Abstract
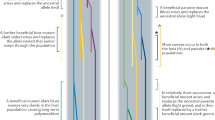
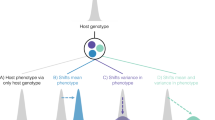
Species from all five kingdoms of life have evolved sophisticated mechanisms to generate diversity in genes that are involved in host–pathogen interactions, conferring reduced levels of parasitism to both individuals and populations. Here, we highlight unifying concepts that underpin these evolutionarily unrelated diversity-generating mechanisms (DGMs). We discuss the mechanisms of and selective forces acting on these diversity-generating immune strategies, as well as their epidemiological and co-evolutionary consequences. We propose that DGMs can be broadly classified into two classes — targeted and untargeted DGMs — which generate different levels of diversity with important consequences for host–parasite co-evolution.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Keesing, F. et al. Impacts of biodiversity on the emergence and transmission of infectious diseases. Nature 468, 647–652 (2010).
Elton, C. S. in The Ecology of Invasions by Animals and Plants (Methuen, 1958).
O'Brien, S. J. et al. Genetic basis for species vulnerability in the cheetah. Science 227, 1428–1434 (1985).
Acevedo-Whitehouse, K., Gulland, F., Greig, D. & Amos, W. Inbreeding: disease susceptibility in California sea lions. Nature 422, 35 (2003).
Altermatt, F. & Ebert, D. Genetic diversity of Daphnia magna populations enhances resistance to parasites. Ecol. Lett. 11, 918–928 (2008).
van Houte, S. et al. The diversity-generating benefits of a prokaryotic adaptive immune system. Nature 532, 358–388 (2016).
Lynch, M. et al. Genetic drift, selection and the evolution of the mutation rate. Nat. Rev. Genet. 17, 704–714 (2016).
Taddei, F. et al. Role of mutator alleles in adaptive evolution. Nature 387, 700–702 (1997).
Zaman, L. et al. Coevolution drives the emergence of complex traits and promotes evolvability. PLoS Biol. 12, e1002023 (2014).
Pal, C., Macia, M. D., Oliver, A., Schachar, I. & Buckling, A. Coevolution with viruses drives the evolution of bacterial mutation rates. Nature 450, 1079–1081 (2007).
Wielgoss, S., Bergmiller, T., Bischofberger, A. M. & Hall, A. R. Adaptation to parasites and costs of parasite resistance in mutator and nonmutator bacteria. Mol. Biol. Evol. 33, 770–782 (2016).
LeClerc, J. E., Li, B., Payne, W. L. & Cebula, T. A. High mutation frequencies among Escherichia coli and Salmonella pathogens. Science 274, 1208–1211 (1996).
Matic, I. et al. Highly variable mutation rates in commensal and pathogenic Escherichia coli. Science 277, 1833–1834 (1997).
Oliver, A., Canton, R., Campo, P., Baquero, F. & Blazquez, J. High frequency of hypermutable Pseudomonas aeruginosa in cystic fibrosis lung infection. Science 288, 1251–1254 (2000).
Lee, H., Popodi, E., Tang, H. & Foster, P. L. Rate and molecular spectrum of spontaneous mutations in the bacterium Escherichia coli as determined by whole-genome sequencing. Proc. Natl Acad. Sci. USA 109, E2774–2783 (2012).
Lynch, M. Rate, molecular spectrum, and consequences of human mutation. Proc. Natl Acad. Sci. USA 107, 961–968 (2010).
Herman, R. K. & Dworkin, N. B. Effect of gene induction on the rate of mutagenesis by ICR-191 in Escherichia coli. J. Bacteriol. 106, 543–550 (1971).
Datta, A. & Jinks-Robertson, S. Association of increased spontaneous mutation rates with high levels of transcription in yeast. Science 268, 1616–1619 (1995).
Makova, K. D. & Hardison, R. C. The effects of chromatin organization on variation in mutation rates in the genome. Nat. Rev. Genet. 16, 213–223 (2015).
Wolfe, K. H., Sharp, P. M. & Li, W. H. Mutation rates differ among regions of the mammalian genome. Nature 337, 283–285 (1989).
