Key Points
-
The term metastasis encompasses specific biological traits that together enable the spread of aggressive tumour cells.
-
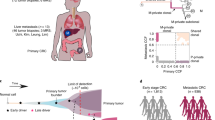
Different tumour types metastasize to distinct secondary organs, reflecting the influence of: the malignant cell of origin, aggressiveness of the primary tumour, the direction of circulation, and the capacity to co-opt supporting components of the microenvironment.
-
Underlying each of these factors are genetic determinants that are largely distinct from those that mediate malignant transformation.
-
Mediators of metastasis can be classified as metastasis initiation, metastasis progression and metastasis virulence genes, on the basis of the tumour stage, location in the body where they act, and their biological function.
-
By combining genome-wide technologies, functional experimentation in model systems and clinical validation, it is becoming possible to identify genetic alterations that are relevant to human metastatic disease.
-
These integrative approaches have uncovered genetic, epigenetic, somatic and inherited alterations, and serve as precedents for future metastasis gene discovery, prognosis and therapy.
Abstract
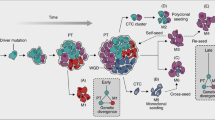
Metastasis can be viewed as an evolutionary process, culminating in the prevalence of rare tumour cells that overcame stringent physiological barriers as they separated from their original environment and developmental fate. This phenomenon brings into focus long-standing questions about the stage at which cancer cells acquire metastatic abilities, the relationship of metastatic cells to their tumour of origin, the basis for metastatic tissue tropism, the nature of metastasis predisposition factors and, importantly, the identity of genes that mediate these processes. With knowledge cemented in decades of research into tumour-initiating events, current experimental and conceptual models are beginning to address the genetic basis for cancer colonization of distant organs.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Weigelt, B., Peterse, J. L. & van 't Veer, L. J. Breast cancer metastasis: markers and models. Nature Rev. Cancer 5, 591–602 (2005).
van de Wouw, A. J., Jansen, R. L., Speel, E. J. & Hillen, H. F. The unknown biology of the unknown primary tumour: a literature review. Ann. Oncol. 14, 191–6 (2003).
Weiss, L. Metastasis of cancer: a conceptual history from antiquity to the 1990s. Cancer Metastasis Rev. 19, 193–383 (2000).
Norton, L. & Massagué, J. Is cancer a disease of self-seeding? Nature Med. 12, 875–878 (2006).
Hanahan, D. & Weinberg, R. A. The hallmarks of cancer. Cell 100, 57–70 (2000).
Christofori, G. New signals from the invasive front. Nature 441, 444–450 (2006).
Fidler, I. J. The pathogenesis of cancer metastasis: the 'seed and soil' hypothesis revisited. Nature Rev. Cancer 3, 453–458 (2003).
Gupta, G. P. & Massagué, J. Cancer metastasis: building a framework. Cell 127, 679–695 (2006).
Mundy, G. R. Metastasis to bone: causes, consequences and therapeutic opportunities. Nature Rev. Cancer 2, 584–593 (2002).
Steeg, P. S. Tumor metastasis: mechanistic insights and clinical challenges. Nature Med. 12, 895–904 (2006).
Capasso, L. L. Antiquity of cancer. Int. J. Cancer 113, 2–13 (2005).
Paget, S. The distribution of secondary growths in cancer of the breast. 1889. Cancer Metastasis Rev. 8, 98–101 (1989).
Ewing, J. Neoplastic Diseases edn 6 (Saunders, Philadelphia, 1928).
Fisher, B. & Fisher, E. R. The interrelationship of hematogenous and lymphatic tumor cell dissemination. Surg. Gynecol. Obstet. 122, 791–798 (1966).
Crespi, B. & Summers, K. Evolutionary biology of cancer. Trends Ecol. Evol. 20, 545–552 (2005).
Nowell, P. C. The clonal evolution of tumor cell populations. Science 194, 23–28 (1976).
Fidler, I. J. Selection of successive tumour lines for metastasis. Nature New Biol. 242, 148–149 (1973).
Fidler, I. J. & Kripke, M. L. Metastasis results from preexisting variant cells within a malignant tumor. Science 197, 893–895 (1977).
Fearon, E. R. & Vogelstein, B. A genetic model for colorectal tumorigenesis. Cell 61, 759–767 (1990).
Vogelstein, B. & Kinzler, K. W. Cancer genes and the pathways they control. Nature Med. 10, 789–799 (2004).
Bernards, R. & Weinberg, R. A. A progression puzzle. Nature 418, 823 (2002).
