Key Points
-
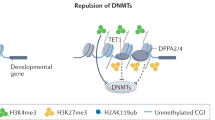
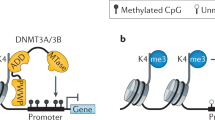
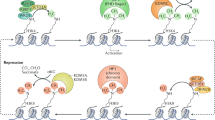
The principal epigenetic mechanisms by which tissue-specific gene-expression patterns and global gene silencing are established and maintained are chromatin modification — including processes such as DNA methylation, histone modification (acetylation, phosphorylation, methylation and ubiquitylation) — and chromatin remodelling.
-
New data show how DNA methylation and histone modification integrate with each other in the epigenetic regulation of genomic imprinting, X inactivation and genome reprogramming.
-
DNA methylation is the heritable epigenetic mark for genomic imprinting, which is established in the developing oocyte and is essential for maintaining the mono-allelic expression of imprinted genes. DNA methyltransferase Dnmt3a and an associated protein Dnmt3l, and the histone methyltransferases Suv39h1 and Suv39h2, are required for spermatogenesis.
-
DNA methylation undergoes dynamic changes during early embryonic development, and genome-wide demethylation (after fertilization) and remethylation (after the implantation of a blastocyst) have an essential role in genome reprogramming.
-
X inactivation is regulated by several factors, including non-coding RNAs (Xist and Tsix), DNA methylation, histone acetylation and deacetylation, histone methylation and Polycomb-group proteins.
-
Reprogramming of somatic cell nuclei in enucleated oocytes by nuclear transfer resets the epigenetic programme for embryonic development.
-
Studies of the epigenetic regulation of chromatin and gene expression in development will help to elucidate the underlying causes of certain human diseases. A greater understanding of these processes will also aid research into the clinical application of pluripotent stem cells or progenitor cells in cell-replacement therapy and into new drugs that target epigenetic regulators for use in treating cancer and certain developmental disorders.
Abstract
The developmental programme of embryogenesis is controlled by both genetic and epigenetic mechanisms. An emerging theme from recent studies is that the regulation of higher-order chromatin structures by DNA methylation and histone modification is crucial for genome reprogramming during early embryogenesis and gametogenesis, and for tissue-specific gene expression and global gene silencing. Disruptions to chromatin modification can lead to the dysregulation of developmental processes, such as X-chromosome inactivation and genomic imprinting, and to various diseases. Understanding the process of epigenetic reprogramming in development is important for studies of cloning and the clinical application of stem-cell therapy.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Bird, A. DNA methylation patterns and epigenetic memory. Genes Dev. 16, 6–21 (2002).
Jenuwein, T. & Allis, C. D. Translating the histone code. Science 293, 1074–1080 (2001).
Zhang, Y. & Reinberg, D. Transcription regulation by histone methylation: interplay between different covalent modifications of the core histone tails. Genes Dev. 15, 2343–2360 (2001).
Narlikar, G. J., Fan, H. Y. & Kingston, R. E. Cooperation between complexes that regulate chromatin structure and transcription. Cell 108, 475–487 (2002).
Bestor, T. H. Activation of mammalian DNA methyltransferase by cleavage of a Zn binding regulatory domain. EMBO J. 11, 2611–2617 (1992).
Li, E., Bestor, T. H. & Jaenisch, R. Targeted mutation of the DNA methyltransferase gene results in embryonic lethality. Cell 69, 915–926 (1992).
Okano, M., Bell, D. W., Haber, D. A. & Li, E. DNA methyltransferases Dnmt3a and Dnmt3b are essential for de novo methylation and mammalian development. Cell 99, 247–257 (1999).This paper provided the first genetic evidence that the Dnmt3 family of methyltransferases is responsible for de novo methylation in mouse ES cells and during development.
Lyko, F. et al. Mammalian (cytosine-5) methyltransferases cause genomic DNA methylation and lethality in Drosophila. Nature Genet. 23, 363–366 (1999).
Hsieh, C. L. In vivo activity of murine de novo methyltransferases, Dnmt3a and Dnmt3b. Mol. Cell. Biol. 19, 8211–8218 (1999).