Hardison, R. C. et al. Covariation in frequencies of substitution, deletion, transposition, and recombination during eutherian evolution. Genome Res. 13, 13–26 (2003).
Chuang, J. H. & Li, H. Functional bias and spatial organization of genes in mutational hot and cold regions in the human genome. PLoS Biol. 2, E29 (2004).
Hamilton, W. D., Axelrod, R. & Tanese, R. Sexual reproduction as an adaptation to resist parasites (a review). Proc. Natl Acad. Sci. USA 87, 3566–3573 (1990).
Lively, C. M. A review of Red Queen models for the persistence of obligate sexual reproduction. J. Hered. 101 (Suppl. 1), S13–S20 (2010).
Lively, C. M. & Morran, L. T. The ecology of sexual reproduction. J. Evol. Biol. 27, 1292–1303 (2014).
Lively, C. Evidence from a New-Zealand snail for the maintenance of sex by parasitism. Nature 328, 519–521 (1987).
King, K. C., Delph, L. F., Jokela, J. & Lively, C. M. The geographic mosaic of sex and the Red Queen. Curr. Biol. 19, 1438–1441 (2009).
King, K. C. & Lively, C. M. Geographic variation in sterilizing parasite species and the Red Queen. Oikos 118, 1416–1420 (2009).
Vergara, D., Jokela, J. & Lively, C. M. Infection dynamics in coexisting sexual and asexual host populations: support for the Red Queen hypothesis. Am. Nat. 184 (Suppl. 1), S22–S30 (2014).
Morran, L. T. et al. Running with the Red Queen: host-parasite coevolution selects for biparental sex. Science 333, 216–218 (2011).
Paigen, K. & Petkov, P. Mammalian recombination hot spots: properties, control and evolution. Nat. Rev. Genet. 11, 221–233 (2010).
Hamilton, W. D. & Zuk, M. Heritable true fitness and bright birds: a role for parasites? Science 218, 384–387 (1982).
Kamiya, T., O'Dwyer, K., Westerdahl, H., Senior, A. & Nakagawa, S. A quantitative review of MHC-based mating preference: the role of diversity and dissimilarity. Mol. Ecol. 23, 5151–5163 (2014).
Reusch, T. B., Haberli, M. A., Aeschlimann, P. B. & Milinski, M. Female sticklebacks count alleles in a strategy of sexual selection explaining MHC polymorphism. Nature 414, 300–302 (2001).
Potts, W. K., Manning, C. J. & Wakeland, E. K. Mating patterns in seminatural populations of mice influenced by MHC genotype. Nature 352, 619–621 (1991).
Leinders-Zufall, T. et al. MHC class I peptides as chemosensory signals in the vomeronasal organ. Science 306, 1033–1037 (2004).
Baer, B. & Schmid-Hempel, P. Experimental variation in polyandry affects parasite loads and fitness in a bumble-bee. Nature 397, 151–154 (1999).
Baer, B. & Schmid-Hempel, P. Bumblebee workers from different sire groups vary in suscetibility to parasite infection. Ecol. Lett. 6, 106–110 (2003).
Baer, B. & Schmid-Hempel, P. Unexpected consequences of polyandry for parasitism and fitness in the bumblebee, Bombus terrestris. Evolution 55, 1639–1643 (2001).
de Visser, J. A. & Elena, S. F. The evolution of sex: empirical insights into the roles of epistasis and drift. Nat. Rev. Genet. 8, 139–149 (2007).
Giraud, A. et al. Costs and benefits of high mutation rates: adaptive evolution of bacteria in the mouse gut. Science 291, 2606–2608 (2001).
Sniegowski, P., Gerrish, P. & Lenski, R. Evolution of high mutation rates in experimental populations of E-coli. Nature 387, 703–705 (1997).
Morran, L. T., Parmenter, M. D. & Phillips, P. C. Mutation load and rapid adaptation favour outcrossing over self-fertilization. Nature 462, 350–352 (2009).
McDonald, M. J., Rice, D. P. & Desai, M. M. Sex speeds adaptation by altering the dynamics of molecular evolution. Nature 531, 233–236 (2016).
