Key Points
-
The availability of genomic data has spurred the development of methods for genome-wide neutrality tests.
-
Genome-wide scans for selection have been applied to a variety of data sets that were collected from global population samples.
-
Both integrative studies that incorporated phenotypic data and regulatory variation, and functional follow-up studies of candidate adaptive variants have increased our understanding of how putatively adaptive genetic variation affects the phenotypic variation that may have a role in reproductive fitness.
-
There are several limitations to these approaches, such as false-positive signatures of adaptation.
-
Other issues include the limited models of adaptation (primarily the classic selective sweep) that have been incorporated into genome-wide scans for selection.
-
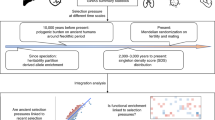
The most comprehensive examples of human adaptation studies take advantage of genome-wide genetic variation with phenotypic data and functional follow-up data to elucidate the role of genetic variants in reproductive fitness in recent human history.
Abstract
The recent availability of genomic data has spurred many genome-wide studies of human adaptation in different populations worldwide. Such studies have provided insights into novel candidate genes and pathways that are putatively involved in adaptation to different environments, diets and disease prevalence. However, much work is needed to translate these results into candidate adaptive variants that are biologically interpretable. In this Review, we discuss methods that may help to identify true biological signals of selection and studies that incorporate complementary phenotypic and functional data. We conclude with recommendations for future studies that focus on opportunities to use integrative genomics methodologies in human adaptation studies.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Scheinfeldt, L. B., Soi, S. & Tishkoff, S. A. Colloquium paper: working toward a synthesis of archaeological, linguistic, and genetic data for inferring African population history. Proc. Natl Acad. Sci. USA 107 (Suppl. 2), 8931–8938 (2010).
Henn, B. M., Cavalli-Sforza, L. L. & Feldman, M. W. The great human expansion. Proc. Natl Acad. Sci. USA 109, 17758–17764 (2012).
Jobling, M. A., Hurles, M. & Tyler-Smith, C. Human Evolutionary Genetics: Origins, Peoples and Disease (Garland Publishing, 2004).
Fraser, H. B. Gene expression drives local adaptation in humans. Genome Res. 23, 1089–1096 (2013). This study systematically evaluates the relative abundances of regulatory variation and coding variation in candidate adaptive regions and includes a novel method for identifying polygenic adaptation.
Sabeti, P. C. et al. Detecting recent positive selection in the human genome from haplotype structure. Nature 419, 832–837 (2002).
Scheinfeldt, L. B. et al. Population genomic analysis of ALMS1 in humans reveals a surprisingly complex evolutionary history. Mol. Biol. Evol. 26, 1357–1367 (2009).
Kleinjan, D. A. & van Heyningen, V. Long-range control of gene expression: emerging mechanisms and disruption in disease. Am. J. Hum. Genet. 76, 8–32 (2005).
Thorisson, G. A. & Stein, L. D. The SNP Consortium website: past, present and future. Nucleic Acids Res. 31, 124–127 (2003).
International HapMap Consortium. A haplotype map of the human genome. Nature 437, 1299–1320 (2005).
Hinds, D. A. et al. Whole-genome patterns of common DNA variation in three human populations. Science 307, 1072–1079 (2005).
Li, J. Z. et al. Worldwide human relationships inferred from genome-wide patterns of variation. Science 319, 1100–1104 (2008).
Jakobsson, M. et al. Genotype, haplotype and copy-number variation in worldwide human populations. Nature 451, 998–1003 (2008).
Sabeti, P. C. et al. Genome-wide detection and characterization of positive selection in human populations. Nature 449, 913–918 (2007).
Voight, B. F., Kudaravalli, S., Wen, X. & Pritchard, J. K. A map of recent positive selection in the human genome. PLoS Biol. 4, e72 (2006).
Wang, E. T., Kodama, G., Baldi, P. & Moyzis, R. K. Global landscape of recent inferred Darwinian selection for Homo sapiens. Proc. Natl Acad. Sci. USA 103, 135–140 (2006).
Williamson, S. H. et al. Localizing recent adaptive evolution in the human genome. PLoS Genet. 3, e90 (2007).
