005).
Zaph, C. et al. Commensal-dependent expression of IL-25 regulates the IL-23–IL-17 axis in the intestine. J. Exp. Med. 205, 2191–2198 (2008).
De Winter, H., Cheroutre, H. & Kronenberg, M. Mucosal immunity and inflammation. II. The yin and yang of T cells in intestinal inflammation: pathogenic and protective roles in a mouse colitis model. Am. J. Physiol. 276, G1317–G1321 (1999).
Simpson, S. J., de Jong, Y. P., Comiskey, M. & Terhorst, C. Pathways of T cell pathology in models of chronic intestinal inflammation. Int. Rev. Immunol. 19, 1–37 (2000).
Elson, C. O. et al. Monoclonal anti-interleukin 23 reverses active colitis in a T cell-mediated model in mice. Gastroenterology 132, 2359–2370 (2007).
Sartor, R. B. The influence of normal microbial flora on the development of chronic mucosal inflammation. Res. Immunol. 148, 567–576 (1997).
Macpherson, A., Khoo, U. Y., Forgacs, I., Philpott-Howard, J. & Bjarnason, I. Mucosal antibodies in inflammatory bowel disease are directed against intestinal bacteria. Gut 38, 365–375 (1996).
Elson, C. O. Commensal bacteria as targets in Crohn's disease. Gastroenterology 119, 254–257 (2000).
Tannock, G. W. Exploring the relationships between intestinal microflora and inflammatory conditions of the human bowel and spine. Antonie Van Leeuwenhoek 81, 529–535 (2002).
Kent, T. H., Summers, R. W., DenBesten, L., Swaner, J. C. & Hrouda, M. Effect of antibiotics on bacterial flora of rats with intestinal blind loops. Proc. Soc. Exp. Biol. Med. 132, 63–67 (1969).
Videla, S. et al. Role of intestinal microflora in chronic inflammation and ulceration of the rat colon. Gut 35, 1090–1097 (1994).
Taurog, J. D. et al. The germfree state prevents development of gut and joint inflammatory disease in HLA-B27 transgenic rats. J. Exp. Med. 180, 2359–2364 (1994).
Rath, H. C. Role of commensal bacteria in chronic experimental colitis: lessons from the HLA-B27 transgenic rat. Pathobiology 70, 131–138 (2002).
Sellon, R. K. et al. Resident enteric bacteria are necessary for development of spontaneous colitis and immune system activation in interleukin-10-deficient mice. Infect. Immun. 66, 5224–5231 (1998).
Cahill, R. J. et al. Inflammatory bowel disease: an immunity-mediated condition triggered by bacterial infection with Helicobacter hepaticus. Infect. Immun. 65, 3126–3131 (1997).
Kullberg, M. C. et al. Induction of colitis by a CD4+ T cell clone specific for a bacterial epitope. Proc. Natl Acad. Sci. USA 100, 15830–15835 (2003).
Barnich, N. et al. CEACAM6 acts as a receptor for adherent-invasive E. coli, supporting ileal mucosa colonization in Crohn disease. J. Clin. Invest. 117, 1566–1574 (2007).
Hampe, J. et al. Association between insertion mutation in NOD2 gene and Crohn's disease in German and British populations. Lancet 357, 1925–1928 (2001).
Kim, S. C., Tonkonogy, S. L., Karrasch, T., Jobin, C. & Sartor, R. B. Dual-association of gnotobiotic Il-10−/− mice with 2 nonpathogenic commensal bacteria induces aggressive pancolitis. Inflamm. Bowel Dis. 13, 1457–1466 (2007).
O'Hara, A. M. & Shanahan, F. The gut flora as a forgotten organ. EMBO Rep. 7, 688–693 (2006).
Ley, R. E., Knight, R. & Gordon, J. I. The human microbiome: eliminating the biomedical/environmental dichotomy in microbial ecology. Environ. Microbiol. 9, 3–4 (2007).
Mazmanian, S. K., Round, J. L. & Kasper, D. L. A microbial symbiosis factor prevents intestinal inflammatory disease. Nature 453, 620–625 (2008).
Glimcher, L. H. Trawling for treasure: tales of T-bet. Nature Immunol. 8, 448–450 (2007).