Lumley, A. J. et al. Sexual selection protects against extinction. Nature 522, 470–473 (2015).
Forsberg, L. A., Gisselsson, D. & Dumanski, J. P. Mosaicism in health and disease — clones picking up speed. Nat. Rev. Genet. 18, 128–142 (2017).
Rast, J. P. & Litman, G. W. T-Cell receptor gene homologs are present in the most primitive jawed vertebrates. Proc. Natl Acad. Sci. USA 91, 9248–9252 (1994).
Weinstein, J. A., Jiang, N., White, R. A.,3rd, Fisher, D. S. & Quake, S. R. High-throughput sequencing of the zebrafish antibody repertoire. Science 324, 807–810 (2009).
Jiang, N. et al. Determinism and stochasticity during maturation of the zebrafish antibody repertoire. Proc. Natl Acad. Sci. USA 108, 5348–5353 (2011).
Boulinier, T. & Staszewski, V. Maternal transfer of antibodies: raising immuno-ecology issues. Trends Ecol. Evol. 23, 282–288 (2008).
Early, P., Huang, H., Davis, M., Calame, K. & Hood, L. An immunoglobulin heavy chain variable region gene is generated from three segments of DNA: VH, D and JH. Cell 19, 981–992 (1980).
Schatz, D. G., Oettinger, M. A. & Baltimore, D. The V(D)J recombination activating gene, RAG-1. Cell 59, 1035–1048 (1989).
Kim, M. S., Lapkouski, M., Yang, W. & Gellert, M. Crystal structure of the V(D)J recombinase RAG1–RAG2. Nature 518, 507–511 (2015).
Schwarz, K. et al. RAG mutations in human B cell-negative SCID. Science 274, 97–99 (1996).
Maizels, N. Diversity achieved by diverse mechanisms: gene conversion in developing B cells of the chicken. Cell 48, 359–360 (1987).
Allen, C. D., Okada, T., Tang, H. L. & Cyster, J. G. Imaging of germinal center selection events during affinity maturation. Science 315, 528–531 (2007).
Victora, G. D. et al. Germinal center dynamics revealed by multiphoton microscopy with a photoactivatable fluorescent reporter. Cell 143, 592–605 (2010).
Gitlin, A. D. et al. Humoral immunity. T cell help controls the speed of the cell cycle in germinal center B cells. Science 349, 643–646 (2015).
Pappas, L. et al. Rapid development of broadly influenza neutralizing antibodies through redundant mutations. Nature 516, 418–422 (2014).
Gitlin, A. D., Shulman, Z. & Nussenzweig, M. C. Clonal selection in the germinal centre by regulated proliferation and hypermutation. Nature 509, 637–640 (2014).
Tas, J. M. et al. Visualizing antibody affinity maturation in germinal centers. Science 351, 1048–1054 (2016).
Kuraoka, M. et al. Complex antigens drive permissive clonal selection in germinal centers. Immunity 44, 542–552 (2016).
Horns, F. et al. Lineage tracing of human B cells reveals the in vivo landscape of human antibody class switching. eLife 5, e16578 (2016).
Pancer, Z. et al. Somatic diversification of variable lymphocyte receptors in the agnathan sea lamprey. Nature 430, 174–180 (2004).
Boehm, T. et al. VLR-based adaptive immunity. Annu. Rev. Immunol. 30, 203–220 (2012).
Alder, M. N. et al. Diversity and function of adaptive immune receptors in a jawless vertebrate. Science 310, 1970–1973 (2005).
Wang, X. H. et al. RNA interference directs innate immunity against viruses in adult Drosophila. Science 312, 452–454 (2006).
van Rij, R. P. et al. The RNA silencing endonuclease Argonaute 2 mediates specific antiviral immunity in Drosophila melanogaster. Genes Dev. 20, 2985–2995 (2006).
Li, H., Li, W. X. & Ding, S. W. Induction and suppression of RNA silencing by an animal virus. Science 296, 1319–1321 (2002).
Hamilton, A. J. & Baulcombe, D. C. A species of small antisense RNA in posttranscriptional gene silencing in plants. Science 286, 950–952 (1999).
