Key Points
-
Different immune cell subsets use diverse metabolic pathways. In general, inflammatory and suppressive cells each utilize glycolysis and oxidative phosphorylation for distinct purposes.
-
Mitochondrial metabolism produces a variety of signalling molecules (such as mitochondrial reactive oxygen species (mROS) and acetyl-CoA) that can drive changes in immune cell function through the regulation of transcription factors and epigenetics.
-
mROS are produced by the mitochondrial electron transport chain as a signal to increase interleukin-2 (IL-2) production in T cells and IL-1β production in macrophages.
-
Acetyl-CoA produced by fatty acid oxidation or pyruvate oxidation in mitochondria can be transported by the citrate shuttle into the cytoplasm, where it can be used for fatty acid synthesis or acetylation reactions. These pathways have crucial roles in immune cell function.
-
M1 macrophages use an altered tricarboxylic acid (TCA) cycle and reverse electron transport to drive inflammation through increased succinate and mROS levels. M2 macrophages have an intact TCA cycle and require the function of the hexosamine branch of glycolysis.
-
Cellular metabolism can be altered by drugs that target mitochondria, such as metformin and mitochondria-targeted antioxidants.
Abstract
Mitochondria are important signalling organelles, and they dictate immunological fate. From T cells to macrophages, mitochondria form the nexus of the various metabolic pathways that define each immune cell subset. In this central position, mitochondria help to control the various metabolic decision points that determine immune cell function. In this Review, we discuss how mitochondrial metabolism varies across different immune cell subsets, how metabolic signalling dictates cell fate and how this signalling could potentially be targeted therapeutically.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Wang, R. et al. The transcription factor Myc controls metabolic reprogramming upon T lymphocyte activation. Immunity 35, 871–882 (2011).
Carr, E. L. et al. Glutamine uptake and metabolism are coordinately regulated by ERK/MAPK during T lymphocyte activation. J. Immunol. 185, 1037–1044 (2010).
Frauwirth, K. A. et al. The CD28 signaling pathway regulates glucose metabolism. Immunity 16, 769–777 (2002).
Michalek, R. D. et al. Estrogen-related receptor-α is a metabolic regulator of effector T-cell activation and differentiation. Proc. Natl Acad. Sci. USA 108, 18348–18353 (2011).
Blagih, J. et al. The energy sensor AMPK regulates T cell metabolic adaptation and effector responses in vivo. Immunity 42, 41–54 (2015).
Ma, E. H. et al. Serine is an essential metabolite for effector T cell expansion. Cell Metab. 25, 345–357 (2017).
Ron-Harel, N. et al. Mitochondrial biogenesis and proteome remodeling promote one-carbon metabolism for T cell activation. Cell Metab. 24, 104–117 (2016).
Chang, C. H. et al. Posttranscriptional control of T cell effector function by aerobic glycolysis. Cell 153, 1239–1251 (2013).
Sena, L. A. et al. Mitochondria are required for antigen-specific T cell activation through reactive oxygen species signaling. Immunity 38, 225–236 (2013). A paper that demonstrates the essential function of mROS that are generated by complex III in T cell activation and antigen-driven responses in vivo.
Macintyre, A. N. et al. The glucose transporter Glut1 is selectively essential for CD4 T cell activation and effector function. Cell Metab. 20, 61–72 (2014).
Kaminski, M. M. et al. T cell activation is driven by an ADP-dependent glucokinase linking enhanced glycolysis with mitochondrial reactive oxygen species generation. Cell Rep. 2, 1300–1315 (2012).
Quintana, A. et al. T cell activation requires mitochondrial translocation to the immunological synapse. Proc. Natl Acad. Sci. USA 104, 14418–14423 (2007).
Tan, H. et al. Integrative proteomics and phosphoproteomics profiling reveals dynamic signaling networks and bioenergetics pathways underlying T cell activation. Immunity 46, 488–503 (2017).
