Key Points
-
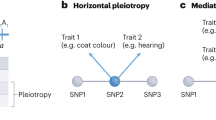
Forward- and reverse-genetic screens in model organisms have revealed that phenotypic traits are typically influenced by the activities of hundreds or thousands of genes and that, conversely, individual genes typically influence many different traits. 'Phenologue' relationships allow these gene–trait connections to be systematically transferred between species.
-
The results of these systematic screens can be used to evaluate and to refine genome-scale methods for linking genes to phenotypes such as integrated 'functional' networks.
-
One reason why the outcome of a particular mutation can vary across individuals is epistasis or genetic interactions between mutations. The systematic mapping of genetic interaction networks in model organisms is providing basic insights into how mutations interact and is leading to the development of computational methods to predict epistasis.
-
The environment can influence the outcome of mutations not only in specific ways, but also in promiscuous ways, such as by altering the activity of molecular chaperones.
-
Whole-genome reverse genetics is the challenge of predicting how individuals vary from their complete genome sequences. Making and experimentally evaluating these predictions in model organisms will lead to the development of improved computational methods for predicting phenotypic variation from genetic variation.
-
In model organisms, mutations often have variable outcomes even in the absence of genetic variation and in a controlled environment. One cause of this is inter-individual variation in the expression or activity of genetic interaction partners, which is termed epigenetic epistasis and occurs, for example, during early embryonic development.
-
Transgenerational genetic and environmental influences can also underlie phenotypic variation. These influences are now being dissected at a molecular level in model organisms.
-
Genetic predictions have both practical and fundamental limitations. More effort should be focused on building clinically useful personalized predictions that incorporate genetic markers and intermediate biomarkers that capture both genetic and non-genetic sources of variance.
Abstract
To what extent can variation in phenotypic traits such as disease risk be accurately predicted in individuals? In this Review, I highlight recent studies in model organisms that are relevant both to the challenge of accurately predicting phenotypic variation from individual genome sequences ('whole-genome reverse genetics') and for understanding why, in many cases, this may be impossible. These studies argue that only by combining genetic knowledge with in vivo measurements of biological states will it be possible to make accurate genetic predictions for individual humans.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Hindorff, L. A. et al. Potential etiologic and functional implications of genome-wide association loci for human diseases and traits. Proc. Natl Acad. Sci. USA 106, 9362–9367 (2009).
Manolio, T. A. et al. Finding the missing heritability of complex diseases. Nature 461, 747–753 (2009).
Burga, A. & Lehner, B. Beyond genotype to phenotype: why the phenotype of an individual cannot always be predicted from their genome sequence and the environment that they experience. FEBS J. 279, 3765–3775 (2012).
Clayton, D. G. Prediction and interaction in complex disease genetics: experience in type 1 diabetes. PLoS Genet. 5, e1000540 (2009).
Roberts, N. J. et al. The predictive capacity of personal genome sequencing. Sci. Transl. Med. 4, 133ra58 (2012). This study provides estimates of the maximum ability of whole-genome sequencing to predict clinically useful risk information for 24 diseases on the basis of analyses of monzygotic twin pairs.
Giaever, G. et al. Functional profiling of the Saccharomyces cerevisiae genome. Nature 418, 387–391 (2002).
Kim, D. U. et al. Analysis of a genome-wide set of gene deletions in the fission yeast Schizosaccharomyces pombe. Nature Biotech. 28, 617–623 (2010).
Baba, T. et al. Construction of Escherichia coli K-12 in-frame, single-gene knockout mutants: the Keio collection. Mol. Syst. Biol. 2, 2006.0008 (2006).
Kamath, R. S. et al. Systematic functional analysis of the Caenorhabditis elegans genome using RNAi. Nature 421, 231–237 (2003). This is the first genome-wide analysis of the effects of gene function inhibition in an animal.
Dietzl, G. et al. A genome-wide transgenic RNAi library for conditional gene inactivation in Drosophila. Nature 448, 151–156 (2007).
Hobert, O. The impact of whole genome sequencing on model system genetics: get ready for the ride. Genetics 184, 317–319 (2010).
