Key Points
-
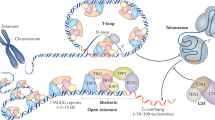
Telomeres protect chromosome ends from unwanted activation of DNA damage response and hazardous replication by a complex and specific network of interactions among DNA, RNA (telomere repeat-containing RNAs), enzymes (telomerase) and specialized proteins (the shelterin complex), as well as general chromatin and repair factors.
-
Specialized telomeric factors such as shelterin components and telomerase can be found outside telomeres, where they have different roles in transcriptional regulation.
-
In budding yeast, delocalization of the telomeric protein Rap1 from critically short telomeres contributes to the transcriptomic changes seen in senescent cells.
-
In mammals, the shelterin subunit RAP1 regulates the expression of gene networks that are related to metabolism and immunity. The shelterin subunit TRF2 regulates neuronal gene expression networks, and telomerase is involved in the transcription of a subset of Wnt/β-catenin and NF-κB target genes.
-
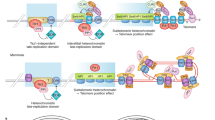
Three non-exclusive mechanisms can be envisaged to explain how telomeres signal their changes through the regulation of the transcriptional properties of their constituents: post-translational modifications, looping and subnuclear compartmentalization.
-
As each telomeric protein may control specific gene networks, the consequences of telomere dysfunction might be more tissue specific than previously expected. Blocking the extratelomeric functions of telomeric proteins should be considered a viable option for drugs against cancer and age-related pathologies without secondary effects due to telomere uncapping.
Abstract
Telomeres protect chromosome ends from degradation and inappropriate DNA damage response activation through their association with specific factors. Interestingly, these telomeric factors are able to localize outside telomeric regions, where they can regulate the transcription of genes involved in metabolism, immunity and differentiation. These findings delineate a signalling pathway by which telomeric changes control the ability of their associated factors to regulate transcription. This mechanism is expected to enable a greater diversity of cellular responses that are adapted to specific cell types and telomeric changes, and may therefore represent a pivotal aspect of development, ageing and telomere-mediated diseases.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Blackburn, E. H. Switching and signaling at the telomere. Cell 106, 661–673 (2001).
de Lange, T. Shelterin: the protein complex that shapes and safeguards human telomeres. Genes Dev. 19, 2100–2110 (2005).
Giraud-Panis, M. J. et al. One identity or more for telomeres? Front. Oncol. 3, 48 (2013).
Lustig, A. J., Kurtz, S. & Shore, D. Involvement of the silencer and UAS binding protein RAP1 in regulation of telomere length. Science 250, 549–553 (1990).
Conrad, M. N., Wright, J. H., Wolf, A. J. & Zakian, V. A. RAP1 protein interacts with yeast telomeres in vivo: overproduction alters telomere structure and decreases chromosome stability. Cell 63, 739–750 (1990).
Gilson, E., Roberge, M., Giraldo, R., Rhodes, D. & Gasser, S. M. Distortion of the DNA double helix by RAP1 at silencers and multiple telomeric binding sites. J. Mol. Biol. 231, 293–310 (1993).
Wotton, D. & Shore, D. A novel Rap1p-interacting factor, Rif2p, cooperates with Rif1p to regulate telomere length in S. cerevisiae. Genes Dev. 11, 748–760 (1997).
Kueng, S., Oppikofer, M. & Gasser, S. M. SIR proteins and the assembly of silent chromatin in budding yeast. Annu. Rev. Genet. 47, 275–306 (2013).
Azzalin, C. M., Reichenbach, P., Khoriauli, L., Giulotto, E. & Lingner, J. Telomeric repeat containing RNA and RNA surveillance factors at mammalian chromosome ends. Science 318, 798–801 (2007).
Schoeftner, S. & Blasco, M. A. Developmentally regulated transcription of mammalian telomeres by DNA-dependent RNA polymerase II. Nature Cell Biol. 10, 228–236 (2008).
Gottschling, D. E., Aparicio, O. M., Billington, B. L. & Zakian, V. A. Position effect at S. cerevisiae telomeres: reversible represssion of Pol II transcription. Cell 63, 751–762 (1990).
Gilson, E. & Geli, V. How telomeres are replicated. Nature Rev. Mol. Cell Biol. 8, 825–838 (2007).
Blackburn, E. H., Greider, C. W. & Szostak, J. W. Telomeres and telomerase: the path from maize, Tetrahymena and yeast to human cancer and aging. Nature Med. 12, 1133–1138 (2006).
Armanios, M. & Blackburn, E. H. The telomere syndromes. Nature Rev. Genet. 13, 693–704 (2012).