Lu, R. et al. Animal virus replication and RNAi-mediated antiviral silencing in Caenorhabditis elegans. Nature 436, 1040–1043 (2005).
Maillard, P. V. et al. Antiviral RNA interference in mammalian cells. Science 342, 235–238 (2013).
Li, Y., Lu, J., Han, Y., Fan, X. & Ding, S. W. RNA interference functions as an antiviral immunity mechanism in mammals. Science 342, 231–234 (2013).
Ding, S. W. & Voinnet, O. Antiviral immunity directed by small RNAs. Cell 130, 413–426 (2007).
Molnar, A. et al. Plant virus-derived small interfering RNAs originate predominantly from highly structured single-stranded viral RNAs. J. Virol. 79, 7812–7818 (2005).
Lafforgue, G. et al. Tempo and mode of plant RNA virus escape from RNA interference-mediated resistance. J. Virol. 85, 9686–9695 (2011).
Martinez, F. et al. Ultradeep sequencing analysis of population dynamics of virus escape mutants in RNAi-mediated resistant plants. Mol. Biol. Evol. 29, 3297–3307 (2012).
Holz, C. L. et al. RNA interference against animal viruses: how morbilliviruses generate extended diversity to escape small interfering RNA control. J. Virol. 86, 786–795 (2012).
Swaney, S., Powers, H., Goodwin, J., Rosales, L. S. & Dougherty, W. G. RNA-mediated resistance with nonstructural genes from the tobacco etch virus genome. Mol. Plant Microbe Interact. 8, 1004–1011 (1995).
Swarts, D. C. et al. The evolutionary journey of Argonaute proteins. Nat. Struct. Mol. Biol. 21, 743–753 (2014).
Barrangou, R. et al. CRISPR provides acquired resistance against viruses in prokaryotes. Science 315, 1709–1712 (2007).
Brouns, S. J. et al. Small CRISPR RNAs guide antiviral defense in prokaryotes. Science 321, 960–964 (2008).
Garneau, J. E. et al. The CRISPR/Cas bacterial immune system cleaves bacteriophage and plasmid DNA. Nature 468, 67–71 (2010).
East-Seletsky, A. et al. Two distinct RNase activities of CRISPR-C2c2 enable guide-RNA processing and RNA detection. Nature 538, 270–273 (2016).
Abudayyeh, O. O. et al. C2c2 is a single-component programmable RNA-guided RNA-targeting CRISPR effector. Science 353, aaf5573 (2016).
Goldberg, G. W., Jiang, W., Bikard, D. & Marraffini, L. A. Conditional tolerance of temperate phages via transcription-dependent CRISPR-Cas targeting. Nature 514, 633–637 (2014).
Samai, P. et al. Co-transcriptional DNA and RNA cleavage during type III CRISPR-Cas immunity. Cell 161, 1164–1174 (2015).
Jiang, F. et al. Structures of a CRISPR-Cas9 R-loop complex primed for DNA cleavage. Science 351, 867–871 (2016).
Richter, C. et al. Priming in the Type I-F CRISPR-Cas system triggers strand-independent spacer acquisition, bi-directionally from the primed protospacer. Nucleic Acids Res. 42, 8516–8526 (2014).
Modell, J. W., Jiang, W. & Marraffini, L. A. CRISPR-Cas systems exploit viral DNA injection to establish and maintain adaptive immunity. Nature 544, 101–104 (2017).
Westra, E. et al. Parasite exposure drives selective evolution of constitutive versus inducible defense. Curr. Biol. 25, 1043–1049 (2015).
Levin, B. R., Moineau, S., Bushman, M. & Barrangou, R. The population and evolutionary dynamics of phage and bacteria with CRISPR-mediated immunity. PLoS Genet. 9, e1003312 (2013).
Bikard, D. & Marraffini, L. A. Innate and adaptive immunity in bacteria: mechanisms of programmed genetic variation to fight bacteriophages. Curr. Opin. Immunol. 24, 15–20 (2012).
van Houte, S., Buckling, A. & Westra, E. R. Evolutionary ecology of prokaryotic immune mechanisms. Microbiol. Mol. Biol. Rev. 80, 745–763 (2016).