Ehrenreich, I. M. et al. Dissection of genetically complex traits with extremely large pools of yeast segregants. Nature 464, 1039–1042 (2010).
Ehrenreich, I. M. et al. Genetic architecture of highly complex chemical resistance traits across four yeast strains. PLoS Genet. 8, e1002570 (2012). This paper describes the detection of more than 800 loci that influence resistance to 13 chemicals in all 6 pairwise crosses of four yeast strains, using extremely large pools of segregants.
Liti, G. & Louis, E. J. Advances in quantitative trait analysis in yeast. PLoS Genet. 8, e1002912 (2012).
Parts, L. et al. Revealing the genetic structure of a trait by sequencing a population under selection. Genome Res. 21, 1131–1138 (2011).
Swinnen, S. et al. Identification of novel causative genes determining the complex trait of high ethanol tolerance in yeast using pooled-segregant whole-genome sequence analysis. Genome Res. 22, 975–984 (2012).
Dudley, A. M., Janse, D. M., Tanay, A., Shamir, R. & Church, G. M. A global view of pleiotropy and phenotypically derived gene function in yeast. Mol. Syst. Biol. 1, 2005.0001 (2005).
Brown, J. A. et al. Global analysis of gene function in yeast by quantitative phenotypic profiling. Mol. Syst. Biol. 2, 2006.0001 (2006).
Hillenmeyer, M. E. et al. The chemical genomic portrait of yeast: uncovering a phenotype for all genes. Science 320, 362–365 (2008).
Wright, S. Physiological and evolutionary theories of dominance. Am. Nat. 68, 24–53 (1934).
McGary, K. L. et al. Systematic discovery of nonobvious human disease models through orthologous phenotypes. Proc. Natl Acad. Sci. USA 107, 6544–6549 (2010). This paper describes the systematic identification of 'phenologues', which are phenotypes in different species that are linked because they are affected by overlapping sets of genes.
Cha, H. J. et al. Evolutionarily repurposed networks reveal the well-known antifungal drug thiabendazole to be a novel vascular disrupting agent. PLoS Biol. 10, e1001379 (2012).
Fraser, A. G. & Marcotte, E. M. A probabilistic view of gene function. Nature Genet. 36, 559–564 (2004).
Lehner, B. & Lee, I. Network-guided genetic screening: building, testing and using gene networks to predict gene function. Brief. Funct. Genom. Proteom. 7, 217–227 (2008).
Lee, I., Date, S. V., Adai, A. T. & Marcotte, E. M. A probabilistic functional network of yeast genes. Science 306, 1555–1558 (2004).
Troyanskaya, O. G., Dolinski, K., Owen, A. B., Altman, R. B. & Botstein, D. A. Bayesian framework for combining heterogeneous data sources for gene function prediction (in Saccharomyces cerevisiae). Proc. Natl Acad. Sci. USA 100, 8348–8353 (2003).
Rhodes, D. R. et al. Probabilistic model of the human protein–protein interaction network. Nature Biotech. 23, 951–959 (2005).
McGary, K. L., Lee, I. & Marcotte, E. M. Broad network-based predictability of Saccharomyces cerevisiae gene loss-of-function phenotypes. Genome Biol. 8, R258 (2007).
Lee, I. et al. A single gene network accurately predicts phenotypic effects of gene perturbation in Caenorhabditis elegans. Nature Genet. 40, 181–188 (2008).
Lee, I., Li, Z. & Marcotte, E. M. An improved, bias-reduced probabilistic functional gene network of baker's yeast, Saccharomyces cerevisiae. PLoS ONE 2, e988 (2007).
Li, Z. et al. Rational extension of the ribosome biogenesis pathway using network-guided genetics. PLoS Biol. 7, e1000213 (2009).
Myers, C. L. et al. Discovery of biological networks from diverse functional genomic data. Genome Biol. 6, R114 (2005).
Chikina, M. D., Huttenhower, C., Murphy, C. T. & Troyanskaya, O. G. Global prediction of tissue-specific gene expression and context-dependent gene networks in Caenorhabditis elegans. PLoS Comput. Biol. 5, e1000417 (2009).