Hayashi, M. T., Cesare, A. J., Fitzpatrick, J. A., Lazzerini-Denchi, E. & Karlseder, J. A telomere-dependent DNA damage checkpoint induced by prolonged mitotic arrest. Nature Struct. Mol. Biol. 19, 387–394 (2012).
von Zglinicki, T., Saretzki, G., Docke, W. & Lotze, C. Mild hyperoxia shortens telomeres and inhibits proliferation of fibroblasts: a model for senescence? Exp. Cell Res. 220, 186–193 (1995).
Fumagalli, M. et al. Telomeric DNA damage is irreparable and causes persistent DNA-damage-response activation. Nature Cell Biol. 14, 355–365 (2012).
Romano, G. H. et al. Environmental stresses disrupt telomere length homeostasis. PLoS Genet. 9, e1003721 (2013).
Choi, J., Fauce, S. R. & Effros, R. B. Reduced telomerase activity in human T lymphocytes exposed to cortisol. Brain Behav. Immun. 22, 600–605 (2008).
Epel, E. S. et al. Accelerated telomere shortening in response to life stress. Proc. Natl Acad. Sci. USA 101, 17312–17315 (2004).
Heidinger, B. J. et al. Telomere length in early life predicts lifespan. Proc. Natl Acad. Sci. USA 109, 1743–1748 (2012).
d'Adda di Fagagna, F. et al. A DNA damage checkpoint response in telomere-initiated senescence. Nature 426, 194–198 (2003).
Takai, H., Smogorzewska, A. & de Lange, T. DNA damage foci at dysfunctional telomeres. Curr. Biol. 13, 1549–1556 (2003).
Nautiyal, S., DeRisi, J. L. & Blackburn, E. H. The genome-wide expression response to telomerase deletion in Saccharomyces cerevisiae. Proc. Natl Acad. Sci. USA 99, 9316–9321 (2002).
Lou, Z. et al. Telomere length regulates ISG15 expression in human cells. Aging (Albany NY) 1, 608–621 (2009).
Low, K. C. & Tergaonkar, V. Telomerase: central regulator of all of the hallmarks of cancer. Trends Biochem. Sci. 38, 426–434 (2013).
Hirashima, K. et al. Telomere length influences cancer cell differentiation in vivo. Mol. Cell. Biol. 33, 2988–2995 (2013).
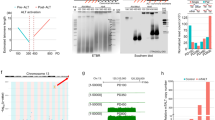
Biroccio, A. et al. TRF2 inhibits a cell-extrinsic pathway through which natural killer cells eliminate cancer cells. Nature Cell Biol. 15, 818–828 (2013). This paper shows that TRF2 binds to and regulates the transcription of HS3ST4 , which is involved in the elimination of cancer cells by natural killer cells. It also reveals an unexpected role of IL-6 in telomere protection.
Martinez, P. et al. RAP1 protects from obesity through its extratelomeric role regulating gene expression. Cell Rep. 3, 2059–2074 (2013). This study shows that mouse RAP1 binds to and regulates the expression of key genes that are involved in energy metabolism.
Yeung, F. et al. Nontelomeric role for Rap1 in regulating metabolism and protecting against obesity. Cell Rep. 3, 1847–1856 (2013). This paper reveals that Rap1 -null mice show important metabolic disorders in the absence of overt telomere damage.
Platt, J. M. et al. Rap1 relocalization contributes to the chromatin-mediated gene expression profile and pace of cell senescence. Genes Dev. 27, 1406–1420 (2013). This is the first study to show that telomere shortening triggers a redistribution of telomeric factors that contributes to the transcriptomic changes of senescent yeast cells.
Gasser, S. M., Hediger, F., Taddei, A., Neumann, F. R. & Gartenberg, M. R. The function of telomere clustering in yeast: the circe effect. Cold Spring Harb. Symp. Quant. Biol. 69, 327–337 (2004).
Maillet, L. et al. Evidence for silencing compartments within the yeast nucleus: a role for telomere proximity and Sir-protein concentration in silencer-mediated repression. Genes Dev. 10, 1796–1811 (1996).
Marcand, S., Buck, S. W., Moretti, P., Gilson, E. & Shore, D. Silencing of genes at nontelomeric sites in yeast is controlled by sequestration of silencing factors at telomeres by Rap1 protein. Genes Dev. 10, 1297–1309 (1996).
Gotta, M. et al. The clustering of telomeres and colocalization with Rap1, Sir3, and Sir4 proteins in wild type S. cerevisiae. J. Cell Biol. 134, 1349–1363 (1996).
