Key Points
-
MicroRNAs (miRNAs) are small RNAs (∼22 nucleotides long) that post-transcriptionally regulate the expression of thousands of genes in a broad range of organisms.
-
miRNA expression profiling is useful for identifying miRNAs that are important in the regulation of a range of processes, including organismal development, tissue differentiation and disease pathology.
-
miRNAs show promise as biomarkers for various diseases.
-
miRNAs are more stable than mRNAs in many specimen types and are more readily measured than proteins. However, sample type, processing and RNA extraction methods can have a substantial impact on the results of miRNA profiling, and therefore quality and quantity assessment is recommended.
-
Biogenesis of miRNAs occurs through multiple steps and includes the intermediate primary miRNA and precursor miRNA forms, as well as post-transcriptional nucleotide additions and deletions, leading to 'isomiRs'. Choice of platform and analysis in miRNA profiling should include consideration of the need to distinguish between different forms of miRNAs.
-
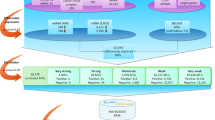
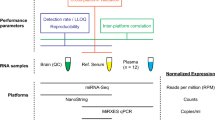
Three main approaches are currently well established for miRNA profiling: quantitative reverse transcription PCR (qRT-PCR), hybridization-based methods (for example, DNA microarrays) and high-throughput sequencing (that is, RNA sequencing). The optimal choice of platform depends on the specific experimental goals.
-
Analysis of miRNA-profiling data typically includes data processing, data quality assessment, data normalization and calculation of differential expression. The optimal approach to data analysis depends on the platform selected and the nature of the experiment.
Abstract
MicroRNAs (miRNAs) are small RNAs that post-transcriptionally regulate the expression of thousands of genes in a broad range of organisms in both normal physiological contexts and in disease contexts. miRNA expression profiling is gaining popularity because miRNAs, as key regulators in gene expression networks, can influence many biological processes and also show promise as biomarkers for disease. Technological advances have spawned a multitude of platforms for miRNA profiling, and an understanding of the strengths and pitfalls of different approaches can aid in their effective use. Here, we review the major considerations for carrying out and interpreting results of miRNA-profiling studies.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Lee, R. C., Feinbaum, R. L. & Ambros, V. The C. elegans heterochronic gene lin-4 encodes small RNAs with antisense complementarity to lin-14. Cell 75, 843–854 (1993).
Wightman, B., Ha, I. & Ruvkun, G. Posttranscriptional regulation of the heterochronic gene lin-14 by lin-4 mediates temporal pattern formation in C. elegans. Cell 75, 855–862 (1993).
Reinhart, B. J. et al. The 21-nucleotide let-7 RNA regulates developmental timing in Caenorhabditis elegans. Nature 403, 901–906 (2000).
Wienholds, E. et al. MicroRNA expression in zebrafish embryonic development. Science 309, 310–311 (2005).
Alvarez-Garcia, I. & Miska, E. A. MicroRNA functions in animal development and human disease. Development 132, 4653–4662 (2005).
Lu, J. et al. MicroRNA expression profiles classify human cancers. Nature 435, 834–838 (2005). This was one of the earliest large-scale studies demonstrating the use of tissue-based miRNA profiles to classify human cancers.
Rosenfeld, N. et al. MicroRNAs accurately identify cancer tissue origin. Nature Biotech. 26, 462–469 (2008). Results from this study were used to create a commercial miRNA profile-based 'tissue of origin' test to aid clinicians in the classification of cancers for which the site of origin is otherwise not discernible.
Boeri, M. et al. MicroRNA signatures in tissues and plasma predict development and prognosis of computed tomography detected lung cancer. Proc. Natl Acad. Sci. USA 108, 3713–3718 (2011).
Tili, E., Michaille, J. J., Costinean, S. & Croce, C. M. MicroRNAs, the immune system and rheumatic disease. Nature Clin. Pract. Rheum. 4, 534–541 (2008).
Courts, C. & Madea, B. Specific micro-RNA signatures for the detection of saliva and blood in forensic body-fluid identification. J. Forensic Sci. 56, 1464–1470 (2011).