Pena-Castillo, L. et al. A critical assessment of Mus musculus gene function prediction using integrated genomic evidence. Genome Biol. 9 (Suppl. 1), S2 (2008).
Lee, I., Ambaru, B., Thakkar, P., Marcotte, E. M. & Rhee, S. Y. Rational association of genes with traits using a genome-scale gene network for Arabidopsis thaliana. Nature Biotech. 28, 149–156 (2010).
Lee, I. et al. Genetic dissection of the biotic stress response using a genome-scale gene network for rice. Proc. Natl Acad. Sci. USA 108, 18548–18553 (2011).
Lage, K. et al. A human phenome–interactome network of protein complexes implicated in genetic disorders. Nature Biotech. 25, 309–316 (2007).
Lee, I., Blom, U. M., Wang, P. I., Shim, J. E. & Marcotte, E. M. Prioritizing candidate disease genes by network-based boosting of genome-wide association data. Genome Res. 21, 1109–1121 (2011).
Gillis, J. & Pavlidis, P. The impact of multifunctional genes on “guilt by association” analysis. PLoS ONE 6, e17258 (2011).
Sopko, R. et al. Mapping pathways and phenotypes by systematic gene overexpression. Mol. Cell 21, 319–330 (2006).
Vavouri, T., Semple, J. I., Garcia-Verdugo, R. & Lehner, B. Intrinsic protein disorder and interaction promiscuity are widely associated with dosage sensitivity. Cell 138, 198–208 (2009).
Birchler, J. A. & Veitia, R. A. Gene balance hypothesis: connecting issues of dosage sensitivity across biological disciplines. Proc. Natl Acad. Sci. USA 109, 14746–14753 (2012).
Moriya, H., Shimizu-Yoshida, Y. & Kitano, H. In vivo robustness analysis of cell division cycle genes in Saccharomyces cerevisiae. PLoS Genet. 2, e111 (2006).
Bernstein, B. E. et al. An integrated encyclopedia of DNA elements in the human genome. Nature 489, 57–74 (2012).
Cookson, W., Liang, L., Abecasis, G., Moffatt, M. & Lathrop, M. Mapping complex disease traits with global gene expression. Nature Rev. Genet. 10, 184–194 (2009).
Francesconi, M., Jelier, R. & Lehner, B. Integrated genome-scale prediction of detrimental mutations in transcription networks. PLoS Genet. 7, e1002077 (2011).
Gertz, J., Siggia, E. D. & Cohen, B. A. Analysis of combinatorial cis-regulation in synthetic and genomic promoters. Nature 457, 215–218 (2009).
Sharon, E. et al. Inferring gene regulatory logic from high-throughput measurements of thousands of systematically designed promoters. Nature Biotech. 30, 521–530 (2012).
Phillips, P. C. Epistasis—the essential role of gene interactions in the structure and evolution of genetic systems. Nature Rev. Genet. 9, 855–867 (2008).
Lehner, B. Molecular mechanisms of epistasis within and between genes. Trends Genet. 27, 323–331 (2011).
Zuk, O., Hechter, E., Sunyaev, S. R. & Lander, E. S. The mystery of missing heritability: genetic interactions create phantom heritability. Proc. Natl Acad. Sci. USA 109, 1193–1198 (2012).
Lehner, B. Modelling genotype–phenotype relationships and human disease with genetic interaction networks. J. Exp. Biol. 210, 1559–1566 (2007).
Drees, B. L. et al. Derivation of genetic interaction networks from quantitative phenotype data. Genome Biol. 6, R38 (2005).
Phillips, P. C. The language of gene interaction. Genetics 149, 1167–1171 (1998).
Costanzo, M. et al. The genetic landscape of a cell. Science 327, 425–431 (2010). The most comprehensive analysis of epistatic interactions in any organism; the effects on growth are quantified for more than 5 million pairs of mutations in yeast.
Frost, A. et al. Functional repurposing revealed by comparing S. pombe and S. cerevisiae genetic interactions. Cell 149, 1339–1352 (2012).
Ryan, C. J. et al. Hierarchical modularity and the evolution of genetic interactomes across species. Mol. Cell 46, 691–704 (2012).