Burgers, W. A., Fuks, F. & Kouzarides, T. DNA methyltransferases get connected to chromatin. Trends Genet. 18, 275–277 (2002).
Takizawa, T. et al. DNA methylation is a critical cell-intrinsic determinant of astrocyte differentiation in the fetal brain. Dev. Cell 1, 749–758 (2001).This paper shows that the demethylation of a promoter element induces tissue-specific gene expression during cellular differentiation.
Nan, X. et al. Transcriptional repression by the methyl-CpG-binding protein MeCP2 involves a histone deacetylase complex. Nature 393, 386–389 (1998).
Jones, P. L. et al. Methylated DNA and MeCP2 recruit histone deacetylase to repress transcription. Nature Genet. 19, 187–191 (1998).References 12 and 13 provided the first molecular evidence linking DNA methylation to histone deacetylation in transcription repression.
Zhang, Y. et al. Analysis of the NuRD subunits reveals a histone deacetylase core complex and a connection with DNA methylation. Genes Dev. 13, 1924–1935 (1999).
Wade, P. A. et al. Mi-2 complex couples DNA methylation to chromatin remodelling and histone deacetylation. Nature Genet. 23, 62–66 (1999).
Ng, H. H. et al. MBD2 is a transcriptional repressor belonging to the MeCP1 histone deacetylase complex. Nature Genet. 23, 58–61 (1999).
Feng, Q. & Zhang, Y. The MeCP1 complex represses transcription through preferential binding, remodeling, and deacetylating methylated nucleosomes. Genes Dev. 15, 827–832 (2001).This study shows how MBD2 might recruit the NuRD remodelling complex to methylated chromatin to remodel the chromatin and to silence transcription.
Murrell, A. et al. An intragenic methylated region in the imprinted Igf2 gene augments transcription. EMBO Rep. 2, 1101–1106 (2001).
Eden, S. et al. An upstream repressor element plays a role in Igf2 imprinting. EMBO J. 20, 3518–3525 (2001).
Strahl, B. D. & Allis, C. D. The language of covalent histone modifications. Nature 403, 41–45 (2000).
Tamaru, H. & Selker, E. U. A histone H3 methyltransferase controls DNA methylation in Neurospora crassa. Nature 414, 277–283 (2001).
Jackson, J. P., Lindroth, A. M., Cao, X. & Jacobsen, S. E. Control of CpNpG DNA methylation by the KRYPTONITE histone H3 methyltransferase. Nature 416, 556–560 (2002).References 21 and 22 provide molecular and genetic evidence that histone H3-K9 methylation is required for DNA methylation.
Jeddeloh, J. A., Stokes, T. L. & Richards, E. J. Maintenance of genomic methylation requires a SWI2/SNF2-like protein. Nature Genet. 22, 94–97 (1999).
Gendrel, A. V., Lippman, Z., Yordan, C., Colot, V. & Martienssen, R. Dependence of heterochromatic histone H3 methylation patterns on the Arabidopsis gene DDM1. Science 20 (in the press).
Gibbons, R. J. et al. Mutations in ATRX, encoding a SWI/SNF-like protein, cause diverse changes in the pattern of DNA methylation. Nature Genet. 24, 368–371 (2000).
Dennis, K., Fan, T., Geiman, T., Yan, Q. & Muegge, K. Lsh, a member of the SNF2 family, is required for genome-wide methylation. Genes Dev. 15, 2940–2944 (2001).References 23–26 show that SWI/SNF-like chromatin-remodelling proteins are intimately involved in regulating DNA methylation and histone methylation.
Bultman, S. et al. A Brg1 null mutation in the mouse reveals functional differences among mammalian SWI/SNF complexes. Mol. Cell 6, 1287–1295 (2000).
Hendrich, B., Guy, J., Ramsahoye, B., Wilson, V. A. & Bird, A. Closely related proteins MBD2 and MBD3 play distinctive but interacting roles in mouse development. Genes Dev. 15, 710–723 (2001).
O'Carroll, D. et al. The polycomb-group gene Ezh2 is required for early mouse development. Mol. Cell. Biol. 21, 4330–4336 (2001).