Zhang, B. et al. MicroRNA-23a curbs necrosis during early T cell activation by enforcing intracellular reactive oxygen species equilibrium. Immunity 44, 568–581 (2016).
Gerriets, V. A. et al. Metabolic programming and PDHK1 control CD4+ T cell subsets and inflammation. J. Clin. Invest. 125, 194–207 (2015).
Michalek, R. D. et al. Cutting edge: distinct glycolytic and lipid oxidative metabolic programs are essential for effector and regulatory CD4+ T cell subsets. J. Immunol. 186, 3299–3303 (2011).
Gerriets, V. A. et al. Foxp3 and Toll-like receptor signaling balance Treg cell anabolic metabolism for suppression. Nat. Immunol. 17, 1459–1466 (2016).
Dang, E. V. et al. Control of TH17/Treg balance by hypoxia-inducible factor 1. Cell 146, 772–784 (2011).
Shi, L. Z. et al. HIF1α-dependent glycolytic pathway orchestrates a metabolic checkpoint for the differentiation of TH17 and Treg cells. J. Exp. Med. 208, 1367–1376 (2011).
Lee, J. H., Elly, C., Park, Y. & Liu, Y. C. E3 ubiquitin ligase VHL regulates hypoxia-inducible factor-1α to maintain regulatory T cell stability and suppressive capacity. Immunity 42, 1062–1074 (2015).
MacIver, N. J. et al. The liver kinase B1 is a central regulator of T cell development, activation, and metabolism. J. Immunol. 187, 4187–4198 (2011).
Shriver, L. P. & Manchester, M. Inhibition of fatty acid metabolism ameliorates disease activity in an animal model of multiple sclerosis. Sci. Rep. 1, 79 (2011).
Berod, L. et al. De novo fatty acid synthesis controls the fate between regulatory T and T helper 17 cells. Nat. Med. 20, 1327–1333 (2014).
Newton, R., Priyadharshini, B. & Turka, L. A. Immunometabolism of regulatory T cells. Nat. Immunol. 17, 618–625 (2016).
Peng, M. et al. Aerobic glycolysis promotes T helper 1 cell differentiation through an epigenetic mechanism. Science 354, 481–484 (2016). An excellent demonstration of how mitochondria-generated citrate pools control histone acetylation and thus affect T cell function
Buck, M. D. et al. Mitochondrial dynamics controls T cell fate through metabolic programming. Cell 166, 63–76 (2016). A key paper demonstrating that mitochondrial structure and form dictate T cell function.
Pearce, E. L. et al. Enhancing CD8 T-cell memory by modulating fatty acid metabolism. Nature 460, 103–107 (2009).
Sukumar, M. et al. Inhibiting glycolytic metabolism enhances CD8+ T cell memory and antitumor function. J. Clin. Invest. 123, 4479–4488 (2013).
van der Windt, G. J. et al. Mitochondrial respiratory capacity is a critical regulator of CD8+ T cell memory development. Immunity 36, 68–78 (2012).
O'Sullivan, D. et al. Memory CD8+ T cells use cell-intrinsic lipolysis to support the metabolic programming necessary for development. Immunity 41, 75–88 (2014).
Cui, G. et al. IL-7-induced glycerol transport and TAG synthesis promotes memory CD8+ T cell longevity. Cell 161, 750–761 (2015).
Pan, Y. et al. Survival of tissue-resident memory T cells requires exogenous lipid uptake and metabolism. Nature 543, 252–256 (2017).
Phan, A. T. et al. Constitutive glycolytic metabolism supports CD8+ T cell effector memory differentiation during viral infection. Immunity 45, 1024–1037 (2016).
Tyrakis, P. A. et al. S-2-hydroxyglutarate regulates CD8+ T-lymphocyte fate. Nature 540, 236–241 (2016). The first demonstration that S -2HG can control immune responses through epigenetic regulation.
Intlekofer, A. M. et al. Hypoxia induces production of L-2-hydroxyglutarate. Cell Metab. 22, 304–311 (2015).