Alizadeh, A. A. et al. Distinct types of diffuse large B-cell lymphoma identified by gene expression profiling. Nature 403, 503–511 (2000).
Golub, T. R. et al. Molecular classification of cancer: class discovery and class prediction by gene expression monitoring. Science 286, 531–537 (1999).
Ramaswamy, S., Ross, K. N., Lander, E. S. & Golub, T. R. A molecular signature of metastasis in primary solid tumors. Nature Genet. 33, 49–54 (2003).
van de Vijver, M. J. et al. A gene-expression signature as a predictor of survival in breast cancer. N. Engl. J. Med. 347, 1999–2009 (2002).
van 't Veer, L. J. et al. Gene expression profiling predicts clinical outcome of breast cancer. Nature 415, 530–536 (2002).
Perou, C. M. et al. Molecular portraits of human breast tumours. Nature 406, 747–752 (2000).
Kerbel, R. S., Waghorne, C., Korczak, B., Lagarde, A. & Breitman, M. L. Clonal dominance of primary tumours by metastatic cells: genetic analysis and biological implications. Cancer Surv. 7, 597–629 (1988).
Steeg, P. S. Metastasis suppressors alter the signal transduction of cancer cells. Nature Rev. Cancer 3, 55–63 (2003).
Schmidt-Kittler, O. et al. From latent disseminated cells to overt metastasis: genetic analysis of systemic breast cancer progression. Proc. Natl Acad. Sci. USA 100, 7737–7742 (2003). A cytogenetic analysis of single tumour cells from the bone marrow of breast cancer patients, leading to the suggestion that metastatic cells disseminate early and evolve independently of their primary tumour.
Schardt, J. A. et al. Genomic analysis of single cytokeratin-positive cells from bone marrow reveals early mutational events in breast cancer. Cancer Cell 8, 227–239 (2005).
Slamon, D. J. et al. Use of chemotherapy plus a monoclonal antibody against HER2 for metastatic breast cancer that overexpresses HER2. N. Engl. J. Med. 344, 783–792 (2001).
Huber, M. A., Kraut, N. & Beug, H. Molecular requirements for epithelial–mesenchymal transition during tumor progression. Curr. Opin. Cell Biol. 17, 548–558 (2005).
Thiery, J. P. Epithelial–mesenchymal transitions in tumour progression. Nature Rev. Cancer 2, 442–454 (2002).
Stupack, D. G. et al. Potentiation of neuroblastoma metastasis by loss of caspase-8. Nature 439, 95–99 (2006).
Gupta, G. P. et al. Mediators of vascular remodelling co-opted for metastatic extravasation. Nature 446, 765–770 (2007).
Minn, A. J. et al. Genes that mediate breast cancer metastasis to lung. Nature 436, 518–524 (2005). References 36 and 37 integrate in vivo selection with clinical validation to identify mediators of lung-specific metastasis, linking aggressive primary tumorigenesis to organ-specific colonization.
Minn, A. J. et al. Lung metastasis genes couple breast tumor size and metastatic spread. Proc. Natl Acad. Sci. USA 104, 6740–6745 (2007).
Richards, F. M. et al. Germline E-cadherin gene (CDH1) mutations predispose to familial gastric cancer and colorectal cancer. Hum. Mol. Genet. 8, 607–610 (1999). This report links the inactivation of a developmentally regulated cell-adhesion gene with predisposition to cancer progression.
Cavallaro, U. & Christofori, G. Cell adhesion and signalling by cadherins and Ig-CAMs in cancer. Nature Rev. Cancer 4, 118–132 (2004).
Kapitanovic, S. et al. nm23-H1 expression and loss of heterozygosity in colon adenocarcinoma. J. Clin. Pathol. 57, 1312–1318 (2004).
Kim, M. et al. Comparative oncogenomics identifies NEDD9 as a melanoma metastasis gene. Cell 125, 1269–1281 (2006). An integrative approach that uses a mouse model to filter human aCGH data and characterize chromosomal aberrations that are associated with melanoma metastasis.
Thompson, E. W. & Newgreen, D. F. Carcinoma invasion and metastasis: a role for epithelial–mesenchymal transition? Cancer Res. 65, 5991–5995 (2005).
Tarin, D. The fallacy of epithelial mesenchymal transition in neoplasia. Cancer Res. 65, 5996–6000 (2005).
Kaelin, W. G. The von Hippel–Lindau tumor suppressor protein: roles in cancer and oxygen sensing. Cold Spring Harb. Symp. Quant. Biol. 70, 159–166 (2005).