Lehner, B., Crombie, C., Tischler, J., Fortunato, A. & Fraser, A. G. Systematic mapping of genetic interactions in Caenorhabditis elegans identifies common modifiers of diverse signaling pathways. Nature Genet. 38, 896–903 (2006).
Byrne, A. B. et al. A global analysis of genetic interactions in Caenorhabditis elegans. J. Biol. 6, 8 (2007).
Horn, T. et al. Mapping of signaling networks through synthetic genetic interaction analysis by RNAi. Nature Methods 8, 341–346 (2011).
Tong, A. H. et al. Systematic genetic analysis with ordered arrays of yeast deletion mutants. Science 294, 2364–2368 (2001).
Gerke, J., Lorenz, K. & Cohen, B. Genetic interactions between transcription factors cause natural variation in yeast. Science 323, 498–501 (2009).
Lorenz, K. & Cohen, B. A. Small- and large-effect quantitative trait locus interactions underlie variation in yeast sporulation efficiency. Genetics 192, 1123–1132 (2012).
Brem, R. B. & Kruglyak, L. The landscape of genetic complexity across 5,700 gene expression traits in yeast. Proc. Natl Acad. Sci. USA 102, 1572–1577 (2005).
Brem, R. B., Storey, J. D., Whittle, J. & Kruglyak, L. Genetic interactions between polymorphisms that affect gene expression in yeast. Nature 436, 701–703 (2005). This paper highlights the importance of epistatic interactions between natural variants that influence gene expression.
Dowell, R. D. et al. Genotype to phenotype: a complex problem. Science 328, 469 (2010). By constructing a gene deletion collection for a second laboratory strain of yeast, the authors identify >40 genes that are essential in this strain but not in a previously analysed strain. In most cases, this 'conditional' essentiality in one strain is due to variation in four or more different modifier loci.
Koch, E. N. et al. Conserved rules govern genetic interaction degree across species. Genome Biol. 13, R57 (2012).
Wong, S. L. et al. Combining biological networks to predict genetic interactions. Proc. Natl Acad. Sci. USA 101, 15682–15687 (2004).
Lee, I. et al. Predicting genetic modifier loci using functional gene networks. Genome Res. 20, 1143–1153 (2010).
Kelley, R. & Ideker, T. Systematic interpretation of genetic interactions using protein networks. Nature Biotech. 23, 561–566 (2005).
Ulitsky, I. & Shamir, R. Pathway redundancy and protein essentiality revealed in the Saccharomyces cerevisiae interaction networks. Mol. Syst. Biol. 3, 104 (2007).
Bellay, J. et al. Putting genetic interactions in context through a global modular decomposition. Genome Res. 21, 1375–1387 (2011).
Hess, D. C. et al. Computationally driven, quantitative experiments discover genes required for mitochondrial biogenesis. PLoS Genet. 5, e1000407 (2009).
Gerke, J., Lorenz, K., Ramnarine, S. & Cohen, B. Gene–environment interactions at nucleotide resolution. PLoS Genet. 6, e1001144 (2010).
St Onge, R. P. et al. Systematic pathway analysis using high-resolution fitness profiling of combinatorial gene deletions. Nature Genet. 39, 199–206 (2007).
Bandyopadhyay, S., Kelley, R., Krogan, N. J. & Ideker, T. Functional maps of protein complexes from quantitative genetic interaction data. PLoS Comput. Biol. 4, e1000065 (2008).
Harrison, R., Papp, B., Pal, C., Oliver, S. G. & Delneri, D. Plasticity of genetic interactions in metabolic networks of yeast. Proc. Natl Acad. Sci. USA 104, 2307–2312 (2007).
Dixon, S. J. et al. Significant conservation of synthetic lethal genetic interaction networks between distantly related eukaryotes. Proc. Natl Acad. Sci. USA 105, 16653–16658 (2008).
Tischler, J., Lehner, B. & Fraser, A. G. Evolutionary plasticity of genetic interaction networks. Nature Genet. 40, 390–391 (2008).
Roguev, A. et al. Conservation and rewiring of functional modules revealed by an epistasis map in fission yeast. Science 322, 405–410 (2008).