Campisi, J. The biology of replicative senescence. Eur. J. Cancer 33, 703–709 (1997).
Martin, S. G., Laroche, T., Suka, N., Grunstein, M. & Gasser, S. M. Relocalization of telomeric Ku and SIR proteins in response to DNA strand breaks in yeast. Cell 97, 621–633 (1999).
Mills, K. D., Sinclair, D. A. & Guarente, L. MEC1-dependent redistribution of the Sir3 silencing protein from telomeres to DNA double-strand breaks. Cell 97, 609–620 (1999).
McAinsh, A. D., Scott-Drew, S., Murray, J. A. & Jackson, S. P. DNA damage triggers disruption of telomeric silencing and Mec1p-dependent relocation of Sir3p. Curr. Biol. 9, 963–966 (1999).
Straatman, K. R. & Louis, E. J. Localization of telomeres and telomere-associated proteins in telomerase-negative Saccharomyces cerevisiae. Chromosome Res. 15, 1033–1050 (2007).
Kennedy, B. K. et al. Redistribution of silencing protein from telomeres to the nucleolus is associated with extension in life span in S. cerevisiae. Cell 89, 381–391 (1997).
Taddei, A. et al. The functional importance of telomere clustering: global changes in gene expression result from SIR factor dispersion. Genome Res. 19, 611–625 (2009). This study shows that Sir proteins released from their nuclear membrane anchors redistribute throughout the genome and change the transcriptome.
Shi, T. et al. Rif1 and Rif2 shape telomere function and architecture through multivalent Rap1 interactions. Cell 153, 1340–1353 (2013).
O'Sullivan, R. J., Kubicek, S., Schreiber, S. L. & Karlseder, J. Reduced histone biosynthesis and chromatin changes arising from a damage signal at telomeres. Nature Struct. Mol. Biol. 17, 1218–1225 (2010).
Ray, A. et al. Sir3p phosphorylation by the Slt2p pathway effects redistribution of silencing function and shortened lifespan. Nature Genet. 33, 522–526 (2003).
Radman-Livaja, M. et al. Dynamics of Sir3 spreading in budding yeast: secondary recruitment sites and euchromatic localization. EMBO J. 30, 1012–1026 (2011).
Kueng, S. et al. Regulating repression: roles for the Sir4 N-terminus in linker DNA protection and stabilization of epigenetic states. PLoS Genet. 8, e1002727 (2012).
Mattarocci, S. et al. Rif1 controls DNA replication timing in yeast through the PP1 phosphatase Glc7. Cell Rep. 7, 62–69 (2014).
Teytelman, L., Thurtle, D. M., Rine, J. & van Oudenaarden, A. Highly expressed loci are vulnerable to misleading ChIP localization of multiple unrelated proteins. Proc. Natl Acad. Sci. USA 110, 18602–18607 (2013).
Bilaud, T. et al. Telomeric localization of TRF2, a novel human telobox protein. Nature Genet. 17, 236–239 (1997).
Broccoli, D., Smogorzewska, A., Chong, L. & de Lange, T. Human telomeres contain two distinct Myb-related proteins, TRF1 and TRF2. Nature Genet. 17, 231–235 (1997).
Li, B., Oestreich, S. & de Lange, T. Identification of human Rap1: implications for telomere evolution. Cell 101, 471–483 (2000).
Martinez, P. et al. Mammalian Rap1 controls telomere function and gene expression through binding to telomeric and extratelomeric sites. Nature Cell Biol. 12, 768–780 (2010). This is the first paper to show that a mammalian shelterin component can be located outside telomeres by acting as a transcription factor.
Yang, D. et al. Human telomeric proteins occupy selective interstitial sites. Cell Res. 21, 1013–1027 (2011). This study compares the genomic binding sites of RAP1 and TRF2 in human cancer cells.
Teo, H. et al. Telomere-independent Rap1 is an IKK adaptor and regulates NF-κB-dependent gene expression. Nature Cell Biol. 12, 758–767 (2010). This paper unveils a role of mammalian RAP1 in NF-κB signalling. It was the first study to show that RAP1 localizes outside telomere by working as an adaptor in the cytoplasm.
Takai, K. K., Hooper, S., Blackwood, S., Gandhi, R. & de Lange, T. In vivo stoichiometry of shelterin components. J. Biol. Chem. 285, 1457–1467 (2010).
Lian, S. et al. PRL-3 activates NF-κB signaling pathway by interacting with RAP1. Biochem. Biophys. Res. Commun. 430, 196–201 (2013).