Mullen, A. R. et al. Oxidation of α-ketoglutarate is required for reductive carboxylation in cancer cells with mitochondrial defects. Cell Rep. 7, 1679–1690 (2014).
Champagne, D. P. et al. Fine-tuning of CD8+ T cell mitochondrial metabolism by the respiratory chain repressor MCJ dictates protection to influenza virus. Immunity 44, 1299–1311 (2016).
Doughty, C. A. et al. Antigen receptor-mediated changes in glucose metabolism in B lymphocytes: role of phosphatidylinositol 3-kinase signaling in the glycolytic control of growth. Blood 107, 4458–4465 (2006).
Garcia-Manteiga, J. M. et al. Metabolomics of B to plasma cell differentiation. J. Proteome Res. 10, 4165–4176 (2011).
Le, A. et al. Glucose-independent glutamine metabolism via TCA cycling for proliferation and survival in B cells. Cell Metab. 15, 110–121 (2012).
Wu, J. L. et al. Temporal regulation of Lsp1 O-GlcNAcylation and phosphorylation during apoptosis of activated B cells. Nat. Commun. 7, 12526 (2016).
Lam, W. Y. et al. Mitochondrial pyruvate import promotes long-term survival of antibody-secreting plasma cells. Immunity 45, 60–73 (2016).
Capasso, M. et al. HVCN1 modulates BCR signal strength via regulation of BCR-dependent generation of reactive oxygen species. Nat. Immunol. 11, 265–272 (2010).
Singh, D. K. et al. The strength of receptor signaling is centrally controlled through a cooperative loop between Ca2+ and an oxidant signal. Cell 121, 281–293 (2005).
Wheeler, M. L. & Defranco, A. L. Prolonged production of reactive oxygen species in response to B cell receptor stimulation promotes B cell activation and proliferation. J. Immunol. 189, 4405–4416 (2012).
Jang, K. J. et al. Mitochondrial function provides instructive signals for activation-induced B-cell fates. Nat. Commun. 6, 6750 (2015). A paper that describes the essential role of mROS in determining B cell fate.
Heinemann, I. U., Jahn, M. & Jahn, D. The biochemistry of heme biosynthesis. Arch. Biochem. Biophys. 474, 238–251 (2008).
Haschemi, A. et al. The sedoheptulose kinase CARKL directs macrophage polarization through control of glucose metabolism. Cell Metab. 15, 813–826 (2012).
Jin, Z., Wei, W., Yang, M., Du, Y. & Wan, Y. Mitochondrial complex I activity suppresses inflammation and enhances bone resorption by shifting macrophage-osteoclast polarization. Cell Metab. 20, 483–498 (2014).
Vats, D. et al. Oxidative metabolism and PGC-1β attenuate macrophage-mediated inflammation. Cell Metab. 4, 13–24 (2006).
Huang, S. C. et al. Cell-intrinsic lysosomal lipolysis is essential for alternative activation of macrophages. Nat. Immunol. 15, 846–855 (2014).
Huang, S. C. et al. Metabolic reprogramming mediated by the mTORC2–IRF4 signaling axis is essential for macrophage alternative activation. Immunity 45, 817–830 (2016).
Chandel, N. S., Schumacker, P. T. & Arch, R. H. Reactive oxygen species are downstream products of TRAF-mediated signal transduction. J. Biol. Chem. 276, 42728–42736 (2001). An early paper demonstrating that immune receptor signalling is dependent on the ETC.
Hall, C. J. et al. Immunoresponsive gene 1 augments bactericidal activity of macrophage-lineage cells by regulating β-oxidation-dependent mitochondrial ROS production. Cell Metab. 18, 265–278 (2013).
West, A. P. et al. TLR signalling augments macrophage bactericidal activity through mitochondrial ROS. Nature 472, 476–480 (2011). An important paper that links TLR signalling to mROS and shows the role of TLR–mROS signalling in macrophage function.