Garrett, W. S. et al. Communicable ulcerative colitis induced by T-bet deficiency in the innate immune system. Cell 131, 33–45 (2007). This study shows that mice that lack T-bet expression in innate immune cells develop spontaneous colitis. Moreover, transfer of the microbiota from these mice is shown to induce disease in wild-type recipient mice.
Turnbaugh, P. J. et al. An obesity-associated gut microbiome with increased capacity for energy harvest. Nature 444, 1027–1031 (2006). These authors show that the microbiome from obese mice has an increased capacity for energy harvest. Transfer of the microbiota to non-obese mice increases their mean fat body weight, suggesting that a change in the microbiota can induce obesity.
Ley, R. E., Turnbaugh, P. J., Klein, S. & Gordon, J. I. Microbial ecology: human gut microbes associated with obesity. Nature 444, 1022–1023 (2006).
Pomp, D., Nehrenberg, D. & Estrada-Smith, D. Complex genetics of obesity in mouse models. Annu. Rev. Nutr. 28, 331–345 (2008).
Lepage, P. et al. Biodiversity of the mucosa-associated microbiota is stable along the distal digestive tract in healthy individuals and patients with IBD. Inflamm. Bowel Dis. 11, 473–480 (2005).
Scanlan, P. D., Shanahan, F., O'Mahony, C. & Marchesi, J. R. Culture-independent analyses of temporal variation of the dominant fecal microbiota and targeted bacterial subgroups in Crohn's disease. J. Clin. Microbiol. 44, 3980–3988 (2006).
Frank, D. N. et al. Molecular-phylogenetic characterization of microbial community imbalances in human inflammatory bowel diseases. Proc. Natl Acad. Sci. USA 104, 13780–13785 (2007). This study indicates that the intestinal microbial populations in patients with IBD and non-IBD patients differ greatly.
Turnbaugh, P. J. et al. The human microbiome project. Nature 449, 804–810 (2007).
Sartor, R. B. Therapeutic manipulation of the enteric microflora in inflammatory bowel diseases: antibiotics, probiotics, and prebiotics. Gastroenterology 126, 1620–1633 (2004).
Di Giacinto, C., Marinaro, M., Sanchez, M., Strober, W. & Boirivant, M. Probiotics ameliorate recurrent Th1-mediated murine colitis by inducing IL-10 and IL-10-dependent TGF-β-bearing regulatory cells. J. Immunol. 174, 3237–3246 (2005).
O'Mahony, C. et al. Commensal-induced regulatory T cells mediate protection against pathogen-stimulated NF-κB activation. PLoS Pathog. 4, e1000112 (2008).
Foligne, B. et al. A key role for dendritic cells in probiotic functionality. PLoS ONE 2, e313 (2007).
Sokol, H. et al. Faecalibacterium prausnitzii is an anti-inflammatory commensal bacterium identified by gut microbiota analysis of Crohn disease patients. Proc. Natl Acad. Sci. USA 105, 16731–16736 (2008). These authors show that F. prausnitzii is specifically reduced in the intestine of patients with Crohn's disease. In addition, this bacterium is shown to have an anti-inflammatory capacity and to protect animals from disease when given orally, suggesting that symbiotic microorganisms may be directly involved in maintaining health.
Mazmanian, S. K., Liu, C. H., Tzianabos, A. O. & Kasper, D. L. An immunomodulatory molecule of symbiotic bacteria directs maturation of the host immune system. Cell 122, 107–118 (2005).
Roncarolo, M. G. & Battaglia, M. Regulatory T-cell immunotherapy for tolerance to self antigens and alloantigens in humans. Nature Rev. Immunol. 7, 585–598 (2007).
Cong, Y. et al. Generation of antigen-specific, Foxp3-expressing CD4+ regulatory T cells by inhibition of APC proteosome function. J. Immunol. 174, 2787–2795 (2005).
Noverr, M. C. & Huffnagle, G. B. Does the microbiota regulate immune responses outside the gut? Trends Microbiol. 12, 562–568 (2004).
Bjorksten, B. The environmental influence on childhood asthma. Allergy 54, S17–S23 (1999).
Penders, J. et al. Gut microbiota composition and development of atopic manifestations in infancy: the KOALA Birth Cohort Study. Gut 56, 661–667 (2007).
Kalliomaki, M. & Isolauri, E. Pandemic of atopic diseases — a lack of microbial exposure in early infancy? Curr. Drug Targets. Infect. Disord. 2, 193–199 (2002).