Smogorzewska, A. et al. Control of human telomere length by TRF1 and TRF2. Mol. Cell. Biol. 20, 1659–1668 (2000).
Krutilina, R. I. et al. Protection of internal (TTAGGG)n repeats in Chinese hamster cells by telomeric protein TRF1. Oncogene 22, 6690–6698 (2003).
Ye, J. et al. TRF2 and apollo cooperate with topoisomerase 2α to protect human telomeres from replicative damage. Cell 142, 230–242 (2010).
Simonet, T. et al. The human TTAGGG repeat factors 1 and 2 bind to a subset of interstitial telomeric sequences and satellite repeats. Cell Res. 21, 1028–1038 (2011). This study shows that TRF1 and TRF2 can colocalize at ITSs and pericentromeric satellite sequences in human cancer cells.
Azzalin, C. M., Nergadze, S. G. & Giulotto, E. Human intrachromosomal telomeric-like repeats: sequence organization and mechanisms of origin. Chromosoma 110, 75–82 (2001).
Lieb, J. D., Liu, X., Botstein, D. & Brown, P. O. Promoter-specific binding of Rap1 revealed by genome-wide maps of protein–DNA association. Nature Genet. 28, 327–334 (2001).
Poulet, A. et al. TRF2 promotes, remodels and protects telomeric Holliday junctions. EMBO J. 28, 641–651 (2009).
Poulet, A. et al. The N-terminal domains of TRF1 and TRF2 regulate their ability to condense telomeric DNA. Nucleic Acids Res. 40, 2566–2576 (2011).
Arat, N. O. & Griffith, J. D. Human Rap1 interacts directly with telomeric DNA and regulates TRF2 localization at the telomere. J. Biol. Chem. 287, 41583–41594 (2012).
Diala, I. et al. Telomere protection and TRF2 expression are enhanced by the canonical Wnt signalling pathway. EMBO Rep. 14, 356–363 (2013).
Mattern, K. A. et al. Dynamics of protein binding to telomeres in living cells: implications for telomere structure and function. Mol. Cell. Biol. 24, 5587–5594 (2004).
Zhang, P. et al. Nontelomeric TRF2–REST interaction modulates neuronal gene silencing and fate of tumor and stem cells. Curr. Biol. 18, 1489–1494 (2008).
Xu, L. & Blackburn, E. H. Human Rif1 protein binds aberrant telomeres and aligns along anaphase midzone microtubules. J. Cell Biol. 167, 819–830 (2004).
Zimmermann, M., Lottersberger, F., Buonomo, S. B., Sfeir, A. & de Lange, T. 53BP1 regulates DSB repair using Rif1 to control 5′ end resection. Science 339, 700–704 (2013).
Cornacchia, D. et al. Mouse Rif1 is a key regulator of the replication-timing programme in mammalian cells. EMBO J. 31, 3678–3690 (2012).
Dan, J. et al. Rif1 maintains telomere length homeostasis of ESCs by mediating heterochromatin silencing. Dev. Cell 29, 7–19 (2014).
Benetti, R., Schoeftner, S., Munoz, P. & Blasco, M. A. Role of TRF2 in the assembly of telomeric chromatin. Cell Cycle 7, 3461–3468 (2008).
Galati, A. et al. TRF2 controls telomeric nucleosome organization in a cell cycle phase-dependent manner. PLoS ONE 7, e34386 (2012).
Kim, H. et al. TRF2 functions as a protein hub and regulates telomere maintenance by recognizing specific peptide motifs. Nature Struct. Mol. Biol. 16, 372–379 (2009).
Giannone, R. J. et al. The protein network surrounding the human telomere repeat binding factors TRF1, TRF2, and POT1. PLoS ONE 5, e12407 (2010).
Lee, O. H. et al. Genome-wide YFP fluorescence complementation screen identifies new regulators for telomere signaling in human cells. Mol. Cell Proteomics 10, M110.001628 (2011).
Deng, Z., Norseen, J., Wiedmer, A., Riethman, H. & Lieberman, P. M. TERRA RNA binding to TRF2 facilitates heterochromatin formation and ORC recruitment at telomeres. Mol. Cell 35, 403–413 (2009).
Buck, M. J. & Lieb, J. D. A chromatin-mediated mechanism for specification of conditional transcription factor targets. Nature Genet. 38, 1446–1451 (2006).
Spiegelman, B. M. in Mitochondrial Biology: New Perspectives, Number 287 60–69 (Wiley, 2007).
Lin, J. et al. Defects in adaptive energy metabolism with CNS-linked hyperactivity in PGC-1α null mice. Cell 119, 121–135 (2004).