Bulua, A. C. et al. Mitochondrial reactive oxygen species promote production of proinflammatory cytokines and are elevated in TNFR1-associated periodic syndrome (TRAPS). J. Exp. Med. 208, 519–533 (2011). An important paper demonstrating that mROS are necessary for hyperinflammatory responses in patients that carry mutations in the gene that encodes TNF receptor type 1.
Jha, A. K. et al. Network integration of parallel metabolic and transcriptional data reveals metabolic modules that regulate macrophage polarization. Immunity 42, 419–430 (2015). A key paper demonstrating that pro-inflammatory and anti-inflammatory macrophages have distinct TCA cycles.
Lampropoulou, V. et al. Itaconate links inhibition of succinate dehydrogenase with macrophage metabolic remodeling and regulation of inflammation. Cell Metab. 24, 158–166 (2016). A study that provides genetic evidence that itaconate is necessary for pro-inflammatory macrophage function.
Mills, E. L. et al. Succinate dehydrogenase supports metabolic repurposing of mitochondria to drive inflammatory macrophages. Cell 167, 457–470.e13 (2016). An important paper demonstrating that succinate-dependent mROS generation is necessary for pro-inflammatory macrophage function.
Tannahill, G. M. et al. Succinate is an inflammatory signal that induces IL-1β through HIF-1α. Nature 496, 238–242 (2013).
Palsson-McDermott, E. M. et al. Pyruvate kinase M2 regulates Hif-1α activity and IL-1β induction and is a critical determinant of the Warburg effect in LPS-activated macrophages. Cell Metab. 21, 65–80 (2015).
Tan, Z. et al. Pyruvate dehydrogenase kinase 1 participates in macrophage polarization via regulating glucose metabolism. J. Immunol. 194, 6082–6089 (2015).
Meiser, J. et al. Pro-inflammatory macrophages sustain pyruvate oxidation through pyruvate dehydrogenase for the synthesis of itaconate and to enable cytokine expression. J. Biol. Chem. 291, 3932–3946 (2016).
Michelucci, A. et al. Immune-responsive gene 1 protein links metabolism to immunity by catalyzing itaconic acid production. Proc. Natl Acad. Sci. USA 110, 7820–7825 (2013).
Naujoks, J. et al. IFNs modify the proteome of Legionella-containing vacuoles and restrict infection via IRG1-derived itaconic acid. PLoS Pathog. 12, e1005408 (2016).
Kannan, Y. et al. TPL-2 regulates macrophage lipid metabolism and M2 differentiation to control TH2-mediated immunopathology. PLoS Pathog. 12, e1005783 (2016).
Mounier, R. et al. AMPKα1 regulates macrophage skewing at the time of resolution of inflammation during skeletal muscle regeneration. Cell Metab. 18, 251–264 (2013).
Carroll, K. C., Viollet, B. & Suttles, J. AMPKα1 deficiency amplifies proinflammatory myeloid APC activity and CD40 signaling. J. Leukoc. Biol. 94, 1113–1121 (2013).
Covarrubias, A. J. et al. Akt–mTORC1 signaling regulates Acly to integrate metabolic input to control of macrophage activation. eLife 5, e11612 (2016).
Nomura, M. et al. Fatty acid oxidation in macrophage polarization. Nat. Immunol. 17, 216–217 (2016).
Seth, R. B., Sun, L. & Chen, Z. J. Antiviral innate immunity pathways. Cell Res. 16, 141–147 (2006).
Galluzzi, L., Kepp, O. & Kroemer, G. Mitochondria: master regulators of danger signalling. Nat. Rev. Mol. Cell Biol. 13, 780–788 (2012).
Dostert, C. et al. Innate immune activation through Nalp3 inflammasome sensing of asbestos and silica. Science 320, 674–677 (2008).
Zhou, R., Yazdi, A. S., Menu, P. & Tschopp, J. A role for mitochondria in NLRP3 inflammasome activation. Nature 469, 221–225 (2011). An early paper that links mitochondria to inflammasome activation.