Donohoe, M. E. et al. Targeted disruption of mouse Yin Yang 1 transcription factor results in peri-implantation lethality. Mol. Cell. Biol. 19, 7237–7244 (1999).
Faust, C., Lawson, K. A., Schork, N. J., Thiel, B. & Magnuson, T. The Polycomb-group gene eed is required for normal morphogenetic movements during gastrulation in the mouse embryo. Development 125, 4495–4506 (1998).
Lagger, G. et al. Essential function of histone deacetylase 1 in proliferation control and CDK inhibitor repression. EMBO J. 21, 2672–2681 (2002).This paper provides genetic evidence that histone deacetylation is essential for embryonic development.
Guy, J., Hendrich, B., Holmes, M., Martin, J. E. & Bird, A. A mouse Mecp2-null mutation causes neurological symptoms that mimic Rett syndrome. Nature Genet. 27, 322–326 (2001).
Chen, R. Z., Akbarian, S., Tudor, M. & Jaenisch, R. Deficiency of methyl-CpG binding protein-2 in CNS neurons results in a Rett-like phenotype in mice. Nature Genet. 27, 327–331 (2001).
Shahbazian, M. D. et al. Mice with truncated MeCP2 recapitulate many Rett syndrome features and display hyperacetylation of histone H3. Neuron 35, 243–254 (2002).
Peters, A. H. et al. Loss of the Suv39h histone methyltransferases impairs mammalian heterochromatin and genome stability. Cell 107, 323–337 (2001).
Tachibana, M. et al. G9a histone methyltransferase plays a dominant role in euchromatic histone H3 lysine 9 methylation and is essential for early embryogenesis. Genes Dev. 16, 1779–1791 (2002).References 36 and 37 show that two histone H3-K9 methyltransferases have distinct functions in mouse development. Suv39h methylates histones in heterochromatin and is dispensable for early development, whereas G9a methylates histones in euchromatin and is essential for early development.
Lachner, M., O'Carroll, D., Rea, S., Mechtler, K. & Jenuwein, T. Methylation of histone H3 lysine 9 creates a binding site for HP1 proteins. Nature 410, 116–120 (2001).
Bannister, A. J. et al. Selective recognition of methylated lysine 9 on histone H3 by the HP1 chromo domain. Nature 410, 120–124 (2001).
Jones, D. O., Cowell, I. G. & Singh, P. B. Mammalian chromodomain proteins: their role in genome organisation and expression. Bioessays 22, 124–137 (2000).
Surani, M. A., Barton, S. C. & Norris, M. L. Development of reconstituted mouse eggs suggests imprinting of the genome during gametogenesis. Nature 308, 548–550 (1984).
McGrath, J. & Solter, D. Completion of mouse embryogenesis requires both the maternal and paternal genomes. Cell 37, 179–183 (1984).
Reik, W. & Walter, J. Genomic imprinting: parental influence on the genome. Nature Rev. Genet. 2, 21–32 (2001).
Tremblay, K. D., Saam, J. R., Ingram, R. S., Tilghman, S. M. & Bartolomei, M. S. A paternal-specific methylation imprint marks the alleles of the mouse H19 gene. Nature Genet. 9, 407–413 (1995).
Thorvaldsen, J. L., Duran, K. L. & Bartolomei, M. S. Deletion of the H19 differentially methylated domain results in loss of imprinted expression of H19 and Igf2. Genes Dev. 12, 3693–3702 (1998).
Yoon, B. J. et al. Regulation of DNA methylation of Rasgrf1. Nature Genet. 30, 92–96 (2002).
Stoger, R. et al. Maternal-specific methylation of the imprinted mouse Igf2r locus identifies the expressed locus as carrying the imprinting signal. Cell 73, 61–71 (1993).
Wutz, A. et al. Imprinted expression of the Igf2r gene depends on an intronic CpG island. Nature 389, 745–749 (1997).
Shemer, R., Birger, Y., Riggs, A. D. & Razin, A. Structure of the imprinted mouse Snrpn gene and establishment of its parental-specific methylation pattern. Proc. Natl Acad. Sci. USA 94, 10267–10272 (1997).