Leone, T. C. et al. PGC-1α deficiency causes multi-system energy metabolic derangements: muscle dysfunction, abnormal weight control and hepatic steatosis. PLoS Biol. 3, e101 (2005).
Arkan, M. C. et al. IKK-β links inflammation to obesity-induced insulin resistance. Nature Med. 11, 191–198 (2005).
Salminen, A. et al. Activation of innate immunity system during aging: NF-κB signaling is the molecular culprit of inflamm-aging. Ageing Res. Rev. 7, 83–105 (2008).
Sfeir, A., Kabir, S., van Overbeek, M., Celli, G. B. & de Lange, T. Loss of Rap1 induces telomere recombination in the absence of NHEJ or a DNA damage signal. Science 327, 1657–1661 (2010).
Jung, Y. et al. TRF2 is in neuroglial cytoplasm and induces neurite-like processes. FEBS Lett. 557, 129–132 (2004).
Cheng, A. et al. Telomere protection mechanisms change during neurogenesis and neuronal maturation: newly generated neurons are hypersensitive to telomere and DNA damage. J. Neurosci. 27, 3722–3733 (2007).
Ovando-Roche, P., Yu, J. S., Testori, S., Ho, C. & Cui, W. TRF2-mediated stabilization of hREST4 is critical for the differentiation and maintenance of neural progenitors. Stem Cells http://dx.doi.org/10.1002/stem.1725 (2014). This paper shows that TRF2 is required for neurogenesis by stabilizing REST4.
Kishi, S. et al. The identification of zebrafish mutants showing alterations in senescence-associated biomarkers. PLoS Genet. 4, e1000152 (2008).
Zhang, P. et al. TRF2 dysfunction elicits DNA damage responses associated with senescence in proliferating neural cells and differentiation of neurons. J. Neurochem. 97, 567–581 (2006).
Zhang, P. et al. Nontelomeric splice variant of telomere repeat-binding factor 2 maintains neuronal traits by sequestering repressor element 1-silencing transcription factor. Proc. Natl Acad. Sci. USA 108, 16434–16439 (2011). This paper proposes a mechanism by which a cytoplasmic form of TRF2 prevents the import of REST and consequently the repression of genes that are involved in neural differentiation.
Renault, V. M. et al. FoxO3 regulates neural stem cell homeostasis. Cell Stem Cell 5, 527–539 (2009).
Webb, A. E. et al. FOXO3 shares common targets with ASCL1 genome-wide and inhibits ASCL1-dependent neurogenesis. Cell Rep. 4, 477–491 (2013).
Chu, C., Qu, K., Zhong, F. L., Artandi, S. E. & Chang, H. Y. Genomic maps of long noncoding RNA occupancy reveal principles of RNA–chromatin interactions. Mol. Cell 44, 667–678 (2011).
Choi, J. et al. TERT promotes epithelial proliferation through transcriptional control of a Myc- and Wnt-related developmental program. PLoS Genet. 4, e10 (2008).
Park, J. I. et al. Telomerase modulates Wnt signalling by association with target gene chromatin. Nature 460, 66–72 (2009).
Shkreli, M. et al. Reversible cell-cycle entry in adult kidney podocytes through regulated control of telomerase and Wnt signaling. Nature Med. 18, 111–119 (2012).
Okamoto, N. et al. Maintenance of tumor initiating cells of defined genetic composition by nucleostemin. Proc. Natl Acad. Sci. USA 108, 20388–20393 (2011).
Hrdlickova, R., Nehyba, J. & Bose, H. R. Jr. Alternatively spliced telomerase reverse transcriptase variants lacking telomerase activity stimulate cell proliferation. Mol. Cell. Biol. 32, 4283–4296 (2012).
Vidal-Cardenas, S. L. & Greider, C. W. Comparing effects of mTR and mTERT deletion on gene expression and DNA damage response: a critical examination of telomere length maintenance-independent roles of telomerase. Nucleic Acids Res. 38, 60–71 (2010).
Strong, M. A. et al. Phenotypes in mTERT+/− and mTERT−/− mice are due to short telomeres, not telomere-independent functions of telomerase reverse transcriptase. Mol. Cell. Biol. 31, 2369–2379 (2011).
Gardano, L., Pucci, F., Christian, L., Le Bihan, T. & Harrington, L. Telomeres, a busy platform for cell signaling. Front. Oncol. 3, 146 (2013).
Listerman, I., Gazzaniga, F. S. & Blackburn, E. H. An investigation of the effects of the telomerase core protein TERT on Wnt signaling in human breast cancer cells. Mol. Cell. Biol. 34, 280–289 (2013).