Muruve, D. A. et al. The inflammasome recognizes cytosolic microbial and host DNA and triggers an innate immune response. Nature 452, 103–107 (2008).
Yu, J. et al. Inflammasome activation leads to caspase-1-dependent mitochondrial damage and block of mitophagy. Proc. Natl Acad. Sci. USA 111, 15514–15519 (2014).
Zhong, Z. et al. NF-κB restricts inflammasome activation via elimination of damaged mitochondria. Cell 164, 896–910 (2016).
Wang, X. et al. RNA viruses promote activation of the NLRP3 inflammasome through a RIP1–RIP3–DRP1 signaling pathway. Nat. Immunol. 15, 1126–1133 (2014).
Moriwaki, K. et al. The mitochondrial phosphatase PGAM5 is dispensable for necroptosis but promotes inflammasome activation in macrophages. J. Immunol. 196, 407–415 (2016).
Garaude, J. et al. Mitochondrial respiratory-chain adaptations in macrophages contribute to antibacterial host defense. Nat. Immunol. 17, 1 037–1045 (2016).
Moon, J. S. et al. NOX4-dependent fatty acid oxidation promotes NLRP3 inflammasome activation in macrophages. Nat. Med. 22, 1002–1012 (2016).
Munoz-Planillo, R. et al. K+ efflux is the common trigger of NLRP3 inflammasome activation by bacterial toxins and particulate matter. Immunity 38, 1142–1153 (2013).
Everts, B. et al. TLR-driven early glycolytic reprogramming via the kinases TBK1–IKKɛ supports the anabolic demands of dendritic cell activation. Nat. Immunol. 15, 323–332 (2014).
Everts, B. et al. Commitment to glycolysis sustains survival of NO-producing inflammatory dendritic cells. Blood 120, 1422–1431 (2012).
Krawczyk, C. M. et al. Toll-like receptor-induced changes in glycolytic metabolism regulate dendritic cell activation. Blood 115, 4742–4749 (2010).
Jantsch, J. et al. Hypoxia and hypoxia-inducible factor-1α modulate lipopolysaccharide-induced dendritic cell activation and function. J. Immunol. 180, 4697–4705 (2008).
Ibrahim, J. et al. Dendritic cell populations with different concentrations of lipid regulate tolerance and immunity in mouse and human liver. Gastroenterology 143, 1061–1072 (2012).
Rehman, A. et al. Role of fatty-acid synthesis in dendritic cell generation and function. J. Immunol. 190, 4640–4649 (2013).
Wu, D. et al. Type 1 interferons induce changes in core metabolism that are critical for immune function. Immunity 44, 1325–1336 (2016).
Ferreira, G. B. et al. Vitamin D3 induces tolerance in human dendritic cells by activation of intracellular metabolic pathways. Cell Rep. 10, 711–725 (2015).
Klotz, L. et al. Peroxisome proliferator-activated receptor γ control of dendritic cell function contributes to development of CD4+ T cell anergy. J. Immunol. 178, 2122–2131 (2007).
Szanto, A. et al. STAT6 transcription factor is a facilitator of the nuclear receptor PPARγ-regulated gene expression in macrophages and dendritic cells. Immunity 33, 699–712 (2010).
Del Prete, A. et al. Role of mitochondria and reactive oxygen species in dendritic cell differentiation and functions. Free Radic. Biol. Med. 44, 1443–1451 (2008).
Zaccagnino, P. et al. An active mitochondrial biogenesis occurs during dendritic cell differentiation. Int. J. Biochem. Cell Biol. 44, 1962–1969 (2012).
Scharping, N. E. et al. The tumor microenvironment represses T cell mitochondrial biogenesis to drive intratumoral T cell metabolic insufficiency and dysfunction. Immunity 45, 374–388 (2016).
Sukumar, M. et al. Mitochondrial membrane potential identifies cells with enhanced stemness for cellular therapy. Cell Metab. 23, 63–76 (2016).