Korona, R. & Levin, B. R. Phage-mediated selection and the evolution and maintenance of restriction-modification. Evolution 47, 556–575 (1993).
Levin, B. R., Antonovics, J. & Sharma, H. Frequency-dependent selection in bacterial populations. Phil. Trans. R. Soc. Series B - Biol. Sci. 319, 459–472 (1988).
Dybvig, K., Sitaraman, R. & French, C. T. A family of phase-variable restriction enzymes with differing specificities generated by high-frequency gene rearrangements. Proc. Natl Acad. Sci. USA 95, 13923–13928 (1998).
Tettelin, H. et al. Complete genome sequence of a virulent isolate of Streptococcus pneumoniae. Science 293, 498–506 (2001).
Sneppen, K., Semsey, S., Seshasayee, A. & Krishna, S. Restriction modification systems as engines of diversity. Frontiers Microbiol. 6, 528 (2015).
Ketting, R. F. The many faces of RNAi. Dev. Cell. 20, 148–161 (2011).
Westra, E. R., Buckling, A. & Fineran, P. C. CRISPR-Cas systems: beyond adaptive immunity. Nat. Rev. Microbiol. 12, 317–326 (2014).
Sampson, T. R., Saroj, S. D., Llewellyn, A. C., Tzeng, Y. & Weiss, D. S. A. CRISPR/Cas system mediates bacterial innate immune evasion and virulence. Nature 497, 254–257 (2013).
Manso, A. S. et al. A random six-phase switch regulates pneumococcal virulence via global epigenetic changes. Nat. Commun. 5, 5055 (2014).
Seo, G. J. et al. Reciprocal inhibition between intracellular antiviral signaling and the RNAi machinery in mammalian cells. Cell. Host Microbe 14, 435–445 (2013).
Morgan, A. D., Bonsall, M. B. & Buckling, A. Impact of bacterial mutation rate on coevolutionary dynamics between bacteria and phages. Evolution 64, 2980–2987 (2010).
Lively, C. M. The effect of host genetic diversity on disease spread. Am. Nat. 175, E149–152 (2010).
Antia, R., Regoes, R. R., Koella, J. C. & Bergstrom, C. T. The role of evolution in the emergence of infectious diseases. Nature 426, 658–661 (2003).
Zhu, Y. et al. Genetic diversity and disease control in rice. Nature 406, 718–722 (2000).
Tubbs, A. & Nussenzweig, A. Endogenous DNA damage as a source of genomic instability in cancer. Cell 168, 644–656 (2017).
Foster, P. L. Stress-induced mutagenesis in bacteria. Crit. Rev. Biochem. Mol. Biol. 42, 373–397 (2007).
Maynard Smith, J. in Group Selection (ed. Williams, G. C.) 163–175 (Aldine-Atherton, 1971).
Maynard Smith, J. in The Evolution of Sex (Cambridge Univ. Press, 1978).
Graham, A., Allen, J. & Read, A. Evolutionary causes and consequences of immunopathology. Annu. Rev. Ecol. Evol. Syst. 36, 373–397 (2005).
Finch, C. E. & Crimmins, E. M. Inflammatory exposure and historical changes in human life-spans. Science 305, 1736–1739 (2004).
Dong, J. et al. Orientation-specific joining of AID-initiated DNA breaks promotes antibody class switching. Nature 525, 134–139 (2015).
Vale, P. F. et al. Costs of CRISPR-Cas-mediated resistance in Streptococcus thermophilus. Proc. Biol. Sci. 282, 20151270 (2015).
Kobasa, D. et al. Aberrant innate immune response in lethal infection of macaques with the 1918 influenza virus. Nature 445, 319–323 (2007).
Graham, A. L. et al. Fitness correlates of heritable variation in antibody responsiveness in a wild mammal. Science 330, 662–665 (2010).
Pleška, M. et al. Bacterial autoimmunity due to a restriction-modification system. Curr. Biol. 26, 1–6 (2016).
Stern, A., Keren, L., Wurtzel, O., Amitai, G. & Sorek, R. Self-targeting by CRISPR: gene regulation or autoimmunity? Trends Genet. 26, 335–340 (2010).