Staller, P. et al. Chemokine receptor CXCR4 downregulated by von Hippel–Lindau tumour suppressor pVHL. Nature 425, 307–311 (2003). An example of how somatic mutations that are acquired during tumour progression can affect the expression of a metastasis-specific gene.
Kang, Y. et al. A multigenic program mediating breast cancer metastasis to bone. Cancer Cell 3, 537–549 (2003).
Muller, A. et al. Involvement of chemokine receptors in breast cancer metastasis. Nature 410, 50–56 (2001). This paper describes how non-immunological tumour cells can express the chemokine receptor CXCR4 and respond to a chemokine source to settle in certain organs.
Feinberg, A. P., Ohlsson, R. & Henikoff, S. The epigenetic progenitor origin of human cancer. Nature Rev. Genet. 7, 21–33 (2006).
Baylin, S. B. & Ohm, J. E. Epigenetic gene silencing in cancer — a mechanism for early oncogenic pathway addiction? Nature Rev. Cancer 6, 107–116 (2006).
Kleer, C. G. et al. EZH2 is a marker of aggressive breast cancer and promotes neoplastic transformation of breast epithelial cells. Proc. Natl Acad. Sci. USA 100, 11606–11611 (2003).
Varambally, S. et al. The polycomb group protein EZH2 is involved in progression of prostate cancer. Nature 419, 624–629 (2002). This paper reports the deregulation of stem-cell epigenetic regulators during metastatic progression.
Kim, J. H. et al. Transcriptional regulation of a metastasis suppressor gene by Tip60 and b-catenin complexes. Nature 434, 921–926 (2005). A study that links the progenitor WNT/β-catenin pathway to the transcriptional repression of a metastasis suppressor gene.
Bandyopadhyay, S. et al. Interaction of KAI1 on tumor cells with DARC on vascular endothelium leads to metastasis suppression. Nature Med. 12, 933–938 (2006).
Frank, S. A. Genetic predisposition to cancer — insights from population genetics. Nature Rev. Genet. 5, 764–772 (2004).
Pharoah, P. D. et al. Polygenic susceptibility to breast cancer and implications for prevention. Nature Genet. 31, 33–36 (2002).
Carey, L. A. et al. Race, breast cancer subtypes, and survival in the Carolina Breast Cancer Study. JAMA 295, 2492–2502 (2006).
Lifsted, T. et al. Identification of inbred mouse strains harboring genetic modifiers of mammary tumor age of onset and metastatic progression. Int. J. Cancer 77, 640–644 (1998).
Park, Y. G. et al. Comparative sequence analysis in eight inbred strains of the metastasis modifier QTL candidate gene Brms1. Mamm. Genome 13, 289–292 (2002).
Park, Y. G. et al. SIPA1 is a candidate for underlying the metastasis efficiency modifier locus MTES1. Nature Genet. 37, 1055–1062 (2005). The first experimental identification of a polymorphism that affects metastatic potential in mice.
Crawford, N. P. et al. Germline polymorphisms in SIPA1 are associated with metastasis and other indicators of poor prognosis in breast cancer. Breast Cancer Res. 8, R16 (2006).
Hiratsuka, S., Watanabe, A., Aburatani, H. & Maru, Y. Tumour-mediated upregulation of chemoattractants and recruitment of myeloid cells predetermines lung metastasis. Nature Cell Biol. 8, 1369–1375 (2006).
Gupta, P. B. et al. The melanocyte differentiation program predisposes to metastasis after neoplastic transformation. Nature Genet. 37, 1047–1054 (2005).
Yu, Y. et al. Expression profiling identifies the cytoskeletal organizer ezrin and the developmental homeoprotein SIX-1 as key metastatic regulators. Nature Med. 10, 175–181 (2004).
Yang, J. et al. Twist, a master regulator of morphogenesis, plays an essential role in tumor metastasis. Cell 117, 927–939 (2004). A gene-expression analysis that identified a role for a transcriptional regulator of embryo development during mouse mammary tumour intravasation.
de Visser, K. E., Eichten, A. & Coussens, L. M. Paradoxical roles of the immune system during cancer development. Nature Rev. Cancer 6, 24–37 (2006).
Karin, M. Nuclear factor-kB in cancer development and progression. Nature 441, 431–436 (2006).
Park, B. K. et al. NF-kB in breast cancer cells promotes osteolytic bone metastasis by inducing osteoclastogenesis via GM-CSF. Nature Med. 13, 62–69 (2007).