Chang, C. H. et al. Metabolic competition in the tumor microenvironment is a driver of cancer progression. Cell 162, 1229–1241 (2015).
Ho, P. C. et al. Phosphoenolpyruvate is a metabolic checkpoint of anti-tumor T cell responses. Cell 162, 1217–1228 (2015).
Mamlouk, S. et al. Loss of prolyl hydroxylase-2 in myeloid cells and T-lymphocytes impairs tumor development. Int. J. Cancer 134, 849–858 (2014).
Hatfield, S. M. et al. Immunological mechanisms of the antitumor effects of supplemental oxygenation. Sci. Transl Med. 7, 277ra30 (2015).
Mellor, A. L. & Munn, D. H. IDO expression by dendritic cells: tolerance and tryptophan catabolism. Nat. Rev. Immunol. 4, 762–774 (2004).
Geiger, R. et al. L-Arginine modulates T cell metabolism and enhances survival and anti-tumor activity. Cell 167, 829–842.e13 (2016).
Fletcher, M. et al. L-Arginine depletion blunts antitumor T-cell responses by inducing myeloid-derived suppressor cells. Cancer Res. 75, 275–283 (2015).
Haas, R. et al. Lactate regulates metabolic and pro-inflammatory circuits in control of T cell migration and effector functions. PLoS Biol. 13, e1002202 (2015).
Colegio, O. R. et al. Functional polarization of tumour-associated macrophages by tumour-derived lactic acid. Nature 513, 559–563 (2014).
Scharping, N. E., Menk, A. V., Whetstone, R. D., Zeng, X. & Delgoffe, G. M. Efficacy of PD-1 blockade is potentiated by metformin-induced reduction of tumor hypoxia. Cancer Immunol. Res. 5, 9–16 (2017). A recent paper that shows how metformin can be used in conjunction with checkpoint blockade therapy to augment tumour immunity.
Wheaton, W. W. et al. Metformin inhibits mitochondrial complex I of cancer cells to reduce tumorigenesis. eLife 3, e02242 (2014).
Fresnak, A. D., June, C. H. & Levine, B. L. Engineered T cells: the promise and challenges of cancer immunotherapy. Nat. Rev. Cancer 16, 566–581 (2016).
Kawalekar, O. U. et al. Distinct signaling of coreceptors regulates specific metabolism pathways and impacts memory development in CAR T cells. Immunity 44, 380–390 (2016).
Tsoyi, K. et al. Metformin inhibits HMGB1 release in LPS-treated RAW 264.7 cells and increases survival rate of endotoxaemic mice. Br. J. Pharmacol. 162, 1498–1508 (2011).
Forslund, K. et al. Disentangling type 2 diabetes and metformin treatment signatures in the human gut microbiota. Nature 528, 262–266 (2015).
Lee, H. & Ko, G. Effect of metformin on metabolic improvement and gut microbiota. Appl. Environ. Microbiol. 80, 5935–5943 (2014).
Anisimov, V. N. et al. Effect of metformin on life span and on the development of spontaneous mammary tumors in HER-2/neu transgenic mice. Exp. Gerontol. 40, 685–693 (2005).
Martin-Montalvo, A. et al. Metformin improves healthspan and lifespan in mice. Nat. Commun. 4, 2192 (2013).
Singhal, A. et al. Metformin as adjunct antituberculosis therapy. Sci. Transl Med. 6, 263ra159 (2014).
Barzilai, N. et al. Metformin as a tool to target aging. Cell Metabolism 23, 1060–1065 (2016).
Kelly, B., Tannahill, G. M., Murphy, M. P. & O'Neill, L. A. Metformin inhibits the production of reactive oxygen species from NADH:ubiquinone oxidoreductase to limit induction of interleukin-1β (IL-1β) and boosts interleukin-10 (IL-10) in lipopolysaccharide (LPS)-activated macrophages. J. Biol. Chem. 290, 20348–20359 (2015).