Lee, J. et al. Erasing genomic imprinting memory in mouse clone embryos produced from day 11.5 primordial germ cells. Development 129, 1807–1817 (2002).
Hajkova, P. et al. Epigenetic reprogramming in mouse primordial germ cells. Mech. Dev. (in the press).References 50 and 51 defined the embryonic stage when imprinting marks are erased in the germ cells.
Bourc'his, D., Xu, G. L., Lin, C. S., Bollman, B. & Bestor, T. H. Dnmt3L and the establishment of maternal genomic imprints. Science 294, 2536–2539 (2001).
Hata, K., Okano, M., Lei, H. & Li, E. Dnmt3L cooperates with the Dnmt3 family of de novo DNA methyltransferases to establish maternal imprints in mice. Development 129, 1983–1993 (2002).
Howell, C. Y. et al. Genomic imprinting disrupted by a maternal effect mutation in the Dnmt1 gene. Cell 104, 829–838 (2001).References 52 – 54 provide genetic evidence that Dnmt3a, Dnmt3b and Dnmt3l but not Dnmt1 are responsible for establishing the imprinting marks through de novo methylation in oocytes.
Hazzouri, M. et al. Regulated hyperacetylation of core histones during mouse spermatogenesis: involvement of histone deacetylases. Eur. J. Cell Biol. 79, 950–960 (2000).
Mayer, W., Niveleau, A., Walter, J., Fundele, R. & Haaf, T. Demethylation of the zygotic paternal genome. Nature 403, 501–502 (2000).
Oswald, J. et al. Active demethylation of the paternal genome in the mouse zygote. Curr. Biol. 10, 475–478 (2000).
Howlett, S. K. & Reik, W. Methylation levels of maternal and paternal genomes during preimplantation development. Development 113, 119–127 (1991).
Kafri, T. et al. Developmental pattern of gene-specific DNA methylation in the mouse embryo and germ line. Genes Dev. 6, 705–714 (1992).
Rougier, N. et al. Chromosome methylation patterns during mammalian preimplantation development. Genes Dev. 12, 2108–2113 (1998).
Santos, F., Hendrich, B., Reik, W. & Dean, W. Dynamic reprogramming of DNA methylation in the early mouse embryo. Dev. Biol. 241, 172–182 (2002).
Chapman, V., Forrester, L., Sanford, J., Hastie, N. & Rossant, J. Cell lineage-specific undermethylation of mouse repetitive DNA. Nature 307, 284–286 (1984).
Rossant, J., Sanford, J. P., Chapman, V. M. & Andrews, G. K. Undermethylation of structural gene sequences in extraembryonic lineages of the mouse. Dev. Biol. 117, 567–573 (1986).
Lei, H. et al. De novo DNA cytosine methyltransferase activities in mouse embryonic stem cells. Development 122, 3195–3205 (1996).
Li, E., Beard, C. & Jaenisch, R. Role for DNA methylation in genomic imprinting. Nature 366, 362–365 (1993).
Beard, C., Li, E. & Jaenisch, R. Loss of methylation activates Xist in somatic but not in embryonic cells. Genes Dev. 9, 2325–2334 (1995).
Caspary, T., Cleary, M. A., Baker, C. C., Guan, X. J. & Tilghman, S. M. Multiple mechanisms regulate imprinting of the mouse distal chromosome 7 gene cluster. Mol. Cell. Biol. 18, 3466–3474 (1998).
Walsh, C. P., Chaillet, J. R. & Bestor, T. H. Transcription of IAP endogenous retroviruses is constrained by cytosine methylation. Nature Genet. 20, 116–117 (1998).
Liang, G. et al. Cooperativity between DNA methyltransferases in the maintenance methylation of repetitive elements. Mol. Cell. Biol. 22, 480–491 (2002).
Rhee, I. et al. DNMT1 and DNMT3b cooperate to silence genes in human cancer cells. Nature 416, 552–556 (2002).