Zhang, C. et al. A whole genome long-range haplotype (WGLRH) test for detecting imprints of positive selection in human populations. Bioinformatics 22, 2122–2128 (2006).
Kelley, J. L., Madeoy, J., Calhoun, J. C., Swanson, W. & Akey, J. M. Genomic signatures of positive selection in humans and the limits of outlier approaches. Genome Res. 16, 980–989 (2006).
Kimura, R., Fujimoto, A., Tokunaga, K. & Ohashi, J. A practical genome scan for population-specific strong selective sweeps that have reached fixation. PLoS ONE 2, e286 (2007).
Pickrell, J. K. et al. Signals of recent positive selection in a worldwide sample of human populations. Genome Res. 19, 826–837 (2009).
Fumagalli, M. et al. Signatures of environmental genetic adaptation pinpoint pathogens as the main selective pressure through human evolution. PLoS Genet. 7, e1002355 (2011).
Brodwin, P. “Bioethics in action” and human population genetics research. Cult. Med. Psychiatry 29, 145–178 (2005).
Clark, A. G., Hubisz, M. J., Bustamante, C. D., Williamson, S. H. & Nielsen, R. Ascertainment bias in studies of human genome-wide polymorphism. Genome Res. 15, 1496–1502 (2005).
Biswas, S., Scheinfeldt, L. B. & Akey, J. M. Genome-wide insights into the patterns and determinants of fine-scale population structure in humans. Am. J. Hum. Genet. 84, 641–650 (2009).
Patterson, N. et al. Ancient admixture in human history. Genetics 192, 1065–1093 (2012).
Pickrell, J. K. et al. The genetic prehistory of southern Africa. Nature Commun. 3, 1143 (2012).
Teshima, K. M., Coop, G. & Przeworski, M. How reliable are empirical genomic scans for selective sweeps? Genome Res. 16, 702–712 (2006).
Tajima, F. Statistical method for testing the neutral mutation hypothesis by DNA polymorphism. Genetics 123, 585–595 (1989).
Fay, J. C. & Wu, C. I. Hitchhiking under positive Darwinian selection. Genetics 155, 1405–1413 (2000).
Fu, Y. X. & Li, W. H. Statistical tests of neutrality of mutations. Genetics 133, 693–709 (1993).
Holsinger, K. E. & Weir, B. S. Genetics in geographically structured populations: defining, estimating and interpreting FST . Nature Rev. Genet. 10, 639–650 (2009).
Shriver, M. D. et al. The genomic distribution of population substructure in four populations using 8,525 autosomal SNPs. Hum. Genom. 1, 274–286 (2004).
Yi, X. et al. Sequencing of 50 human exomes reveals adaptation to high altitude. Science 329, 75–78 (2010).
Chen, H., Patterson, N. & Reich, D. Population differentiation as a test for selective sweeps. Genome Res. 20, 393–402 (2010).
Grossman, S. R. et al. A composite of multiple signals distinguishes causal variants in regions of positive selection. Science 327, 883–886 (2010).
Liu, X. et al. Detecting and characterizing genomic signatures of positive selection in global populations. Am. J. Hum. Genet. 92, 866–881 (2013).
Sabeti, P. C. et al. Positive natural selection in the human lineage. Science 312, 1614–1620 (2006).
Oleksyk, T. K., Smith, M. W. & O'Brien, S. J. Genome-wide scans for footprints of natural selection. Phil. Trans. R. Soc. B 365, 185–205 (2010).
Lohmueller, K. E. et al. Natural selection affects multiple aspects of genetic variation at putatively neutral sites across the human genome. PLoS Genet. 7, e1002326 (2011). This paper demonstrates that deleterious mutations and hitch-hiking of neighbouring genetic variants are affecting patterns of nucleotide diversity across the human genome.
Price, A. L. et al. Long-range LD can confound genome scans in admixed populations. Am. J. Hum. Genet. 83, 132–135 (2008).
Tang, H. et al. Response to Price et al. Am. J. Hum. Genet. 83, 135–139 (2008).
Przeworski, M., Coop, G. & Wall, J. D. The signature of positive selection on standing genetic variation. Evolution 59, 2312–2323 (2005).
Innan, H. & Kim, Y. Detecting local adaptation using the joint sampling of polymorphism data in the parental and derived populations. Genetics 179, 1713–1720 (2008).