Luo, J. L. et al. Nuclear cytokine-activated IKKα controls prostate cancer metastasis by repressing Maspin. Nature 18 March 2007 (doi:10.1038/nature05656).
Kang, Y. et al. Breast cancer bone metastasis mediated by the Smad tumor suppressor pathway. Proc. Natl Acad. Sci. USA 102, 13909–13914 (2005).
Olsson, A. K., Dimberg, A., Kreuger, J. & Claesson-Welsh, L. VEGF receptor signalling — in control of vascular function. Nature Rev. Mol. Cell Biol. 7, 359–371 (2006).
Weis, S., Cui, J., Barnes, L. & Cheresh, D. Endothelial barrier disruption by VEGF-mediated Src activity potentiates tumor cell extravasation and metastasis. J. Cell Biol. 167, 223–229 (2004).
Kaplan, R. N. et al. VEGFR1-positive haematopoietic bone marrow progenitors initiate the pre-metastatic niche. Nature 438, 820–827 (2005).
Bierie, B. & Moses, H. L. Tumour microenvironment: TGFβ: the molecular Jekyll and Hyde of cancer. Nature Rev. Cancer 6, 506–520 (2006).
Siegel, P. M. & Massagué, J. Cytostatic and apoptotic actions of TGFβ in homeostasis and cancer. Nature Rev. Cancer 3, 807–821 (2003).
Carmeliet, P. Angiogenesis in life, disease and medicine. Nature 438, 932–936 (2005).
Carmeliet, P. & Jain, R. K. Angiogenesis in cancer and other diseases. Nature 407, 249–257 (2000).
Debnath, J. & Brugge, J. S. Modelling glandular epithelial cancers in three-dimensional cultures. Nature Rev. Cancer 5, 675–688 (2005).
Van Dyke, T. & Jacks, T. Cancer modeling in the modern era: progress and challenges. Cell 108, 135–144 (2002).
Guy, C. T., Cardiff, R. D. & Muller, W. J. Induction of mammary tumors by expression of polyomavirus middle T oncogene: a transgenic mouse model for metastatic disease. Mol. Cell. Biol. 12, 954–961 (1992).
Gingrich, J. R. et al. Metastatic prostate cancer in a transgenic mouse. Cancer Res. 56, 4096–4102 (1996).
Jackson, E. L. et al. The differential effects of mutant p53 alleles on advanced murine lung cancer. Cancer Res. 65, 10280–10288 (2005).
Kim, C. F. et al. Mouse models of human non-small-cell lung cancer: raising the bar. Cold Spring Harb. Symp. Quant. Biol. 70, 241–250 (2005).
Nathoo, N., Toms, S. A. & Barnett, G. H. Metastases to the brain: current management perspectives. Expert Rev. Neurother. 4, 633–640 (2004).
Douma, S. et al. Suppression of anoikis and induction of metastasis by the neurotrophic receptor TrkB. Nature 430, 1034–1039 (2004).
Brumby, A. M. & Richardson, H. E. Using Drosophila melanogaster to map human cancer pathways. Nature Rev. Cancer 5, 626–639 (2005).
Woodhouse, E. C. et al. Drosophila screening model for metastasis: Semaphorin 5c is required for l(2)gl cancer phenotype. Proc. Natl Acad. Sci. USA 100, 11463–11468 (2003).
Pagliarini, R. A. & Xu, T. A genetic screen in Drosophila for metastatic behavior. Science 302, 1227–1231 (2003).
Dupuy, A. J., Akagi, K., Largaespada, D. A., Copeland, N. G. & Jenkins, N. A. Mammalian mutagenesis using a highly mobile somatic Sleeping Beauty transposon system. Nature 436, 221–226 (2005).
Collier, L. S., Carlson, C. M., Ravimohan, S., Dupuy, A. J. & Largaespada, D. A. Cancer gene discovery in solid tumours using transposon-based somatic mutagenesis in the mouse. Nature 436, 272–276 (2005).
Dickins, R. A. et al. Probing tumor phenotypes using stable and regulated synthetic microRNA precursors. Nature Genet. 37, 1289–1295 (2005).
Ellsworth, R. E. et al. Allelic imbalance in primary breast carcinomas and metastatic tumors of the axillary lymph nodes. Mol. Cancer Res. 3, 71–77 (2005).
Sebat, J. et al. Large-scale copy number polymorphism in the human genome. Science 305, 525–528 (2004).
Laird, P. W. The power and the promise of DNA methylation markers. Nature Rev. Cancer 3, 253–266 (2003).