McDowell, T. L. et al. Localization of a putative transcriptional regulator (ATRX) at pericentromeric heterochromatin and the short arms of acrocentric chromosomes. Proc. Natl Acad. Sci. USA 96, 13983–13988 (1999).
Gregory, R. I. et al. DNA methylation is linked to deacetylation of histone H3, but not H4, on the imprinted genes Snrpn and U2af1-rs1. Mol. Cell. Biol. 21, 5426–5436 (2001).
Grandjean, V., O'Neill, L., Sado, T., Turner, B. & Ferguson-Smith, A. Relationship between DNA methylation, histone H4 acetylation and gene expression in the mouse imprinted Igf2-H19 domain. FEBS Lett. 488, 165–169 (2001).
Sleutels, F., Zwart, R. & Barlow, D. P. The non-coding Air RNA is required for silencing autosomal imprinted genes. Nature 415, 810–813 (2002).This study shows that a non-coding RNA is involved in regulating genomic imprinting.
Avner, P. & Heard, E. X-chromosome inactivation: counting, choice and initiation. Nature Rev. Genet. 2, 59–67 (2001).
Penny, G. D., Kay, G. F., Sheardown, S. A., Rastan, S. & Brockdorff, N. Requirement for Xist in X chromosome inactivation. Nature 379, 131–137 (1996).
Marahrens, Y., Panning, B., Dausman, J., Strauss, W. & Jaenisch, R. Xist-deficient mice are defective in dosage compensation but not spermatogenesis. Genes Dev. 11, 156–166 (1997).
Wutz, A. & Jaenisch, R. A shift from reversible to irreversible X inactivation is triggered during ES cell differentiation. Mol. Cell 5, 695–705 (2000).
Lee, J. T., Davidow, L. S. & Warshawsky, D. Tsix, a gene antisense to Xist at the X-inactivation centre. Nature Genet. 21, 400–404 (1999).
Luikenhuis, S., Wutz, A. & Jaenisch, R. Antisense transcription through the Xist locus mediates Tsix function in embryonic stem cells. Mol. Cell. Biol. 21, 8512–8520 (2001).
Sado, T., Wang, Z., Sasaki, H. & Li, E. Regulation of imprinted X-chromosome inactivation in mice by Tsix. Development 128, 1275–1286 (2001).
Stavropoulos, N., Lu, N. & Lee, J. T. A functional role for Tsix transcription in blocking Xist RNA accumulation but not in X-chromosome choice. Proc. Natl Acad. Sci. USA 98, 10232–10237 (2001).
Lee, J. T. Disruption of imprinted X inactivation by parent-of-origin effects at Tsix. Cell 103, 17–27 (2000).
Wutz, A., Rasmussen, T. P. & Jaenisch, R. Chromosomal silencing and localization are mediated by different domains of Xist RNA. Nature Genet. 30, 167–174 (2002).
Beletskii, A., Hong, Y. K., Pehrson, J., Egholm, M. & Strauss, W. M. PNA interference mapping demonstrates functional domains in the noncoding RNA Xist. Proc. Natl Acad. Sci. USA 98, 9215–9220 (2001).
Boggs, B. A. et al. Differentially methylated forms of histone H3 show unique association patterns with inactive human X chromosomes. Nature Genet. 30, 73–76 (2002).
Heard, E. et al. Methylation of histone H3 at Lys-9 is an early mark on the X chromosome during X inactivation. Cell 107, 727–738 (2001).References 86 and 87 show that H3-K9 methylation correlates with the inactive X, whereas H3-K4 methylation correlates with the active X. H3-K9 methylation is also an early event in the initiation of X inactivation.
Sado, T. et al. X inactivation in the mouse embryo deficient for Dnmt1: distinct effect of hypomethylation on imprinted and random X inactivation. Dev. Biol. 225, 294–303 (2000).
Brown, C. J. & Willard, H. F. The human X-inactivation centre is not required for maintenance of X-chromosome inactivation. Nature 368, 154–156 (1994).
Csankovszki, G., Panning, B., Bates, B., Pehrson, J. R. & Jaenisch, R. Conditional deletion of Xist disrupts histone macroH2A localization but not maintenance of X inactivation. Nature Genet. 22, 323–324 (1999).