Pritchard, J. K., Pickrell, J. K. & Coop, G. The genetics of human adaptation: hard sweeps, soft sweeps, and polygenic adaptation. Curr. Biol. 20, R208–R215 (2010).
Cutter, A. D. & Payseur, B. A. Genomic signatures of selection at linked sites: unifying the disparity among species. Nature Rev. Genet. 14, 262–274 (2013).
Coop, G. et al. The role of geography in human adaptation. PLoS Genet. 5, e1000500 (2009).
Daub, J. T. et al. Evidence for polygenic adaptation to pathogens in the human genome. Mol. Biol. Evol. 30, 1544–1558 (2013).
Hernandez, R. D. et al. Classic selective sweeps were rare in recent human evolution. Science 331, 920–924 (2011). This study demonstrates that amino acid substitutions and putative regulatory sites are not significantly enriched in alleles that are highly differentiated between populations, suggesting that classic sweeps were not a dominant mode of human adaptation over the past ~250 kya.
Simonson, T. S. et al. Genetic evidence for high-altitude adaptation in Tibet. Science 329, 72–75 (2010).
Hancock, A. M. et al. Adaptations to climate-mediated selective pressures in humans. PLoS Genet. 7, e1001375 (2010). This study looks at correlations between SNP allele frequencies and climate-related variables across global populations (a method that may be useful for detecting polygenic adaptation), demonstrating an enrichment for gene sets that are related to ultraviolet radiation, cancer, infection and immunity.
Moore, L. G. et al. Maternal adaptation to high-altitude pregnancy: an experiment of nature — a review. Placenta 25 (Suppl. A), S60–S71 (2004).
Buzbas, E. O., Joyce, P. & Rosenberg, N. A. Inference on the strength of balancing selection for epistatically interacting loci. Theor. Popul. Biol. 79, 102–113 (2011).
Scheinfeldt, L. B., Biswas, S., Madeoy, J., Connelly, C. F. & Akey, J. M. Clusters of adaptive evolution in the human genome. Frontiers Genet. 2, 50 (2011).
Akey, J. M. Constructing genomic maps of positive selection in humans: where do we go from here? Genome Res. 19, 711–722 (2009). This paper evaluates results across several genome-wide scans for selection and highlights the degree to which false positives are common in these studies.
Li, J. et al. Joint analysis of demography and selection in population genetics: where do we stand and where could we go? Mol. Ecol. 21, 28–44 (2012).
Granka, J. M. et al. Limited evidence for classic selective sweeps in African populations. Genetics 192, 1049–1064 (2012).
Quach, H. et al. Signatures of purifying and local positive selection in human miRNAs. Am. J. Hum. Genet. 84, 316–327 (2009).
Andres, A. M. et al. Targets of balancing selection in the human genome. Mol. Biol. Evol. 26, 2755–2764 (2009).
Nielsen, R. et al. Darwinian and demographic forces affecting human protein coding genes. Genome Res. 19, 838–849 (2009).
Bazin, E., Dawson, K. J. & Beaumont, M. A. Likelihood-free inference of population structure and local adaptation in a Bayesian hierarchical model. Genetics 185, 587–602 (2010).
Sousa, V. M., Carneiro, M., Ferrand, N. & Hey, J. Identifying loci under selection against gene flow in isolation with migration models. Genetics 194, 211–233 (2013).
Bersaglieri, T. et al. Genetic signatures of strong recent positive selection at the lactase gene. Am. J. Hum. Genet. 74, 1111–1120 (2004).
Tishkoff, S. A. et al. Convergent adaptation of human lactase persistence in Africa and Europe. Nature Genet. 39, 31–40 (2007).
Lamason, R. L. et al. SLC24A5, a putative cation exchanger, affects pigmentation in zebrafish and humans. Science 310, 1782–1786 (2005).
Jablonski, N. G. & Chaplin, G. The evolution of human skin coloration. J. Hum. Evol. 39, 57–106 (2000).
Beleza, S. et al. Genetic architecture of skin and eye color in an African-European admixed population. PLoS Genet. 9, e1003372 (2013).
Lango Allen, H. et al. Hundreds of variants clustered in genomic loci and biological pathways affect human height. Nature 467, 832–838 (2010).