Lindquist, S. Protein folding sculpting evolutionary change. Cold Spring Harb. Symp. Quant. Biol. 74, 103–108 (2009).
Zhao, R. et al. Navigating the chaperone network: an integrative map of physical and genetic interactions mediated by the hsp90 chaperone. Cell 120, 715–727 (2005).
Rutherford, S. L. & Lindquist, S. Hsp90 as a capacitor for morphological evolution. Nature 396, 336–342 (1998).
Queitsch, C., Sangster, T. A. & Lindquist, S. Hsp90 as a capacitor of phenotypic variation. Nature 417, 618–624 (2002).
Casanueva, M. O., Burga, A. & Lehner, B. Fitness trade-offs and environmentally induced mutation buffering in isogenic C. elegans. Science 335, 82–85 (2012).
Jelier, R., Semple, J. I., Garcia-Verdugo, R. & Lehner, B. Predicting phenotypic variation in yeast from individual genome sequences. Nature Genet. 43, 1270–1274 (2011). This paper reports the application of whole-genome reverse genetics: phenotypic predictions are made from the complete genome sequences of yeast strains and the accuracy of these predictions are evaluated by experimentation. Predictions are generally good, even for genetically complex traits, when the genes linked to the trait are evaluated as reliable using a gene network.
Liti, G. et al. Population genomics of domestic and wild yeasts. Nature 458, 337–341 (2009).
Baker, M. Functional genomics: the changes that count. Nature 482, 257–262 (2012).
Gartner, K. A third component causing random variability beside environment and genotype. A reason for the limited success of a 30 year long effort to standardize laboratory animals? Lab. Anim. 24, 71–77 (1990).
Eldar, A. et al. Partial penetrance facilitates developmental evolution in bacteria. Nature 460, 510–514 (2009).
Raj, A., Rifkin, S. A., Andersen, E. & van Oudenaarden, A. Variability in gene expression underlies incomplete penetrance. Nature 463, 913–918 (2010).
Burga, A., Casanueva, M. O. & Lehner, B. Predicting mutation outcome from early stochastic variation in genetic interaction partners. Nature 480, 250–253 (2011). Whether an inherited mutation affects genetically identical individuals or not is predicted by inter-individual variation in the expression of a specific and a promiscuous genetic interaction partner during early embryonic development.
Hales, C. N. & Barker, D. J. Type 2 (non-insulin-dependent) diabetes mellitus: the thrifty phenotype hypothesis. Diabetologia 35, 595–601 (1992).
Wang, T. J. et al. Metabolite profiles and the risk of developing diabetes. Nature Med. 17, 448–453 (2011).
Seidel, H. S., Rockman, M. V. & Kruglyak, L. Widespread genetic incompatibility in C. elegans maintained by balancing selection. Science 319, 589–594 (2008).
Xing, Y. et al. Evidence for transgenerational transmission of epigenetic tumor susceptibility in Drosophila. PLoS Genet. 3, 1598–1606 (2007).
Frazier, H. N. & Roth, M. B. Adaptive sugar provisioning controls survival of C. elegans embryos in adverse environments. Curr. Biol. 19, 859–863 (2009).
Ng, S. F. et al. Chronic high-fat diet in fathers programs β-cell dysfunction in female rat offspring. Nature 467, 963–966 (2010).
Carone, B. R. et al. Paternally induced transgenerational environmental reprogramming of metabolic gene expression in mammals. Cell 143, 1084–1096 (2010).
Jablonka, E. & Raz, G. Transgenerational epigenetic inheritance: prevalence, mechanisms, and implications for the study of heredity and evolution. Q. Rev. Biol. 84, 131–176 (2009).
Painter, R. C. et al. Transgenerational effects of prenatal exposure to the Dutch famine on neonatal adiposity and health in later life. BJOG 115, 1243–1249 (2008).
Ferguson-Smith, A. C. Genomic imprinting: the emergence of an epigenetic paradigm. Nature Rev. Genet. 12, 565–575 (2011).