Ghosh, A. et al. Telomerase directly regulates NF-κB-dependent transcription. Nature Cell Biol. 14, 1270–1281 (2012).
Hoffmeyer, K. et al. Wnt/β-catenin signaling regulates telomerase in stem cells and cancer cells. Science 336, 1549–1554 (2012).
Yin, L., Hubbard, A. K. & Giardina, C. NF-κB regulates transcription of the mouse telomerase catalytic subunit. J. Biol. Chem. 275, 36671–36675 (2000).
Ding, D., Xi, P., Zhou, J., Wang, M. & Cong, Y. S. Human telomerase reverse transcriptase regulates MMP expression independently of telomerase activity via NF-κB-dependent transcription. FASEB J. 27, 4375–4383 (2013).
Walker, J. R. & Zhu, X. D. Post-translational modifications of TRF1 and TRF2 and their roles in telomere maintenance. Mech. Ageing Dev. 133, 421–434 (2012).
Smith, S., Giriat, I., Schmitt, A. & de Lange, T. Tankyrase, a poly(ADP-ribose) polymerase at human telomeres. Science 282, 1484–1487 (1998).
Seimiya, H., Muramatsu, Y., Smith, S. & Tsuruo, T. Functional subdomain in the ankyrin domain of tankyrase 1 required for poly(ADP-ribosyl)ation of TRF1 and telomere elongation. Mol. Cell. Biol. 24, 1944–1955 (2004).
Chang, W., Dynek, J. N. & Smith, S. TRF1 is degraded by ubiquitin-mediated proteolysis after release from telomeres. Genes Dev. 17, 1328–1333 (2003).
Dantzer, F. et al. Functional interaction between poly(ADP-ribose) polymerase 2 (PARP-2) and TRF2: PARP activity negatively regulates TRF2. Mol. Cell. Biol. 24, 1595–1607 (2004).
Gomez, M. et al. PARP1 is a TRF2-associated poly(ADP-ribose) polymerase and protects eroded telomeres. Mol. Biol. Cell 17, 1686–1696 (2006).
Fujita, K. et al. Positive feedback between p53 and TRF2 during telomere-damage signalling and cellular senescence. Nature Cell Biol. 12, 1205–1212 (2010).
Her, Y. R. & Chung, I. K. p300-mediated acetylation of TRF2 is required for maintaining functional telomeres. Nucleic Acids Res. 41, 2267–2283 (2013).
Griffith, J. D. et al. Mammalian telomeres end in a large duplex loop. Cell 97, 503–514 (1999).
Strahl-Bolsinger, S., Hecht, A., Luo, K. & Grunstein, M. SIR2 and SIR4 interactions differ in core and extended telomeric heterochromatin in yeast. Genes Dev. 11, 83–93 (1997).
de Bruin, D., Zaman, Z., Liberatore, R. A. & Ptashne, M. Telomere looping permits gene activation by a downstream UAS in yeast. Nature 409, 109–113 (2001).
Lebrun, E., Fourel, G., Defossez, P. A. & Gilson, E. A methyltransferase targeting assay reveals silencer–telomere interactions in budding yeast. Mol. Cell. Biol. 23, 1498–1508 (2003).
Bystricky, K., Laroche, T., van Houwe, G., Blaszczyk, M. & Gasser, S. M. Chromosome looping in yeast: telomere pairing and coordinated movement reflect anchoring efficiency and territorial organization. J. Cell Biol. 168, 375–387 (2005).
Wang, R. C., Smogorzewska, A. & de Lange, T. Homologous recombination generates T-loop-sized deletions at human telomeres. Cell 119, 355–368 (2004).
Vannier, J. B., Depeiges, A., White, C. & Gallego, M. E. ERCC1/XPF protects short telomeres from homologous recombination in Arabidopsis thaliana. PLoS Genet. 5, e1000380 (2009).
Fourel, G., Revardel, E., Koering, C. E. & Gilson, E. Cohabitation of insulators and silencing elements in yeast subtelomeric regions. EMBO J. 18, 2522–2537 (1999).
de Lange, T. Human telomeres are attached to the nuclear matrix. EMBO J. 11, 717–724 (1992).
Voronin, A. P., Lobov, I. B., Gilson, E. & Podgornaya, O. I. A telomere-binding protein (TRF2/MTBP) from mouse nuclear matrix with motives of an intermediate filament-type rod domain. J. Anti Aging Med. 6, 205–218 (2003).