Hansen, R. S. et al. Escape from gene silencing in ICF syndrome: evidence for advanced replication time as a major determinant. Hum. Mol. Genet. 9, 2575–2587 (2000).
Jeppesen, P. & Turner, B. M. The inactive X chromosome in female mammals is distinguished by a lack of histone H4 acetylation, a cytogenetic marker for gene expression. Cell 74, 281–289 (1993).
Csankovszki, G., Nagy, A. & Jaenisch, R. Synergism of Xist RNA, DNA methylation, and histone hypoacetylation in maintaining X chromosome inactivation. J. Cell Biol. 153, 773–784 (2001).
Wang, J. et al. Imprinted X inactivation maintained by a mouse Polycomb group gene. Nature Genet. 28, 371–375 (2001).This work shows that a Polycomb group gene is required for maintaining imprinted X inactivation.
Wilmut, I., Schnieke, A. E., McWhir, J., Kind, A. J. & Campbell, K. H. Viable offspring derived from fetal and adult mammalian cells. Nature 385, 810–813 (1997).
Wakayama, T., Perry, A. C., Zuccotti, M., Johnson, K. R. & Yanagimachi, R. Full-term development of mice from enucleated oocytes injected with cumulus cell nuclei. Nature 394, 369–374 (1998).
Tanaka, S. et al. Placentomegaly in cloned mouse concepti caused by expansion of the spongiotrophoblast layer. Biol. Reprod. 65, 1813–1821 (2001).
Tamashiro, K. L. et al. Cloned mice have an obese phenotype not transmitted to their offspring. Nature Med. 8, 262–267 (2002).
Ogonuki, N. et al. Early death of mice cloned from somatic cells. Nature Genet. 30, 253–254 (2002).
Tian, X. C., Xu, J. & Yang, X. Normal telomere lengths found in cloned cattle. Nature Genet. 26, 272–273 (2000).
Kang, Y. K. et al. Aberrant methylation of donor genome in cloned bovine embryos. Nature Genet. 28, 173–177 (2001).
Dean, W. et al. Conservation of methylation reprogramming in mammalian development: aberrant reprogramming in cloned embryos. Proc. Natl Acad. Sci. USA 98, 13734–13738 (2001).
Bourc'his, D. et al. Delayed and incomplete reprogramming of chromosome methylation patterns in bovine cloned embryos. Curr. Biol. 11, 1542–1546 (2001).
Kang, Y. K. et al. Limited demethylation leaves mosaic-type methylation states in cloned bovine pre-implantation embryos. EMBO J. 21, 1092–1100 (2002).
Boiani, M., Eckardt, S., Scholer, H. R. & McLaughlin, K. J. Oct4 distribution and level in mouse clones: consequences for pluripotency. Genes Dev. 16, 1209–1219 (2002).
Niwa, H., Miyazaki, J. & Smith, A. G. Quantitative expression of Oct-3/4 defines differentiation, dedifferentiation or self-renewal of ES cells. Nature Genet. 24, 372–376 (2000).
Inoue, K. et al. Faithful expression of imprinted genes in cloned mice. Science 295, 297 (2002).
Rideout, W. M. III, Eggan, K. & Jaenisch, R. Nuclear cloning and epigenetic reprogramming of the genome. Science 293, 1093–1098 (2001).
Hochedlinger, K. & Jaenisch, R. Monoclonal mice generated by nuclear transfer from mature B and T donor cells. Nature 415, 1035–1038 (2002).
Rideout, W. M. III, Hochedlinger, K., Kyba, M., Daley, G. Q. & Jaenisch, R. Correction of a genetic defect by nuclear transplantation and combined cell and gene therapy. Cell 109, 17–27 (2002).This work shows the feasibility of therapeutic cloning in a mouse model.
Tada, M., Takahama, Y., Abe, K., Nakatsuji, N. & Tada, T. Nuclear reprogramming of somatic cells by in vitro hybridization with ES cells. Curr. Biol. 11, 1553–1558 (2001).