Lu, J. et al. MicroRNA expression profiles classify human cancers. Nature 435, 834–838 (2005).
Varambally, S. et al. Integrative genomic and proteomic analysis of prostate cancer reveals signatures of metastatic progression. Cancer Cell 8, 393–406 (2005).
Sjoblom, T. et al. The consensus coding sequences of human breast and colorectal cancers. Science 314, 268–274 (2006).
Wang, Y. et al. Gene-expression profiles to predict distant metastasis of lymph-node-negative primary breast cancer. Lancet 365, 671–679 (2005).
Fan, C. et al. Concordance among gene-expression-based predictors for breast cancer. N. Engl. J. Med. 355, 560–569 (2006). An analysis of five prominent breast cancer gene signatures that shows that, despite little overlap in gene identity, these signatures can classify similar subsets of patient who are at risk for metastatic relapse.
Massagué, J. Sorting out breast-cancer gene signatures. N. Engl. J. Med. 356, 294–297 (2007).
Chang, H. Y. et al. Robustness, scalability, and integration of a wound-response gene expression signature in predicting breast cancer survival. Proc. Natl Acad. Sci. USA 102, 3738–3743 (2005).
Liu, R. et al. The prognostic role of a gene signature from tumorigenic breast-cancer cells. N. Engl. J. Med. 356, 217–226 (2007).
Rhodes, D. R. & Chinnaiyan, A. M. Integrative analysis of the cancer transcriptome. Nature Genet. 37, S31–S37 (2005).
Luzzi, K. J. et al. Multistep nature of metastatic inefficiency: dormancy of solitary cells after successful extravasation and limited survival of early micrometastases. Am. J. Pathol. 153, 865–873 (1998).
Fidler, I. J. & Nicolson, G. L. Fate of recirculating B16 melanoma metastatic variant cells in parabiotic syngeneic recipients. J. Natl Cancer Inst. 58, 1867–1872 (1977).
Khanna, C. & Hunter, K. Modeling metastasis in vivo. Carcinogenesis 26, 513–523 (2005).
Clark, E. A., Golub, T. R., Lander, E. S. & Hynes, R. O. Genomic analysis of metastasis reveals an essential role for RhoC. Nature 406, 532–535 (2000). The first study to combine genomic profiling and in vivo selection for the identification of metastasis genes.
Khanna, C. et al. The membrane–cytoskeleton linker ezrin is necessary for osteosarcoma metastasis. Nature Med. 10, 182–186 (2004).
Minn, A. J. et al. Distinct organ-specific metastatic potential of individual breast cancer cells and primary tumors. J. Clin. Invest. 115, 44–55 (2005).
Pinkel, D. & Albertson, D. G. Comparative genomic hybridization. Annu. Rev. Genomics Hum. Genet. 6, 331–354 (2005).
Adler, A. S. et al. Genetic regulators of large-scale transcriptional signatures in cancer. Nature Genet. 38, 421–430 (2006).
Chin, K. et al. Genomic and transcriptional aberrations linked to breast cancer pathophysiologies. Cancer Cell 10, 529–541 (2006).
Zender, L. et al. Identification and validation of oncogenes in liver cancer using an integrative oncogenomic approach. Cell 125, 1253–1267 (2006).
Sweet-Cordero, A. et al. An oncogenic KRAS2 expression signature identified by cross-species gene-expression analysis. Nature Genet. 37, 48–55 (2005).
Ellwood-Yen, K. et al. Myc-driven murine prostate cancer shares molecular features with human prostate tumors. Cancer Cell 4, 223–238 (2003).
Tarin, D., Vass, A. C., Kettlewell, M. G. & Price, J. E. Absence of metastatic sequelae during long-term treatment of malignant ascites by peritoneo-venous shunting. A clinico-pathological report. Invasion Metastasis 4, 1–12 (1984).
Joyce, J. A. Therapeutic targeting of the tumor microenvironment. Cancer Cell 7, 513–520 (2005).
Nierodzik, M. L. & Karpatkin, S. Thrombin induces tumor growth, metastasis, and angiogenesis: evidence for a thrombin-regulated dormant tumor phenotype. Cancer Cell 10, 355–362 (2006).
Sorlie, T. et al. Gene expression patterns of breast carcinomas distinguish tumor subclasses with clinical implications. Proc. Natl Acad. Sci. USA 98, 10869–10874 (2001).
Paik, S. et al. A multigene assay to predict recurrence of tamoxifen-treated, node-negative breast cancer. N. Engl. J. Med. 351, 2817–2826 (2004).