Krol, J., Loedige, I. & Filipowicz, W. The widespread regulation of microRNA biogenesis, function and decay. Nature Rev. Genet. 11, 597–610 (2010). This comprehensive Review nicely outlines aspects of miRNA biogenesis that are beyond the scope of the present Review.
Davis-Dusenbery, B. N. & Hata, A. Mechanisms of control of microRNA biogenesis. J. Biochem. 148, 381–392 (2010).
Ragan, C., Zuker, M. & Ragan, M. A. Quantitative prediction of miRNA-mRNA interaction based on equilibrium concentrations. PLoS Comput. Biol. 7, e1001090 (2011).
Liang, Y., Ridzon, D., Wong, L. & Chen, C. Characterization of microRNA expression profiles in normal human tissues. BMC Genomics 8, 166 (2007).
Griffiths-Jones, S., Grocock, R. J., van Dongen, S., Bateman, A. & Enright, A. J. miRBase: microRNA sequences, targets and gene nomenclature. Nucleic Acids Res. 34, D140–D144 (2006).
Griffiths-Jones, S., Saini, H. K., van Dongen, S. & Enright, A. J. miRBase: tools for microRNA genomics. Nucleic Acids Res. 36, D154–D158 (2008).
Griffiths-Jones, S. The microRNA Registry. Nucleic Acids Res. 32, D109–D111 (2004).
Katoh, T. et al. Selective stabilization of mammalian microRNAs by 3′ adenylation mediated by the cytoplasmic poly(A) polymerase GLD-2. Genes Dev. 23, 433–438 (2009).
Jones, M. R. et al. Zcchc11-dependent uridylation of microRNA directs cytokine expression. Nature Cell Biol. 11, 1157–1163 (2009).
Kozomara, A. & Griffiths-Jones, S. miRBase: integrating microRNA annotation and deep-sequencing data. Nucleic Acids Res. 39, D152–D157 (2011).
Wyman, S. K. et al. Post-transcriptional generation of miRNA variants by multiple nucleotidyl transferases contributes to miRNA transcriptome complexity. Genome Res. 21, 1450–1461 (2011).
Cloonan, N. et al. MicroRNAs and their isomiRs function cooperatively to target common biological pathways. Genome Biol. 12, R126 (2011).
Ibberson, D., Benes, V., Muckenthaler, M. U. & Castoldi, M. RNA degradation compromises the reliability of microRNA expression profiling. BMC Biotechnol. 9, 102 (2009).
Podolska, A., Kaczkowski, B., Litman, T., Fredholm, M. & Cirera, S. How the RNA isolation method can affect microRNA microarray results. Acta Biochim. Pol. 58, 535–540 (2011).
Wang, W. X. et al. Focus on RNA isolation: obtaining RNA for microRNA (miRNA) expression profiling analyses of neural tissue. Biochim. Biophys. Acta 1779, 749–757 (2008).
Hammerle-Fickinger, A. et al. Validation of extraction methods for total RNA and miRNA from bovine blood prior to quantitative gene expression analyses. Biotechnol. Lett. 32, 35–44 (2010).
Weber, J. A. et al. The microRNA spectrum in 12 body fluids. Clin. Chem. 56, 1733–1741 (2010).
Accerbi, M. et al. Methods for isolation of total RNA to recover miRNAs and other small RNAs from diverse species. Methods Mol. Biol. 592, 31–50 (2010).
Castoldi, M., Benes, V., Hentze, M. W. & Muckenthaler, M. U. miChip: a microarray platform for expression profiling of microRNAs based on locked nucleic acid (LNA) oligonucleotide capture probes. Methods 43, 146–152 (2007).
Aravin, A. & Tuschl, T. Identification and characterization of small RNAs involved in RNA silencing. FEBS Lett. 579, 5830–5840 (2005).
Doleshal, M. et al. Evaluation and validation of total RNA extraction methods for microRNA expression analyses in formalin-fixed, paraffin-embedded tissues. J. Mol. Diagn. 10, 203–211 (2008).
Xi, Y. et al. Systematic analysis of microRNA expression of RNA extracted from fresh frozen and formalin-fixed paraffin-embedded samples. RNA 13, 1668–1674 (2007).
Dahlgaard, J. et al. Analytical variables influencing the performance of a miRNA based laboratory assay for prediction of relapse in stage I non-small cell lung cancer (NSCLC). BMC Res. Notes 4, 424 (2011).