Teng, G. et al. RAG represents a widespread threat to the lymphocyte genome. Cell 162, 751–765 (2015).
Ashby, B. & King, K. C. Diversity and the maintenance of sex by parasites. J. Evol. Biol. 28, 511–520 (2015).
Morgan, A. D., Gandon, S. & Buckling, A. The effect of migration on local adaptation in a coevolving host-parasite system. Nature 437, 253–256 (2005).
Buckling, A. & Rainey, P. B. Antagonistic coevolution between a bacterium and a bacteriophage. Proc. Biol. Sci. 269, 931–936 (2002).
Luijckx, P., Ben-Ami, F., Mouton, L., Du Pasquier, L. & Ebert, D. Cloning of the unculturable parasite Pasteuria ramosa and its Daphnia host reveals extreme genotype-genotype interactions. Ecol. Lett. 14, 125–131 (2011).
Luijckx, P., Fienberg, H., Duneau, D. & Ebert, D. A matching-allele model explains host resistance to parasites. Curr. Biol. 23, 1085–1088 (2013).
Lively, C. M. & Dybdahl, M. F. Parasite adaptation to locally common host genotypes. Nature 405, 679–681 (2000).
Lively, C., Craddock, C. & Vrijenhoek, R. Red Queen hypothesis supported by parasitism in sexual and clonal fish. Nature 344, 864–866 (1990).
Decaestecker, E. et al. Host-parasite 'Red Queen' dynamics archived in pond sediment. Nature 450, 870–873 (2007).
Gomez, P. & Buckling, A. Bacteria-phage antagonistic coevolution in soil. Science 332, 106–109 (2011).
Deveau, H. et al. Phage response to CRISPR-encoded resistance in Streptococcus thermophilus. J. Bacteriol. 190, 1390–1400 (2008).
Morley, D., Broniewski, J. M., Westra, E. R., Buckling, A. & van Houte, S. Host diversity limits the evolution of parasite local adaptation. Mol. Ecol. 26, 1756–1763 (2016).
Chabas, H., van Houte, S., Hoyland-Kroghsbo, N. M., Buckling, A. & Westra, E. R. Immigration of susceptible hosts triggers the evolution of alternative parasite defence strategies. Proc. Biol. Sci. 283, 20160721 (2016).
Griffith, F. The significance of pneumococcal types. J. Hyg. (Lond.) 27, 113–159 (1928).
Schwede, A., Macleod, O. J., MacGregor, P. & Carrington, M. How does the VSG coat of bloodstream form african trypanosomes interact with external proteins? PLoS Pathog. 11, e1005259 (2015).
Su, X. Z. et al. The large diverse gene family var encodes proteins involved in cytoadherence and antigenic variation of Plasmodium falciparum-infected erythrocytes. Cell 82, 89–100 (1995).
Gerber, M., Isel, C., Moules, V. & Marquet, R. Selective packaging of the influenza A genome and consequences for genetic reassortment. Trends Microbiol. 22, 446–455 (2014).
Paul, B. G. et al. Targeted diversity generation by intraterrestrial archaea and archaeal viruses. Nat. Commun. 6, 6585 (2015).
Doulatov, S. et al. Tropism switching in Bordetella bacteriophage defines a family of diversity-generating retroelements. Nature 431, 476–481 (2004).
Gemmill, A., Viney, M. & Read, A. Host immune status determines sexuality in a parasitic nematode. Evolution 51, 393–401 (1997).
Pumplin, N. & Voinnet, O. RNA silencing suppression by plant pathogens: defence, counter-defence and counter-counter-defence. Nat. Rev. Microbiol. 11, 745–760 (2013).
Young, D., Hussell, T. & Dougan, G. Chronic bacterial infections: living with unwanted guests. Nat. Immunol. 3, 1026–1032 (2002).
Belkaid, Y., Piccirillo, C. A., Mendez, S., Shevach, E. M. & Sacks, D. L. CD4+CD25+ regulatory T cells control Leishmania major persistence and immunity. Nature 420, 502–507 (2002).