Ottaviani, A. et al. Identification of a perinuclear positioning element in human subtelomeres that requires A-type lamins and CTCF. EMBO J. 28, 2428–2436 (2009).
Crabbe, L., Cesare, A. J., Kasuboski, J. M., Fitzpatrick, J. A. & Karlseder, J. Human telomeres are tethered to the nuclear envelope during postmitotic nuclear assembly. Cell Rep. 2, 1521–1529 (2012).
Kyrion, G., Liu, K. & Lustig, A. J. RAP1 and telomere structure regulate telomere position effects in Saccharomyces cerevisiae. Genes Dev. 7, 1146–1159 (1993).
de Bruin, D., Kantrow, S. M., Liberatore, R. A. & Zakian, V. A. Telomere folding is required for the stable maintenance of telomere position effects in yeast. Mol. Cell. Biol. 20, 7991–8000 (2000).
Mercier, G. et al. A haploid-specific transcriptional response to irradiation in Saccharomyces cerevisiae. Nucleic Acids Res. 33, 6635–6643 (2005).
Ruault, M., de Meyer, A., Loiodice, I. & Taddei, A. Clustering heterochromatin: Sir3 promotes telomere clustering independently of silencing in yeast. J. Cell Biol. 192, 417–431 (2011).
Laroche, T. et al. Mutation of yeast Ku genes disrupts the subnuclear organization of telomeres. Curr. Biol. 8, 653–656 (1998).
Maillet, L. et al. Ku-deficient yeast strains exhibit alternative states of silencing competence. EMBO Rep. 2, 203–210 (2001).
Baur, J. A., Zou, Y., Shay, J. W. & Wright, W. E. Telomere position effect in human cells. Science 292, 2075–2077 (2001).
Benetti, R., Garcia-Cao, M. & Blasco, M. A. Telomere length regulates the epigenetic status of mammalian telomeres and subtelomeres. Nature Genet. 39, 243–250 (2007).
Arnoult, N., Van Beneden, A. & Decottignies, A. Telomere length regulates TERRA levels through increased trimethylation of telomeric H3K9 and HP1α. Nature Struct. Mol. Biol. 19, 948–956 (2012). This study shows that the expression of non-coding TERRA is regulated by TPE in human cells.
Stadler, G. et al. Telomere position effect regulates DUX4 in human facioscapulohumeral muscular dystrophy. Nature Struct. Mol. Biol. 20, 671–678 (2013).
Schoeftner, S. et al. Telomere shortening relaxes X chromosome inactivation and forces global transcriptome alterations. Proc. Natl Acad. Sci. USA 106, 19393–19398 (2009).
Tennen, R. I., Bua, D. J., Wright, W. E. & Chua, K. F. SIRT6 is required for maintenance of telomere position effect in human cells. Nature Commun. 2, 433 (2011).
Koering, C. E. et al. Human telomeric position effect is determined by chromosomal context and telomeric chromatin integrity. EMBO Rep. 3, 1055–1061 (2002).
Fanti, L., Giovinazzo, G., Berloco, M. & Pimpinelli, S. The heterochromatin protein 1 prevents telomere fusions in Drosophila. Mol. Cell 2, 527–538 (1998).
Mitchell, T. R. & Zhu, X. D. Methylated TRF2 associates with the nuclear matrix and serves as a potential biomarker for cellular senescence. Aging (Albany NY) 6, 248–263 (2014).
Bradshaw, P. S., Stavropoulos, D. J. & Meyn, M. S. Human telomeric protein TRF2 associates with genomic double-strand breaks as an early response to DNA damage. Nature Genet. 37, 193–197 (2005).
Guo, N. et al. Short telomeres compromise β-cell signaling and survival. PLoS ONE 6, e17858 (2011).
Celli, G. B. & de Lange, T. DNA processing is not required for ATM-mediated telomere damage response after TRF2 deletion. Nature Cell Biol. 7, 712–718 (2005).
Sahin, E. & Depinho, R. A. Linking functional decline of telomeres, mitochondria and stem cells during ageing. Nature 464, 520–528 (2010).
Murillo-Ortiz, B. et al. Telomere length and type 2 diabetes in males, a premature aging syndrome. Aging Male 15, 54–58 (2012).
Martinez, P. & Blasco, M. A. Telomeric and extra-telomeric roles for telomerase and the telomere-binding proteins. Nature Rev. Cancer 11, 161–176 (2011).
Tazumi, A. et al. Telomere-binding protein Taz1 controls global replication timing through its localization near late replication origins in fission yeast. Genes Dev. 26, 2050–2062 (2012).