Mitchell, P. S. et al. Circulating microRNAs as stable blood-based markers for cancer detection. Proc. Natl Acad. Sci. USA 105, 10513–10518 (2008). This study demonstrated that miRNAs are present at high concentrations in human serum and plasma, are surprisingly stable and are changed in the setting of disease.
McDonald, J. S., Milosevic, D., Reddi, H. V., Grebe, S. K. & Algeciras-Schimnich, A. Analysis of circulating microRNA: preanalytical and analytical challenges. Clin. Chem. 57, 833–840 (2011).
Duttagupta, R., Jiang, R., Gollub, J., Getts, R. C. & Jones, K. W. Impact of cellular miRNAs on circulating miRNA biomarker signatures. PLoS ONE 6, e20769 (2011).
Pritchard, C. C. et al. Blood cell origin of circulating microRNAs: a cautionary note for cancer biomarker studies. Cancer Prev. Res. 5, 492–497 (2011).
Arroyo, J. D. et al. Argonaute2 complexes carry a population of circulating microRNAs independent of vesicles in human plasma. Proc. Natl Acad. Sci. USA 108, 5003–5008 (2011).
Becker, C., Hammerle-Fickinger, A., Riedmaier, I. & Pfaffl, M. W. mRNA and microRNA quality control for RT-qPCR analysis. Methods 50, 237–243 (2010).
Kroh, E. M., Parkin, R. K., Mitchell, P. S. & Tewari, M. Analysis of circulating microRNA biomarkers in plasma and serum using quantitative reverse transcription-PCR (qRT-PCR). Methods 50, 298–301 (2010).
Wark, A. W., Lee, H. J. & Corn, R. M. Multiplexed detection methods for profiling microRNA expression in biological samples. Angew. Chem. Int. Edn Engl. 47, 644–652 (2008).
Newman, M. A., Mani, V. & Hammond, S. M. Deep sequencing of microRNA precursors reveals extensive 3′ end modification. RNA 17, 1795–1803 (2011). This study demonstrated the usefulness of RNA-seq for pre-miRNA analysis and identified end modifications to the miRNA precursor, which may lead to sequence variation of mature miRNA forms as well.
Chiang, H. R. et al. Mammalian microRNAs: experimental evaluation of novel and previously annotated genes. Genes Dev. 24, 992–1009 (2010).
Wei, C., Salichos, L., Wittgrove, C. M., Rokas, A. & Patton, J. G. Transcriptome-wide analysis of small RNA expression in early zebrafish development. RNA (2012).
Burroughs, A. M. et al. A comprehensive survey of 3′ animal miRNA modification events and a possible role for 3′ adenylation in modulating miRNA targeting effectiveness. Genome Res. 20, 1398–1410 (2010).
Ach, R. A., Wang, H. & Curry, B. Measuring microRNAs: comparisons of microarray and quantitative PCR measurements, and of different total RNA prep methods. BMC Biotechnology 8, 69 (2008). This study systematically addressed the effect of different RNA preparation methods on miRNA profiling.
Chen, Y., Gelfond, J. A., McManus, L. M. & Shireman, P. K. Reproducibility of quantitative RT-PCR array in miRNA expression profiling and comparison with microarray analysis. BMC Genomics 10, 407 (2009).
Jensen, S. G. et al. Evaluation of two commercial global miRNA expression profiling platforms for detection of less abundant miRNAs. BMC Genomics 12, 435 (2011).
Git, A. et al. Systematic comparison of microarray profiling, real-time PCR, and next-generation sequencing technologies for measuring differential microRNA expression. RNA 16, 991–1006 (2010).
Pradervand, S. et al. Concordance among digital gene expression, microarrays, and qPCR when measuring differential expression of microRNAs. BioTechniques 48, 219–222 (2010).
Chamnongpol, S., Maroney, P. A. & Nilsen, T. W. A rapid, quantitative assay for direct detection of microRNAs and other small RNAs using splinted ligation. Methods Mol. Biol. 667, 3–17 (2010).
Maroney, P. A., Chamnongpol, S., Souret, F. & Nilsen, T. W. Direct detection of small RNAs using splinted ligation. Nature Protoc. 3, 279–287 (2008).