Atanasiu, C., Su, T. J., Sturrock, S. S. & Dryden, D. T. Interaction of the ocr gene 0.3 protein of bacteriophage T7 with EcoKI restriction/modification enzyme. Nucleic Acids Res. 30, 3936–3944 (2002).
Bondy-Denomy, J., Pawluk, A., Maxwell, K. L. & Davidson, A. R. Bacteriophage genes that inactivate the CRISPR/Cas bacterial immune system. Nature 493, 429–432 (2013).
Pawluk, A. et al. Inactivation of CRISPR-Cas systems by anti-CRISPR proteins in diverse bacterial species. Nat. Microbiol. 1, 16085 (2016).
Bondy-Denomy, J. et al. Multiple mechanisms for CRISPR-Cas inhibition by anti-CRISPR proteins. Nature 526, 136–139 (2015).
Pawluk, A. et al. Naturally occurring off-switches for CRISPR-Cas9. Cell 167, 1829–1838.e9 (2016).
Rauch, B. J. et al. Inhibition of CRISPR-Cas9 with bacteriophage proteins. Cell 168, 150–158.e10 (2017).
Obbard, D. J., Jiggins, F. M., Halligan, D. L. & Little, T. J. Natural selection drives extremely rapid evolution in antiviral RNAi genes. Curr. Biol. 16, 580–585 (2006).
Acknowledgements
E.R.W. and A.B. acknowledge the Natural Environment Research Council, the Biotechnology and Biological Sciences Research Council, the Royal Society, the Leverhulme Trust, the Wellcome Trust, the European Research Council and the AXA research fund for funding.
Author information
Authors and Affiliations
Contributions
E.R.W., D.S. and M.L. researched data for the article, contributed to discussion of the content, wrote the article and reviewed and/or edited the manuscript before submission. A.B. contributed to discussion of the content and reviewed and/or edited the manuscript before submission.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Glossary
- Monoculture effect
-
The increased incidence of diseases in monocultures of the same crop.
- Indels
-
The insertion or deletion of bases in the DNA of an organism.
- Transposable element
-
A DNA sequence that can mobilize to a new position within the genome.
- Genetic drift
-
A change in allele frequencies as a result of the random sampling of gametes that form the next generation.
- Parthenogenic
-
Reproducing in an asexual manner.
- Germinal centres
-
Sites within secondary lymphoid tissue where B cell proliferation, selection and maturation take place during antibody responses.
- CRISPR escape phage
-
Phage that acquire mutations at positions in the protospacer (the sequence matching the CRISPR spacer) or the protospacer adjacent motif (a short DNA sequence required for CRISPR activity) that allow them to overcome CRISPR-Cas immunity.
- Arms-race dynamics
-
(ARD). Co-evolutionary dynamics that are characterized by the increase of both host resistance and pathogen infectivity ranges: hosts evolve resistance to a broader range of pathogen genotypes and pathogens evolve infectivity to a broader range of host genotypes.
- Fluctuating selection dynamics
-
(FSD). Co-evolutionary dynamics that are characterized by fluctuations in host and pathogen genotypes owing to frequency dependent selection, whereby the fitness of host genotypes is inversely correlated with their frequency in the population.
Rights and permissions
About this article
Cite this article
Westra, E., Sünderhauf, D., Landsberger, M. et al. Mechanisms and consequences of diversity-generating immune strategies. Nat Rev Immunol 17, 719–728 (2017). https://doi.org/10.1038/nri.2017.78
Published:
Issue Date:
DOI: https://doi.org/10.1038/nri.2017.78
This article is cited by
-
Phenotype Design Space Provides a Mechanistic Framework Relating Molecular Parameters to Phenotype Diversity Available for Selection
Journal of Molecular Evolution (2023)
-
High-throughput sequencing analysis reveals genomic similarity in phenotypic heterogeneous Photorhabdus luminescens cell populations
Annals of Microbiology (2022)
-
Horizontal transfer of a plasmid possessing mcr-1 marked with a single nucleotide mutation between Escherichia coli isolates from community residents
BMC Research Notes (2022)
-
Breaking the law: unconventional strategies for antibody diversification
Nature Reviews Immunology (2019)