Yin, Y. et al. Normalization of CD4+ T cell metabolism reverses lupus. Sci. Transl Med. 7, 274ra18 (2015).
Zhao, W. et al. The peroxisome proliferator-activated receptor γ agonist pioglitazone improves cardiometabolic risk and renal inflammation in murine lupus. J. Immunol. 183, 2729–2740 (2009).
Johnson, K. M. et al. Identification and validation of the mitochondrial F1F0-ATPase as the molecular target of the immunomodulatory benzodiazepine Bz-423. Chem. Biol. 12, 485–496 (2005).
Lee, C. F. et al. Preventing allograft rejection by targeting immune metabolism. Cell Rep. 13, 760–770 (2015).
Gatza, E. et al. Manipulating the bioenergetics of alloreactive T cells causes their selective apoptosis and arrests graft-versus-host disease. Sci. Transl Med. 3, 67ra8 (2011).
Orr, A. L. et al. Suppressors of superoxide production from mitochondrial complex III. Nat. Chem. Biol. 11, 834–836 (2015).
Brand, M. D. et al. Suppressors of superoxide-H2O2 production at site IQ of mitochondrial complex I protect against stem cell hyperplasia and ischemia–reperfusion injury. Cell Metab. 24, 582–592 (2016).
Chouchani, E. T. et al. Mitochondrial ROS regulate thermogenic energy expenditure and sulfenylation of UCP1. Nature 532, 112–116 (2016).
Acknowledgements
This work was supported by the US National Institutes of Health (R35 CA197532, PO1 AG04966502 and PO1 HL071643 to N.S.C.; T32 CA9560 to M.M.M.; and T32 T32HL076139 to S.E.W.).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Glossary
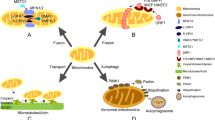
- Mitochondrial permeability transition pores
-
High-conductance inner mitochondrial membrane channels. Persistent opening of these pores irreversibly commits cells to death by causing mitochondrial depolarization (which blocks oxidative phosphorylation and reactive oxygen species production), matrix swelling and cristae unfolding, and results in the release of stored calcium+ and of apoptogenic proteins.
- Mitophagy
-
A special form of autophagy, in which mitochondria (in a damaged or depolarized state) are engulfed by autophagosomes and degraded.
- Autophagy
-
A cellular process, by which cytoplasmic organelles and macromolecular complexes are engulfed by double membrane-bound vesicles for delivery to lysosomes and subsequent degradation. This process is involved in the constitutive turnover of proteins and organelles, and is central to cellular activities that maintain a balance between the synthesis and breakdown of various proteins.
- Pentose phosphate pathway
-
(PPP). A pathway that uses glucose to generate NADPH and pentose sugars (such as ribose). The first (oxidative) phase converts glucose-6-phosphate to ribulose-5-phosphate and generates NADPH. The second (non-oxidative) phase synthesizes other sugars from ribulose-5-phosphate.
Rights and permissions
About this article
Cite this article
Mehta, M., Weinberg, S. & Chandel, N. Mitochondrial control of immunity: beyond ATP. Nat Rev Immunol 17, 608–620 (2017). https://doi.org/10.1038/nri.2017.66
Published:
Issue Date:
DOI: https://doi.org/10.1038/nri.2017.66
This article is cited by
-
Stem Leydig cells support macrophage immunological homeostasis through mitochondrial transfer in mice
Nature Communications (2024)
-
Bioorthogonal photocatalytic proximity labeling in primary living samples
Nature Communications (2024)
-
Long-chain acyl-CoA synthetase 4-mediated mitochondrial fatty acid metabolism and dendritic cell antigen presentation
Inflammation Research (2024)
-
Immune response stability to the SARS-CoV-2 mRNA vaccine booster is influenced by differential splicing of HLA genes
Scientific Reports (2024)
-
Inflamed macrophages sans mitochondrial pyruvate carrier?
Nature Metabolism (2023)