Kalliomaki, M. & Isolauri, E. Role of intestinal flora in the development of allergy. Curr. Opin. Allergy Clin. Immunol. 3, 15–20 (2003).
Sakaguchi, S. et al. Foxp3+ CD25+CD4+ natural regulatory T cells in dominant self-tolerance and autoimmune disease. Immunol. Rev. 212, 8–27 (2006).
Eckburg, P. B. et al. Diversity of the human intestinal microbial flora. Science 308, 1635–1638 (2005).
Gill, S. R. et al. Metagenomic analysis of the human distal gut microbiome. Science 312, 1355–1359 (2006).
Palmer, C., Bik, E. M., Digiulio, D. B., Relman, D. A. & Brown, P. O. Development of the human infant intestinal microbiota. PLoS Biol. 5, e177 (2007).
Izcue, A. et al. Interleukin-23 restrains regulatory T cell activity to drive T cell-dependent colitis. Immunity 28, 559–570 (2008). This report shows that FOXP3-deficient T cells can induce colitis in IL-23-deficient recipients, suggesting that disease can occur in the absence of regulation.
Moreau, M. C., Ducluzeau, R., Guy-Grand, D. & Muller, M. C. Increase in the population of duodenal immunoglobulin A plasmocytes in axenic mice associated with different living or dead bacterial strains of intestinal origin. Infect. Immun. 21, 532–539 (1978).
Suzuki, K. et al. Aberrant expansion of segmented filamentous bacteria in IgA-deficient gut. Proc. Natl Acad. Sci. USA 101, 1981–1986 (2004).
Kroese, F. G., de Waard, R. & Bos, N. A. B-1 cells and their reactivity with the murine intestinal microflora. Semin. Immunol. 8, 11–18 (1996).
Macpherson, A. J. & Uhr, T. Induction of protective IgA by intestinal dendritic cells carrying commensal bacteria. Science 303, 1662–1665 (2004).
Macpherson, A. J., Geuking, M. B. & McCoy, K. D. Immune responses that adapt the intestinal mucosa to commensal intestinal bacteria. Immunology 115, 153–162 (2005).
He, B. et al. Intestinal bacteria trigger T cell-independent immunoglobulin A2 class switching by inducing epithelial-cell secretion of the cytokine APRIL. Immunity 26, 812–826 (2007).
Cerutti, A. The regulation of IgA class switching. Nature Rev. Immunol. 8, 421–434 (2008).
Tezuka, H. et al. Regulation of IgA production by naturally occurring TNF/iNOS-producing dendritic cells. Nature 448, 929–933 (2007).
Peterson, D. A., McNulty, N. P., Guruge, J. L. & Gordon, J. I. IgA response to symbiotic bacteria as a mediator of gut homeostasis. Cell Host Microbe 2, 328–339 (2007).
O'Mahony, S. M. et al. Early life stress alters behavior, immunity, and microbiota in rats: implications for irritable bowel syndrome and psychiatric illnesses. Biol. Psychiatry 65, 263–267 (2009).
Acknowledgements
We thank members of the Mazmanian laboratory for their critical review of the manuscript. We apologize to those whose work could not be mentioned owing to space limitations and the scope of the manuscript, in particular the vast clinical data on inflammatory bowel disease. J.L.R is a Merck Fellow of the Jane Coffin Child's Memorial Fund. S.K.M. is a Searle Scholar. Work in our laboratory is supported by funding from the National Institutes of Health, USA, Damon Runyon Cancer Research Foundation and the Crohn's & Colitis Foundation of America to S.K.M.
Author information
Authors and Affiliations
Corresponding author
Related links
Glossary
- Mutualism
-
A symbiotic association in which both members benefit from the relationship.
- Pathogen
-
An opportunistic organism that rarely comes into contact with the host, but causes acute or chronic disease following infection. Derived from the Greek word 'pathos', which means suffering.
- Microbiome
-
The collective genomes of a microbiota.
- Microbiota
-
The amalgam of microorganisms that make up a complex and diverse community living within a given anatomical niche (usually an environmentally exposed surface of the body).
- Inflammatory bowel disease
-
A chronic condition of the intestine that is characterized by severe inflammation and mucosal destruction. The most common forms in humans are ulcerative colitis and Crohn's disease.