Ying, Q. L., Nichols, J., Evans, E. P. & Smith, A. G. Changing potency by spontaneous fusion. Nature 416, 545–548 (2002).
Jiang, Y. et al. Pluripotency of mesenchymal stem cells derived from adult marrow. Nature 418, 41–49 (2002).
Jones, P. A. & Baylin, S. B. The fundamental role of epigenetic events in cancer. Nature Rev. Genet. 3, 415–428 (2002).
Okano, M., Xie, S. & Li, E. Dnmt2 is not required for de novo and maintenance methylation of viral DNA in embryonic stem cells. Nucleic Acids Res. 26, 2536–2540 (1998).
Hendrich, B., Hardeland, U., Ng, H. H., Jiricny, J. & Bird, A. The thymine glycosylase MBD4 can bind to the product of deamination at methylated CpG sites. Nature 401, 301–304 (1999).
Riccio, A. et al. The DNA repair gene MBD4 (MED1) is mutated in human carcinomas with microsatellite instability. Nature Genet. 23, 266–268 (1999).
Prokhortchouk, A. et al. The p120 catenin partner Kaiso is a DNA methylation-dependent transcriptional repressor. Genes Dev. 15, 1613–1618 (2001).
Wang, H. et al. Purification and functional characterization of a histone H3-lysine 4-specific methyltransferase. Mol. Cell 8, 1207–1217 (2001).
Nishioka, K. et al. Set9, a novel histone H3 methyltransferase that facilitates transcription by precluding histone tail modifications required for heterochromatin formation. Genes Dev. 16, 479–489 (2002).
Rea, S. et al. Regulation of chromatin structure by site-specific histone H3 methyltransferases. Nature 406, 593–599 (2000).
Yang, L. et al. Molecular cloning of ESET, a novel histone H3-specific methyltransferase that interacts with ERG transcription factor. Oncogene 21, 148–152 (2002).
Schultz, D. C., Ayyanathan, K., Negorev, D., Maul, G. G. & Rauscher, F. J. III . SETDB1: a novel KAP-1-associated histone H3, lysine 9-specific methyltransferase that contributes to HP1-mediated silencing of euchromatic genes by KRAB zinc-finger proteins. Genes Dev. 16, 919–932 (2002).
Ogawa, H., Ishiguro, K., Gaubatz, S., Livingston, D. M. & Nakatani, Y. A complex with chromatin modifiers that occupies E2F- and Myc-responsive genes in G0 cells. Science 296, 1132–1136 (2002).
Peterson, C. L. Chromatin remodeling: nucleosomes bulging at the seams. Curr. Biol. 12, R245–R247 (2002).
Reyes, J. C. et al. Altered control of cellular proliferation in the absence of mammalian brahma (SNF2α). EMBO J. 17, 6979–6991 (1998).
Roberts, C. W., Galusha, S. A., McMenamin, M. E., Fletcher, C. D. & Orkin, S. H. Haploinsufficiency of Snf5 (integrase interactor 1) predisposes to malignant rhabdoid tumors in mice. Proc. Natl Acad. Sci. USA 97, 13796–13800 (2000).
Kim, J. K. et al. Srg3, a mouse homolog of yeast SWI3, is essential for early embryogenesis and involved in brain development. Mol. Cell. Biol. 21, 7787–7795 (2001).
Acknowledgements
I am indebted to members of my lab for stimulating discussion and comments on the manuscript and to many colleagues in the field for discussion during the preparation of this review. Research in my laboratory is supported by the National Institutes of Health.
Author information
Authors and Affiliations
Related links
Related links
DATABASES
LocusLink
OMIM
The <i>Arabidopsis</i> Information Resource
Glossary
- HISTONES
-
Small, highly conserved basic proteins, found in the chromatin of all eukaryotic cells, which associate with DNA to form a nucleosome.
- CORE HISTONES
-
These are histones H2A, H2B, H3 and H4. A nucleosome contains two copies of each of the core histones wrapped by 146-bp DNA.
- EUCHROMATIN
-
The lightly staining regions of the nucleus that generally contain decondensed, transcriptionally active regions of the genome.