Chapin, S. C. & Doyle, P. S. Ultrasensitive multiplexed microRNA quantification on encoded gel microparticles using rolling circle amplification. Anal. Chem. 83, 7179–7185 (2011).
Harcourt, E. M. & Kool, E. T. Amplified microRNA detection by templated chemistry. Nucleic Acids Res. 25 Jan 2012 (doi:10.1093/nar/gkr1313).
Mashimo, Y., Mie, M., Suzuki, S. & Kobatake, E. Detection of small RNA molecules by a combination of branched rolling circle amplification and bioluminescent pyrophosphate assay. Anal. Bioanal. Chem. 401, 221–227 (2011).
Zhou, Y. et al. A dumbbell probe-mediated rolling circle amplification strategy for highly sensitive microRNA detection. Nucleic Acids Res. 38, e156 (2010).
Chan, H. M., Chan, L. S., Wong, R. N. & Li, H. W. Direct quantification of single-molecules of microRNA by total internal reflection fluorescence microscopy. Anal. Chem. 82, 6911–6918 (2010).
Jiang, L., Duan, D., Shen, Y. & Li, J. Direct microRNA detection with universal tagged probe and time-resolved fluorescence technology. Biosens. Bioelectron. 34, 291–295 (2012).
Nasheri, N. et al. An enzyme-linked assay for the rapid quantification of microRNAs based on the viral suppressor of RNA silencing protein p19. Anal. Biochem. 412, 165–172 (2011).
Peng, Y. & Gao, Z. Amplified detection of microRNA based on ruthenium oxide nanoparticle-initiated deposition of an insulating film. Anal. Chem. 83, 820–827 (2011).
Persat, A., Chivukula, R. R., Mendell, J. T. & Santiago, J. G. Quantification of global microRNA abundance by selective isotachophoresis. Anal. Chem. 82, 9631–9635 (2010).
Persat, A. & Santiago, J. G. MicroRNA profiling by simultaneous selective isotachophoresis and hybridization with molecular beacons. Anal. Chem. 83, 2310–2316 (2011).
Qavi, A. J., Kindt, J. T., Gleeson, M. A. & Bailey, R. C. Anti-DNA:RNA antibodies and silicon photonic microring resonators: increased sensitivity for multiplexed microRNA detection. Anal. Chem. 83, 5949–5956 (2011).
Sipova, H. et al. Surface plasmon resonance biosensor for rapid label-free detection of microribonucleic acid at subfemtomole level. Anal. Chem. 82, 10110–10115 (2010).
Nelson, P. T., Wang, W. X., Wilfred, B. R. & Tang, G. Technical variables in high-throughput miRNA expression profiling: much work remains to be done. Biochim. Biophys. Acta 1779, 758–765 (2008).
Castoldi, M. et al. A sensitive array for microRNA expression profiling (miChip) based on locked nucleic acids (LNA). RNA 12, 913–920 (2006).
Liu, C. G., Calin, G. A., Volinia, S. & Croce, C. M. MicroRNA expression profiling using microarrays. Nature Protoc. 3, 563–578 (2008).
Yin, J. Q., Zhao, R. C. & Morris, K. V. Profiling microRNA expression with microarrays. Trends Biotechnol. 26, 70–76 (2008).
Geiss, G. K. et al. Direct multiplexed measurement of gene expression with color-coded probe pairs. Nature Biotech. 26, 317–325 (2008).
Metzker, M. L. Sequencing technologies — the next generation. Nature Rev. Genet. 11, 31–46 (2010). This is an excellent Review of the major next-generation sequencing platforms.
Ambros, V. et al. A uniform system for microRNA annotation. RNA 9, 277–279 (2003).
Ruby, J. G. et al. Large-scale sequencing reveals 21U-RNAs and additional microRNAs and endogenous siRNAs in C. elegans. Cell 127, 1193–207 (2006).
Linsen, S. E. et al. Limitations and possibilities of small RNA digital gene expression profiling. Nature Methods 6, 474–476 (2009).
Tian, G. et al. Sequencing bias: comparison of different protocols of microRNA library construction. BMC Biotechnology 10, 64 (2010).
Hafner, M. et al. RNA-ligase-dependent biases in miRNA representation in deep-sequenced small RNA cDNA libraries. RNA 17, 1697–1712 (2011).