Kang, S. J. et al. Genome-wide association of anthropometric traits in African- and African-derived populations. Hum. Mol. Genet. 19, 2725–2738 (2010).
Jarvis, J. P. et al. Patterns of ancestry, signatures of natural selection, and genetic association with stature in Western African Pygmies. PLoS Genet. 8, e1002641 (2012). This study integrates signatures of natural selection together with GWASs to identify candidate loci that have a role in the short-stature trait in African pygmies.
Migliano, A. B., Vinicius, L. & Lahr, M. M. Life history trade-offs explain the evolution of human pygmies. Proc. Natl Acad. Sci. USA 104, 20216–20219 (2007).
Perry, G. H. & Dominy, N. J. Evolution of the human Pygmy phenotype. Trends Ecol. Evol. 24, 218–225 (2009).
Mendizabal, I., Marigorta, U. M., Lao, O. & Comas, D. Adaptive evolution of loci covarying with the human African Pygmy phenotype. Hum. Genet. 131, 1305–1317 (2012).
Turchin, M. C. et al. Evidence of widespread selection on standing variation in Europe at height-associated SNPs. Nature Genet. 44, 1015–1019 (2012). This paper demonstrates evidence for polygenic selection that acts on genetic variants influencing height in Northern Europeans compared to Southern Europeans.
Bigham, A. W. et al. Andean and Tibetan patterns of adaptation to high altitude. Am. J. Hum. Biol. 25, 190–197 (2013).
Bigham, A. W. et al. Identifying positive selection candidate loci for high-altitude adaptation in Andean populations. Hum. Genom. 4, 79–90 (2009).
Peng, Y. et al. Genetic variations in Tibetan populations and high-altitude adaptation at the Himalayas. Mol. Biol. Evol. 28, 1075–1081 (2011).
Beall, C. M. et al. Natural selection on EPAS1 (HIF2α) associated with low hemoglobin concentration in Tibetan highlanders. Proc. Natl Acad. Sci. USA 107, 11459–11464 (2010).
Scheinfeldt, L. B. et al. Genetic adaptation to high altitude in the Ethiopian highlands. Genome Biol. 13, R1 (2012).
Alkorta-Aranburu, G. et al. The genetic architecture of adaptations to high altitude in Ethiopia. PLoS Genet. 8, e1003110 (2012).
Huerta-Sanchez, E. et al. Genetic signatures reveal high-altitude adaptation in a set of Ethiopian populations. Mol. Biol. Evol. 30, 1877–1888 (2013).
Scheinfeldt, L. B. & Tishkoff, S. A. Living the high life: high-altitude adaptation. Genome Biol. 11, 133 (2010).
Enattah, N. S. et al. Independent introduction of two lactase-persistence alleles into human populations reflects different history of adaptation to milk culture. Am. J. Hum. Genet. 82, 57–72 (2008).
Imtiaz, F. et al. The T/G 13915 variant upstream of the lactase gene (LCT) is the founder allele of lactase persistence in an urban Saudi population. J. Med. Genet. 44, e89 (2007).
Ingram, C. J. et al. A novel polymorphism associated with lactose tolerance in Africa: multiple causes for lactase persistence? Hum. Genet. 120, 779–788 (2007).
Ingram, C. J., Mulcare, C. A., Itan, Y., Thomas, M. G. & Swallow, D. M. Lactose digestion and the evolutionary genetics of lactase persistence. Hum. Genet. 124, 579–591 (2009).
Ingram, C. J. et al. Multiple rare variants as a cause of a common phenotype: several different lactase persistence associated alleles in a single ethnic group. J. Mol. Evol. 69, 579–588 (2009).
Norton, H. L. et al. Genetic evidence for the convergent evolution of light skin in Europeans and East Asians. Mol. Biol. Evol. 24, 710–722 (2007).
Henn, B. M. et al. Hunter-gatherer genomic diversity suggests a southern African origin for modern humans. Proc. Natl Acad. Sci. USA 108, 5154–5162 (2011).