Dave, A., Cooley, C., Garg, M. & Bianchi, A. Protein phosphatase 1 recruitment by Rif1 regulates DNA replication origin firing by counteracting DDK activity. Cell Rep. 7, 53–61 (2014).
Hiraga, S. et al. Rif1 controls DNA replication by directing protein phosphatase 1 to reverse Cdc7-mediated phosphorylation of the MCM complex. Genes Dev. 28, 372–383 (2014).
Chen, L. Y. et al. Mitochondrial localization of telomeric protein TIN2 links telomere regulation to metabolic control. Mol. Cell 47, 839–850 (2012).
Listerman, I., Sun, J., Gazzaniga, F. S., Lukas, J. L. & Blackburn, E. H. The major reverse transcriptase-incompetent splice variant of the human telomerase protein inhibits telomerase activity but protects from apoptosis. Cancer Res. 73, 2817–2828 (2013).
Zhang, L. F. et al. Telomeric RNAs mark sex chromosomes in stem cells. Genetics 182, 685–698 (2009).
Acknowledgements
The authors thank the discussions and suggestions of all the members of the E.G. team, as well as M. Shkreli, V. Géli, P. Lopez and E. van Obberghen. They are also grateful to the anonymous reviewers for helpful suggestions. The E.G. team is supported by La Ligue Nationale contre le Cancer ('Equipe labellisée'), the French Institut National du Cancer (programmes TELOFUN and TELOCHROM), the French National Research Agency (ANR) (programmes TELOREP and INNATELO) and the Investments for the Future LABEX SIGNALIFE (programme reference # ANR-11-LABX-0028-01). They apologize for all the important works that have not been cited owing to space limitation.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Glossary
- DNA damage response
-
(DDR). A pathway triggered by DNA lesions, in which phosphatidylinositol-3 kinases (ATM, ATR or DNA-PK) activate appropriate repair pathways and/or cell cycle arrest.
- Senescence
-
A permanent arrest in the cell cycle.
- Apoptosis
-
A form of programmed cell death.
- Extracellular stress response pathways
-
Pathways that are triggered by extracellular stimuli and that involve mitogen-activated protein kinases such as JNK and p38 to activate an adaptive response programme, which maximizes cell cycle progression and survival.
- NF-κB signalling
-
A pathway mediated by nuclear factor-κB (NF-κB) that regulates many physiological processes, including innate and adaptive immune responses, cell death and inflammation.
- Mass action law
-
A mathematical model based on the principle that the rate of a chemical reaction is proportional to the molecular concentrations of the reacting substances.
- Wnt/β-catenin signalling
-
A fundamental and complex pathway that is required for metazoan development and tissue homeostasis; it is often dysregulated in cancer.
- Luciferase reporter assays
-
Assays commonly used to assess the transcriptional activity in cells that are transfected with a genetic construct containing the luciferase gene under the control of a promoter of interest.
- Glucose intolerance
-
A metabolic disorder associated with insulin resistance, which is considered to be a pre-diabetic state.
- Hepatic steatosis
-
The accumulation of stored lipids — most notably, triglycerides — to abnormally high levels in the liver.
- Oxidative phosphorylation
-
A metabolic pathway in which the mitochondria use energy released by the oxidation of nutrients to generate ATP.
- White fat
-
Adipose tissue used as energy storage.
- Poly(ADP-ribose) polymerases
-
A family of proteins involved in various cellular processes, including DNA repair, regulation of chromatin structure and regulation of cell cycle checkpoints.
- E3 ubiquitin ligase
-
An enzyme that couples the small (7 kDa) protein ubiquitin to lysine residues on proteins targeted for degradation by the proteasome.
- Natural killer cells
-
Cytotoxic cells that are crucial for eliminating infected, damaged or cancerous cells.
Rights and permissions
About this article
Cite this article
Ye, J., Renault, V., Jamet, K. et al. Transcriptional outcome of telomere signalling. Nat Rev Genet 15, 491–503 (2014). https://doi.org/10.1038/nrg3743
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrg3743
This article is cited by
-
Telomeres and telomerase: active but complex players in life-history decisions
Biogerontology (2024)
-
Non-canonical telomere protection role of FOXO3a of human skeletal muscle cells regulated by the TRF2-redox axis
Communications Biology (2023)
-
Sexual Dimorphism in Telomere Length in Childhood Autism
Journal of Autism and Developmental Disorders (2023)
-
Heterochromatin replication goes hand in hand with telomere protection
Nature Structural & Molecular Biology (2020)
-
Key role of piRNAs in telomeric chromatin maintenance and telomere nuclear positioning in Drosophila germline
Epigenetics & Chromatin (2018)