Kapranov, P. et al. New class of gene-termini-associated human RNAs suggests a novel RNA copying mechanism. Nature 466, 642–646 (2010).
Schmittgen, T. D. et al. Real-time PCR quantification of precursor and mature microRNA. Methods 44, 31–38 (2008).
Benes, V. & Castoldi, M. Expression profiling of microRNA using real-time quantitative PCR, how to use it and what is available. Methods 50, 244–249 (2010).
Jiang, J., Lee, E. J., Gusev, Y. & Schmittgen, T. D. Real-time expression profiling of microRNA precursors in human cancer cell lines. Nucleic Acids Res. 33, 5394–5403 (2005).
Schmittgen, T. D., Jiang, J., Liu, Q. & Yang, L. A high-throughput method to monitor the expression of microRNA precursors. Nucleic Acids Res. 32, e43 (2004). This was one of the first studies to describe methods for profiling pre-miRNAs.
Chugh, P., Tamburro, K. & Dittmer, D. P. Profiling of pre-micro RNAs and microRNAs using quantitative real-time PCR (qPCR) arrays. J. Vis. Exp. 3 Dec 2010 (doi: 10.3791/2210).
Thomas, M., Lieberman, J. & Lal, A. Desperately seeking microRNA targets. Nature Struct. Mol. Biol. 17, 1169–1174 (2010).
Thomson, D. W., Bracken, C. P. & Goodall, G. J. Experimental strategies for microRNA target identification. Nucleic Acids Res. 39, 6845–6853 (2011).
Garber, M., Grabherr, M. G., Guttman, M. & Trapnell, C. Computational methods for transcriptome annotation and quantification using RNA-seq. Nature Methods 8, 469–477 (2011).
Sarkar, D. et al. Quality assessment and data analysis for microRNA expression arrays. Nucleic Acids Res. 37, e17 (2009).
Leek, J. T. et al. Tackling the widespread and critical impact of batch effects in high-throughput data. Nature Rev. Genet. 11, 733–739 (2010).
Hutson, S. Data handling errors spur debate over clinical trial. Nature Med. 16, 618 (2010).
Meyer, S. U., Pfaffl, M. W. & Ulbrich, S. E. Normalization strategies for microRNA profiling experiments: a 'normal' way to a hidden layer of complexity? Biotechnol. Lett. 32, 1777–1788 (2010).
Peltier, H. J. & Latham, G. J. Normalization of microRNA expression levels in quantitative RT-PCR assays: identification of suitable reference RNA targets in normal and cancerous human solid tissues. RNA 14, 844–852 (2008).
Pradervand, S. et al. Impact of normalization on miRNA microarray expression profiling. RNA 15, 493–501 (2009).
Wylie, D., Shelton, J., Choudhary, A. & Adai, A. T. A novel mean-centering method for normalizing microRNA expression from high-throughput RT-qPCR data. BMC Res. Notes 4, 555 (2011).
Risso, D., Massa, M. S., Chiogna, M. & Romualdi, C. A modified LOESS normalization applied to microRNA arrays: a comparative evaluation. Bioinformatics 25, 2685–2691 (2009).
Mestdagh, P. et al. A novel and universal method for microRNA RT-qPCR data normalization. Genome Biol. 10, R64 (2009).
Gunaratne, P. H., Creighton, C. J., Watson, M. & Tennakoon, J. B. Large-scale integration of microRNA and gene expression data for identification of enriched microRNA–mRNA associations in biological systems. Methods Mol. Biol. 667, 297–315 (2010).
Sales, G. et al. MAGIA, a web-based tool for miRNA and genes integrated analysis. Nucleic Acids Res. 38, W352–W359 (2010).
Huang, G. T., Athanassiou, C. & Benos, P. V. mirConnX: condition-specific mRNA–microRNA network integrator. Nucleic Acids Res. 39, W416–W423 (2011).
Jiang, Q. et al. miR2Disease: a manually curated database for microRNA deregulation in human disease. Nucleic Acids Res. 37, D98–D104 (2009).
Lagana, A. et al. miRo: a miRNA knowledge base. Database 2009, bap008 (2009).
Ulitsky, I. et al. Expander: from expression microarrays to networks and functions. Nature Protoc. 5, 303–322 (2010).