Kudaravalli, S., Veyrieras, J. B., Stranger, B. E., Dermitzakis, E. T. & Pritchard, J. K. Gene expression levels are a target of recent natural selection in the human genome. Mol. Biol. Evol. 26, 649–658 (2009). One of the first studies to integrate eQTLs with candidate adaptive loci that have been identified using genome-wide SNP data, to explore the degree to which regulatory variation has been involved in human adaptation.
Grossman, S. R. et al. Identifying recent adaptations in large-scale genomic data. Cell 152, 703–713 (2013). This paper integrates candidate adaptive loci identified in WGS data with several different types of regulatory variation (eQTLs, lincRNAs, enhancers and promoters).
Lambert, C. A. & Tishkoff, S. A. Genetic structure in African populations: implications for human demographic history. Cold Spring Harb. Symp. Quant. Biol. 74, 395–402 (2009).
Luzzatto, L. Sickle cell anaemia and malaria. Mediterr. J. Hematol. Infect. Dis 4, e2012065 (2012).
Vernot, B. et al. Personal and population genomics of human regulatory variation. Genome Res. 22, 1689–1697 (2012). This study integrates candidate adaptive loci with DNase I-hypersensitive sites that have been identified in the ENCODE project.
Olds, L. C. & Sibley, E. Lactase persistence DNA variant enhances lactase promoter activity in vitro: functional role as a cis regulatory element. Hum. Mol. Genet. 12, 2333–2340 (2003).
Hughes, D. A. et al. Parallel selection on TRPV6 in human populations. PLoS ONE 3, e1686 (2008).
Akey, J. M., Swanson, W. J., Madeoy, J., Eberle, M. & Shriver, M. D. TRPV6 exhibits unusual patterns of polymorphism and divergence in worldwide populations. Hum. Mol. Genet. 15, 2106–2113 (2006).
Kamberov, Y. G. et al. Modeling recent human evolution in mice by expression of a selected EDAR variant. Cell 152, 691–702 (2013). This study is an example of how mouse models combined with human GWASs can be used to determine the functional significance of an adaptive variant.
Miller, C. T. et al. cis-regulatory changes in Kit ligand expression and parallel evolution of pigmentation in sticklebacks and humans. Cell 131, 1179–1189 (2007).
Lao, O., de Gruijter, J. M., van Duijn, K., Navarro, A. & Kayser, M. Signatures of positive selection in genes associated with human skin pigmentation as revealed from analyses of single nucleotide polymorphisms. Ann. Hum. Genet. 71, 354–369 (2007).
Varemo, L., Nielsen, J. & Nookaew, I. Enriching the gene set analysis of genome-wide data by incorporating directionality of gene expression and combining statistical hypotheses and methods. Nucleic Acids Res. 41, 4378–4391 (2013).
Grundberg, E. et al. Population genomics in a disease targeted primary cell model. Genome Res. 19, 1942–1952 (2009).
Xiong, Q., Ancona, N., Hauser, E. R., Mukherjee, S. & Furey, T. S. Integrating genetic and gene expression evidence into genome-wide association analysis of gene sets. Genome Res. 22, 386–397 (2012).
Zaykin, D. V., Zhivotovsky, L. A., Czika, W., Shao, S. & Wolfinger, R. D. Combining p-values in large-scale genomics experiments. Pharm. Statist. 6, 217–226 (2007).
Mo, Q. et al. Pattern discovery and cancer gene identification in integrated cancer genomic data. Proc. Natl Acad. Sci. USA 110, 4245–4250 (2013).
Schadt, E. E. et al. An integrative genomics approach to infer causal associations between gene expression and disease. Nature Genet. 37, 710–717 (2005).
Orr, H. A. Testing natural selection versus genetic drift in phenotypic evolution using quantitative trait locus data. Genetics 149, 2099–2104 (1998).
Fraser, H. B. et al. Systematic detection of polygenic cis-regulatory evolution. PLoS Genet. 7, e1002023 (2011).
Leinonen, T., McCairns, R. J., O'Hara, R. B. & Merila, J. QST–FST comparisons: evolutionary and ecological insights from genomic heterogeneity. Nature Rev. Genet. 14, 179–190 (2013).
Lachance, J. et al. Evolutionary history and adaptation from high-coverage whole-genome sequences of diverse African hunter-gatherers. Cell 150, 457–469 (2012). This is the first high-coverage WGS study across diverse African populations, demonstrating signatures of local adaptation, including candidate genes for the short-stature trait in pygmies.