Chang, H. Y. et al. Gene expression signature of fibroblast serum response predicts human cancer progression: similarities between tumors and wounds. PLoS Biol. 2, e7 (2004).
Chi, J. T. et al. Gene expression programs in response to hypoxia: cell type specificity and prognostic significance in human cancers. PLoS Med. 3, e47 (2006).
Bild, A. H. et al. Oncogenic pathway signatures in human cancers as a guide to targeted therapies. Nature 439, 353–357 (2006).
Acknowledgements
We apologize for omitting primary references and the work of those we could not cite owing to space limitations. We would like to thank A. Incassati, A. Chiang, P. Bos, D. Padua, G. Gupta and S.Tavazoie for insightful discussions. J.M. was funded by the National Institutes of Health, USA, grant P01-94060, and by a grant of the Keck Foundation, USA. D.X.N. is a Berlex postdoctoral fellow of the Damon Runyon Cancer Research Foundation, USA. J.M. is an Investigator of the Howard Hughes Medical Institute, USA.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Related links
Glossary
- Primary tumour
-
Cancer that arises from the malignant conversion of cells from an initial organ site.
- Histological grade
-
Morphologically identifiable steps of tumour progression that are used to classify disease stage.
- Organ tropism
-
A predilection of a primary tumour to spread to specific secondary organs.
- Angiogenesis
-
The pathological growth of new blood vessels to support tumour growth.
- Intravasation
-
Entry of tumour cells into the bloodstream.
- Extravasation
-
Exit of tumour cells out of capillary beds into the parenchyma of an organ.
- Parenchyma
-
The main functional portion of an organ.
- Emboli
-
Clumped tumour cells that typically lodge in blood vessels.
- Haematogenous dissemination
-
The spread of cancer cells through the bloodstream.
- Melanoma
-
Skin cancer that is initiated by the transformation of melanocytes.
- Metastasis suppressor gene
-
A gene in which loss of function specifically enhances metastasis without affecting primary tumour growth.
- Mesenteric circulation
-
Blood flow from the intestines.
- Peritoneal cavity
-
The space within the abdomen that contains the intestines, the stomach and the liver.
- Palliative shunt
-
Diversion of malignant fluid to alleviate symptoms that arise from its accumulation in the body cavity.
- Ascites
-
A detectable accumulation of free fluid in the peritoneal cavity.
- Metastasis initiation genes
-
A gene that is engaged in the invasion and intravasation of metastatic cells.
- Metastasis progression gene
-
A gene that has dual functions in mediating primary tumorigenesis and metastatic colonization.
- Metastasis virulence gene
-
A gene that is exclusively involved in distant organ colonization.
- Focal contacts
-
Dynamic cell-adhesion structures.
- Chemokine
-
A group of small signalling proteins (cytokines) that are usually secreted by immune cells.
- Neural crest cell
-
Highly motile cells that originate from the ectoderm during development.
- Rhabdomyosarcoma
-
Paediatric malignancy that arises from skeletal muscle cells.
- Stromal activation
-
Stimulation and mobilization of host cells in the microenvironment that surrounds a tumour.
- Osteoclastogenesis
-
The differentiation and activation of osteoclasts that mediates bone resorption.
- Xenograft
-
Implantation of human tumour cells into an immunocompromized animal.
- Orthotopic site
-
Transplant of tumour cells into the anatomical location of an animal that best recapitulates the original source of primary tumorigenesis.
Rights and permissions
About this article
Cite this article
Nguyen, D., Massagué, J. Genetic determinants of cancer metastasis. Nat Rev Genet 8, 341–352 (2007). https://doi.org/10.1038/nrg2101
Issue Date:
DOI: https://doi.org/10.1038/nrg2101
This article is cited by
-
miRNA-383-5p Regulated Migration and Invasion of Tumor Cells by Inhibiting NCKAP1 Expression in Gastric Cancer
Biochemical Genetics (2024)
-
New insights into the regulation of METTL3 and its role in tumors
Cell Communication and Signaling (2023)
-
Multidimensional outlook on the pathophysiology of cervical cancer invasion and metastasis
Molecular and Cellular Biochemistry (2023)
-
MicroRNA profiles in aqueous humor between pseudoexfoliation glaucoma and normal tension glaucoma patients in a Korean population
Scientific Reports (2022)
-
High expression of oncogene cadherin-6 correlates with tumor progression and a poor prognosis in gastric cancer
Cancer Cell International (2021)