Bizuayehu, T. T. et al. Differential expression patterns of conserved miRNAs and isomiRs during Atlantic halibut development. BMC Genomics 13, 11 (2012).
Yao, Y. et al. Cloning and characterization of microRNAs from wheat (Triticum aestivum L.). Genome Biol. 8, R96 (2007).
Ule, J., Jensen, K., Mele, A. & Darnell, R. B. CLIP: a method for identifying protein-RNA interaction sites in living cells. Methods 37, 376–386 (2005).
Ule, J. et al. CLIP identifies Nova-regulated RNA networks in the brain. Science 302, 1212–1215 (2003).
Hafner, M. et al. PAR-CliP—a method to identify transcriptome-wide the binding sites of RNA binding proteins. J. Vis. Exp. 2 Jul 2010 (doi:10.3791/2034).
Hafner, M. et al. Transcriptome-wide identification of RNA-binding protein and microRNA target sites by PAR-CLIP. Cell 141, 129–141 (2010).
Lopes, C. T. et al. Cytoscape Web: an interactive web-based network browser. Bioinformatics 26, 2347–2348 (2010).
Buckley, P. G. et al. Chromosomal and microRNA expression patterns reveal biologically distinct subgroups of 11q- neuroblastoma. Clin. Cancer Res. 16, 2971–2978 (2010).
Ji, J. et al. MicroRNA expression, survival, and response to interferon in liver cancer. New Engl. J. Med. 361, 1437–1447 (2009).
Lai, C. Y. et al. MicroRNA expression aberration as potential peripheral blood biomarkers for schizophrenia. PLoS ONE 6, e21635 (2011).
Saal, S. & Harvey, S. J. MicroRNAs and the kidney: coming of age. Curr. Opin. Nephrol. Hypertens. 18, 317–323 (2009).
Guerau-de-Arellano, M., Alder, H., Ozer, H. G., Lovett-Racke, A. & Racke, M. K. miRNA profiling for biomarker discovery in multiple sclerosis: from microarray to deep sequencing. J. Neuroimmunol. 9 Nov 2011 (doi:http://dx.doi.org/10.1016/j.jneuroim.2011.10.006).
Ferracin, M. et al. MicroRNA profiling for the identification of cancers with unknown primary tissue-of-origin. J. Pathol. 225, 43–53 (2011).
Varadhachary, G. R. et al. Prospective gene signature study using microRNA to identify the tissue of origin in patients with carcinoma of unknown primary. Clin. Cancer Res. 17, 4063–4070 (2011).
Lawrie, C. H. et al. Detection of elevated levels of tumour-associated microRNAs in serum of patients with diffuse large B-cell lymphoma. Br. J. Haematol. 141, 672–675 (2008).
Chen, X. et al. Characterization of microRNAs in serum: a novel class of biomarkers for diagnosis of cancer and other diseases. Cell Res. 18, 997–1006 (2008).
Kim, D. J. et al. Plasma components affect accuracy of circulating cancer-related microRNA quantitation. J. Mol. Diagn. 14, 71–80 (2012).
Watson, A. K. & Witwer, K. W. Do platform-specific factors explain microRNA profiling disparities? Clin. Chem. 58, 472–474 (2011).
Zubakov, D. et al. MicroRNA markers for forensic body fluid identification obtained from microarray screening and quantitative RT-PCR confirmation. Int. J. Legal Med. 124, 217–226 (2010).
Stark, M. S. et al. Characterization of the melanoma miRNAome by deep sequencing. PLoS ONE 5, e9685 (2010).
Hoefig, K. P. & Heissmeyer, V. Measuring microRNA expression in size-limited FACS-sorted and microdissected samples. Methods Mol. Biol. 667, 47–63 (2010).
Acknowledgements
We thank J. Tait for helpful comments on the manuscript. We acknowledge the many authors whose work could not be cited owing to space constraints. M.T. acknowledges generous support from a Damon Runyon-Rachleff Innovation Award, a Stand Up To Cancer Innovative Research Grant (Innovative Research Grant SU2C-AACR-IRG1109), a Prostate Cancer Foundation Creativity Award and funding from the US National Institutes of Health (R01DK085714; P50 CA83636; 5 P30 CA015704; P50 CA97186) and Department of Defense (PC074012; OC080159).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
Muneesh Tewari is a named inventor on patent applications relating to circulating microRNA. He has served on the scientific advisory boards of Wafergen, Inc. and Combimatrix, Inc. within the last three years and has had a past research collaboration with scientists at Nanostring, Inc. Neither Colin C. Pritchard nor Heather H. Cheng declares any competing financial interests.