Lanktree, M. B. et al. Meta-analysis of dense genecentric association studies reveals common and uncommon variants associated with height. Am. J. Hum. Genet. 88, 6–18 (2011).
Acknowledgements
The authors thank S. Soi and J. Lachance for their discussions and suggestions. This work was funded by the US National Institutes of Health (NIH) Pioneer Award (DP1OD0644S), the NIH (R01GM076637) and the US National Science Foundation Hominid grant (BCS0827436) to S.A.T.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Related links
Glossary
- Reproductive fitness
-
A measure of how well an organism survives and reproduces; it is generally determined by the genetic contribution of an individual to the gene pool of the next generation.
- Gene pathways
-
Sets of interacting gene products that are related to particular functions, including signalling and metabolic pathways.
- Demographic history
-
The study of population-level changes over time, including changes in population size, migration, and gene flow between populations.
- International HapMap Project
-
A publically available genome-wide data set of common single nucleotide polymorphisms generated from global populations.
- Linkage disequilibrium
-
The non-random association of two or more genetic variants.
- Positive selection
-
A type of natural selection in which a trait that increases reproductive fitness becomes more common over time in a population.
- FST
-
A statistical measure of population structure based on differences in variant frequencies between populations using genotypic data.
- Locus-specific branch length
-
A measure of population structure in one population sample relative to two other population samples based on FST values; it is useful for identifying population-specific adaptation.
- Haplotype
-
A set of genetic variants that are inherited together on a single chromosome.
- Convergent evolution
-
The independent evolution of similar phenotypic traits.
- Locus
-
A region of the genome that contains a particular gene or genetic sequence.
- Pygmy
-
Typically used in the genetic literature to refer to an individual member of a population in which the average male and female stature is <150 cm and <140 cm, respectively. Historically, the term has been used in a pejorative manner; however, some recent populations that culturally identify themselves as Pygmies have revived the term themselves.
- Pleiotropic
-
Pertaining to a gene: affecting multiple phenotypes.
- Epigenetic
-
Pertaining to an inherited phenotypic change: caused by mechanisms other than changes in the underlying DNA sequence, such as DNA methylation or histone modification.
- Metabolomic
-
Pertaining to the metabolome, which is the combined set of metabolites that are present in a given tissue at a given time.
- Transcriptomic
-
Pertaining to the transcriptome, which is the combined set of RNA transcripts that are present in a given tissue at a given time.
- Expression quantitative trait loci
-
Regions of the genome containing genetic polymorphisms that either alter or are in linkage disequilibrium with variants which affect gene regulation to influence the levels of RNA or protein produced.
- Enhancers
-
Regulatory DNA elements that usually bind several transcription factors; they can activate transcription at long distances from the promoter and in an orientation-independent manner.
- DNase I-hypersensitive sites
-
Genetic regions that are sensitive to the DNase I enzyme. These are important regulatory regions because they are available for binding by gene-regulatory proteins.
- Induced pluripotent stem cells
-
Somatically derived cells that have been synthetically induced to behave as pluripotent stem cells.
- Integrative genomics
-
The integration of data from multiple 'omics' platforms.
- QST
-
A statistical measure of population structure or differences in quantitative trait measurements between populations using phenotypic data.
Rights and permissions
About this article
Cite this article
Scheinfeldt, L., Tishkoff, S. Recent human adaptation: genomic approaches, interpretation and insights. Nat Rev Genet 14, 692–702 (2013). https://doi.org/10.1038/nrg3604
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrg3604
This article is cited by
-
Star allele search: a pharmacogenetic annotation database and user-friendly search tool of publicly available 1000 Genomes Project biospecimens
BMC Genomics (2024)
-
Adaptive selection drives TRPP3 loss-of-function in an Ethiopian population
Scientific Reports (2020)
-
Genomes reveal marked differences in the adaptive evolution between orangutan species
Genome Biology (2018)
-
Adaptation of Arabidopsis thaliana to the Yangtze River basin
Genome Biology (2017)
-
Haplotype selection as an adaptive mechanism in the protozoan pathogen Leishmania donovani
Nature Ecology & Evolution (2017)