Ashe, A. et al. piRNAs can trigger a multigenerational epigenetic memory in the germline of C. elegans. Cell 150, 88–99 (2012).
Shirayama, M. et al. piRNAs initiate an epigenetic memory of nonself RNA in the C. elegans germline. Cell 150, 65–77 (2012).
Buckley, B. A. et al. A nuclear Argonaute promotes multigenerational epigenetic inheritance and germline immortality. Nature 489, 447–451 (2012). References 103–105 establish that piRNA-triggered gene silencing is stably transmitted across many generations in C. elegans.
Greer, E. L. et al. Transgenerational epigenetic inheritance of longevity in Caenorhabditis elegans. Nature 479, 365–371 (2011).
Huang, N., Lee, I., Marcotte, E. M. & Hurles, M. E. Characterising and predicting haploinsufficiency in the human genome. PLoS Genet. 6, e1001154 (2010).
Liu, C., van Dyk, D., Li, Y., Andrews, B. & Rao, H. A genome-wide synthetic dosage lethality screen reveals multiple pathways that require the functioning of ubiquitin-binding proteins Rad23 and Dsk2. BMC Biol. 7, 75 (2009).
Burga, A. & Lehner, B. Predicting phenotypes from genotypes, phenotypes and a combination of the two. Curr. Opin. Biotech. (in the press).
Acknowledgements
Our research is funded by the European Research Council (ERC), MINECO Plan Nacional grants BFU2008-00365 and BFU2011-26206, ERASysBio+ ERANET project EUI2009-04059 GRAPPLE, the European Molecular Biology Organization (EMBO) Young Investigator Program, EU Framework 7 project 277899 4DCellFate and the EMBL/CRG Systems Biology Program.
Author information
Authors and Affiliations
Ethics declarations
Competing interests
The author declares no competing financial interests.
Related links
FURTHER INFORMATION
Glossary
- Modules
-
Groups of genes or proteins in a network that have strong interactions among themselves and that carry out particular functions largely independently of other genes or proteins. Mutations in genes from a module often have similar phenotypic consequences.
- Orthologous
-
A gene in one species is orthologous to a gene in another species if they are derived from a common ancestor.
- Disordered regions
-
Regions of proteins that are intrinsically unfolded; that is, they are without a well-defined tertiary structure under physiological conditions.
- Expression quantitative trait loci
-
(eQTLs). Regions of the genome containing genetic polymorphisms that alter how genes are regulated, influencing how much RNA or protein they produce.
- Major- and minor-effect loci
-
Regions of the genome containing genetic polymorphisms that account for a large or small proportion of variance in a particular phenotype, respectively.
- Isogenic
-
Lacking genetic variation. Some laboratory animals, such as Caenorhabditis elegans and mice, are inbred and so siblings have identical genome sequences except for de novo mutations arising in each generation.
- Haploinsufficiency
-
A gene is haploinsufficient if removal of one of the two copies in a diploid organism has a detectable effect on fitness or a phenotype.
- Dominance
-
The extent to which one allele of a gene exerts its effects irrespective of a second allele in diploid organisms. Complete dominance implies that the heterozgygote has a phenotype that is indistinguishable from that of the dominant homozygote. Overdominance implies that the phenotype of the heterozygote lies outside the range of both homozygote parents.
Rights and permissions
About this article
Cite this article
Lehner, B. Genotype to phenotype: lessons from model organisms for human genetics. Nat Rev Genet 14, 168–178 (2013). https://doi.org/10.1038/nrg3404
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrg3404
This article is cited by
-
Heterogeneity in maternal mRNAs within clutches of eggs in response to thermal stress during the embryonic stage
BMC Ecology and Evolution (2024)
-
DeepGAMI: deep biologically guided auxiliary learning for multimodal integration and imputation to improve genotype–phenotype prediction
Genome Medicine (2023)
-
The Insertion in the 3′ UTR of Pmel17 Is the Causal Variant for Golden Skin Color in Tilapia
Marine Biotechnology (2022)
-
Intellectual disability genomics: current state, pitfalls and future challenges
BMC Genomics (2021)
-
Biophysical ambiguities prevent accurate genetic prediction
Nature Communications (2020)