- HETEROCHROMATIN
-
A cytologically defined genomic component that contains repetitive DNA (highly repetitive satellite DNA, transposable elements and ribosomal DNA gene clusters) and some protein-coding genes.
- EPIGENETIC
-
Any heritable influence (in the progeny of cells or of individuals) on chromosome or gene function that is not accompanied by a change in DNA sequence. Examples of epigenetic events include mammalian X-chromosome inactivation, imprinting, centromere inactivation and position effect variegation.
- CHROMATIN MODIFICATION
-
Includes processes such as DNA methylation and histone modification (acetylation, phosphorylation, methylation and ubiquitylation).
- CHROMATIN REMODELLING
-
Transient changes in chromatin accessibility.
- NUCLEOSOME
-
The fundamental unit into which DNA and histones are packaged in eukaryotic cells. It is the basic structural subunit of chromatin and consists of 200 bp of DNA and an octamer of histone proteins, comprising two of each core histone.
- SET DOMAIN
-
(Suvar3–9, Enhancer-of-zeste, Trithorax) domain. An evolutionarily conserved sequence motif that was initially identified in the Drosophila position effect variegation suppressor Su(var)3–9, the Polycomb-group protein Enhancer-of-zeste, and the Trithorax-group protein Trithorax. It is present in many histone methyltransferases and is required for enzyme activity.
- HETEROCHROMATIN PROTEIN 1
-
(HP1). A protein that binds to highly repetitive, heterochromatic satellite DNA at centromeres and telomeres.
- CHROMODOMAIN
-
A highly conserved sequence motif that has been identified in various animal and plant species. Chromodomain proteins are often structural components of large macromolecular chromatin complexes or are involved in remodelling chromatin structure.
- GASTRULATION
-
A morphogenetic process that leads to the formation of the mesoderm layer between the endoderm and ectoderm layers and to the formation of embryonic body patterns.
- DIFFERENTIALLY METHYLATED REGION
-
(DMR). DNA segments in imprinted genes that show different methylation patterns between paternal and maternal alleles. Some DMRs acquire DNA methylation in the germ cells, whereas others acquire DNA methylation during embryogenesis.
- PRONUCLEUS
-
The sperm nucleus or the egg nucleus in a fertilized egg before a single nucleus forms.
- BLASTOCYST
-
A pre-implantation embryo that contains a fluid-filled cavity called a blastocoel.
- INNER CELL MASS
-
(ICM). A small clump of apparently undifferentiated cells in the blastocyst, which gives rise to the entire fetus and some of its extra-embryonic membranes.
- TROPHOBLAST
-
The post-implantation derivatives of the trophectoderm, which make up most of the fetal part of the placenta.
- PRIMITIVE ENDODERM
-
An early differentiated cell type that lines the inner surface of the blastocyst cavity. It gives rise to the endoderm component of the extra-embryonic membranes.
- YOLK SAC
-
An extra-embryonic membrane that consists of an outer endoderm layer and an inner mesoderm layer, which surrounds the developing embryo.
- TROPHECTODERM
-
The outer epithelial layer of the blastocyst.
- PLURIPOTENCY
-
The ability of a cell to contribute to several tissues in a developing organism. If a cell is able to contribute to all tissues, it is said to be totipotent.
Rights and permissions
About this article
Cite this article
Li, E. Chromatin modification and epigenetic reprogramming in mammalian development. Nat Rev Genet 3, 662–673 (2002). https://doi.org/10.1038/nrg887
Issue Date:
DOI: https://doi.org/10.1038/nrg887
This article is cited by
-
Obesity-associated epigenetic alterations and the obesity-breast cancer axis
Oncogene (2024)
-
IDH1 mutation impairs antiviral response and potentiates oncolytic virotherapy in glioma
Nature Communications (2023)
-
Effect of lipotoxicity on mitochondrial function and epigenetic programming during bovine in vitro embryo production
Scientific Reports (2023)
-
Plant deubiquitinases: from structure and activity to biological functions
Plant Cell Reports (2023)
-
Embryonic exposure to decitabine induces multiple neural tube defects in developing zebrafish
Fish Physiology and Biochemistry (2023)