Supplementary information
Supplementary information S1 (figure)
miRNA Biogenesis. (PDF 200 kb)
Supplementary information S2 (Box)
MicroRNA Nomenclature (PDF 178 kb)
Supplementary information S3 (Table)
Selected Emerging Technologies for miRNA Profiling* (PDF 198 kb)
Glossary
- Primary miRNA
-
(pri-miRNA). The initial transcription product of microRNA (miRNA) genes. Pri-miRNAs are generally >100 nucleotides long (frequently a few kilobases long) and may contain one or more miRNA stem–loops that are processed by the miRNA biogenesis machinery.
- Precursor miRNA
-
(pre-miRNA). A hairpin precursor of microRNA (miRNA) that is formed by the cleavage of the primary miRNA transcript by the Drosha–DGCR8 protein complex. Precursor miRNAs are typically ∼70–100 nucleotides long.
- miRBase
-
A comprehensive database of microRNA sequences and nomenclature from plant and animal species.
- Pre-analytic variables
-
Variables that occur before sample assay. For example, the time elapsed between when a blood sample is drawn from the patient and when it is processed by the laboratory is a pre-analytic variable.
- Argonaute
-
(AGO). These proteins are the central components of RNA-silencing mechanisms. They provide the platform for target–mRNA recognition by short guide RNA strands (for example, miRNAs) and, in the case of AGO2 (in humans), the catalytic activity for mRNA cleavage.
- Seed region
-
The six or seven nucleotides between the nucleotides positions 2–7 or 2–8 of the microRNA (miRNA) 5′ end that determine, in large measure, miRNA target selection by virtue of sequence complementarity to the miRNA seed region.
- Laser capture microdissection
-
(LCM). A method for capturing specific cells of interest from heterogeneous tissue samples. Cells for capture are chosen by the operator using a microscope and are cut out from the tissue using a laser. The isolated cells can be used for various analyses, including miRNA profiling.
- Splinted ligation
-
A method of labelling in which ligation of an oligonucleotide to the 3′ end of a microRNA is facilitated by a 'bridge' oligonucleotide that hybridizes to both the 3′ end of the miRNA and the 5 end of the oligonucleotide to be ligated.
- Microfluidic cards
-
Typically, disposable cards in which fluid pressure is used to move input samples and reagents through microfabricated channels into specific locations (akin to 'wells') with high precision, permitting highly parallel and low-sample-volume, real-time PCR.
- Locked nucleic acids
-
(LNAs). A class of RNA analogues in which the 2′ oxygen and the 4′ carbon positions in the ribose ring are connected or 'locked' to create increased thermal stability relative to DNA or RNA when they are complexed with complementary DNA or RNA.
- Next-generation sequencing
-
(NGS). Any of several technologies that sequence large numbers of DNA fragments in parallel, producing millions or billions of short reads in a single run of an automated sequencer.
- PIWI-interacting RNAs
-
(piRNAs). Small (23–32 nt) RNAs that are associated with PIWI clade proteins of the Argonaute family. They ensure genome stability in the germline of flies, mice and zebrafish by silencing transposable and repetitive elements.
Rights and permissions
About this article
Cite this article
Pritchard, C., Cheng, H. & Tewari, M. MicroRNA profiling: approaches and considerations. Nat Rev Genet 13, 358–369 (2012). https://doi.org/10.1038/nrg3198
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrg3198
This article is cited by
-
Understanding molecular mechanisms and miRNA-based targets in diabetes foot ulcers
Molecular Biology Reports (2024)
-
Emerging roles of plant microRNAs during Colletotrichum spp. infection
Planta (2024)
-
Correlation of PTEN signaling pathway and miRNA in breast cancer
Molecular Biology Reports (2024)
-
Non-coding RNA methylation modifications in hepatocellular carcinoma: interactions and potential implications
Cell Communication and Signaling (2023)
-
Exploring the potential of microRNA as a diagnostic tool for gestational diabetes
Journal of Translational Medicine (2023)