- Gut-associated lymphoid tissue
-
Lymphoid structures and aggregates associated with the intestinal mucosa, specifically the tonsils, Peyer's patches, lymphoid follicles, appendix or caecal patch and mesenteric lymph nodes. They are enriched in conventional and unconventional lymphocytes and specialized dendritic cell and macrophage subsets. They provide the first line of defence against entry of pathogens across the mucosal barrier.
- Peyer's patches
-
Groups of lymphoid nodules that are present in the small intestine (usually the ileum). They occur massed together on the intestinal wall, opposite the line of attachment of the mesentery. Peyer's patches consist of a dome area, B cell follicles and interfollicular T cell areas. High endothelial venules are present mainly in the interfollicular areas.
- Mesenteric lymph node
-
(MLN). A lymph node that is located at the base of the mesentery. MLNs collect lymph (including cells and antigens) draining from the intestinal mucosa.
- Specific pathogen free
-
Conditions in which animals are reared and maintained in an environment with an unknown complex microbiota that is free from specific known pathogens.
- Isolated lymphoid follicles
-
Small lymphoid aggregates located in the anti-mesenteric wall of the small intestine, which contain B cells, dendritic cells, stromal cells and some T cells. They may contain germinal centres. They are thought to have a role in maintaining equilibrium between the immune system and the microbiota.
- Pattern recognition receptor
-
A host receptor (such as Toll-like receptors) that can sense pathogen-associated molecular patterns and initiate signalling cascades (which involve the activation of nuclear factor-κB) that lead to an innate immune response.
- Commensal
-
A microorganism that benefits from an association with no known effects on the host. Derived from the Latin phrase 'com mensa', meaning to share a table.
- Crohn's disease
-
A form of chronic inflammatory bowel disease that can affect the entire gastrointestinal tract, but is most common in the colon and terminal ileum. It is characterized by transmural inflammation, strictures and granuloma formation, and is believed to result from an abnormal T cell-mediated immune response to commensal bacteria.
- Ulcerative colitis
-
A chronic disease that is characterized by inflammation of the mucosa and sub-mucosa of the large intestine.
- Regulatory T (TReg) cell
-
A specialized type of CD4+ T cell that can suppress the responses of other T cells. These cells provide a crucial mechanism for the maintenance of peripheral self tolerance and are characterized by expression of CD25 (the α-chain of the interleukin-2 receptor) and the transcription factor forkhead box P3 (FOXP3).
- Parasite
-
An opportunistic organism that maintains a prolonged and close association with the host, which benefits the parasite at the expense of the host.
- Pathobiont
-
A symbiont that does not normally elicit an inflammatory response but under particular conditions (environmentally induced) has the potential to cause dysregulated inflammation and lead to disease.
- VSL#3
-
A mixture of bacteria consisting of four strains of Lactobacillus (L. casei, L. plantarum, L. acidophilus and L. delbrueckii subspecies bulgaricus), three strains of Bifidobacterium (B. longum, B. breve and B. infantis) and Streptococcus salivarius subspecies thermophilus.
- Symbiont
-
An organism that lives in association with a host (usually for a lifetime) without obvious benefit or harm to either member.
- Symbiosis
-
A constant and intimate relationship that occurs between dissimilar species, which was originally defined as 'living together'. Although it is often used to describe a beneficial relationship, symbiosis does not necessarily imply that either partner gains an advantage.
Rights and permissions
About this article
Cite this article
Round, J., Mazmanian, S. The gut microbiota shapes intestinal immune responses during health and disease. Nat Rev Immunol 9, 313–323 (2009). https://doi.org/10.1038/nri2515
Issue Date:
DOI: https://doi.org/10.1038/nri2515
This article is cited by
-
The role of gut microbiota in human metabolism and inflammatory diseases: a focus on elderly individuals
Annals of Microbiology (2024)
-
Microbes little helpers and suppliers for therapeutic asthma approaches
Respiratory Research (2024)
-
Monolithic-to-focal evolving biointerfaces in tissue regeneration and bioelectronics
Nature Chemical Engineering (2024)
-
Intestinal epithelial Krüppel-like factor 4 alleviates endotoxemia and atherosclerosis through improving NF-κB/miR-34a-mediated intestinal permeability
Acta Pharmacologica Sinica (2024)
-
Seasonal changes in gut microbiota of sea cucumber over natural aestivation cycle
Journal of Oceanology and Limnology (2024)