Key Points
-
It is now well established that mutations within cis-regulatory regions are responsible for various significant evolutionary differences in organismal traits.
-
It has been proposed that cis-regulatory mutations make a qualitatively distinct contribution to trait evolution, in part because they might result in more limited pleiotropy and therefore fewer functional trade-offs, and in part because they might commonly be co-dominant and therefore immediately visible to selection.
-
Many species-specific differences in cuticular colouration in fruitflies are the product of mutations in the cis-regulatory regions of yellow (y), ebony (e) and bric a brac 1/2 (bab1/bab2).
-
The loss of pelvic armour in stickleback fish has evolved repeatedly through cis-regulatory mutations in paired-like homeodomain transcription factor 1 (pitx1).
-
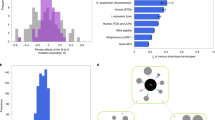
Many traits that are unique to humans or that vary among them are also due to mutations within cis-reguatory regions, including functionally significant mutations near Duffy blood group, chemokine receptor (DARC), lactase (LCT) and prodynorphin (PDYN).
-
Taken collectively, the evidence from the best-studied cases supports the idea that cis-regulatory mutations make a qualitatively distinctive contribution to trait evolution.
Abstract
For decades, evolutionary biologists have argued that changes in cis-regulatory sequences constitute an important part of the genetic basis for adaptation. Although originally based on first principles, this claim is now empirically well supported: numerous studies have identified cis-regulatory mutations with functionally significant consequences for morphology, physiology and behaviour. The focus has now shifted to considering whether cis-regulatory and coding mutations make qualitatively different contributions to phenotypic evolution. Cases in which parallel mutations have produced parallel trait modifications in particular suggest that some phenotypic changes are more likely to result from cis-regulatory mutations than from coding mutations.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Jacob, F. & Monod, J. Genetic regulatory mechanisms in the synthesis of proteins. J. Mol. Biol. 3, 318–356 (1961).
Monod, J. & Jacob, F. General conclusions — teleonomic mechanisms in cellular metabolism, growth, and differentiation. Cold Spring Harb. Symp. Quant. Biol. 26, 389–401 (1961).
Britten, R. J. & Davidson, E. H. Repetitive and non-repetitive DNA sequences and a speculation on the origins of evolutionary novelty. Q. Rev. Biol. 46, 111–138 (1971).
King, M. C. & Wilson, A. C. Evolution at two levels in humans and chimpanzees. Science 188, 107–116 (1975). This classic paper proposed that phenotypic differences between humans and chimpanzees are largely due to cis -regulatory mutations.
Carroll, S. B., Grenier, J. & Weatherbee, S. From DNA to Diversity (Blackwell Publishing, Malden, 2004).
Stern, D. L. Evolutionary developmental biology and the problem of variation. Evolution 54, 1079–1091 (2000).
Wilkins, A. S. The Evolution of Developmental Pathways (Sinauer Associates, Sunderland, 2002).
Wray, G. A. et al. The evolution of transcriptional regulation in eukaryotes. Mol. Biol. Evol. 20, 1377–1419 (2003).
Davidson, E. H. The Regulatory Genome: Gene Regulatory Networks in Development and Evolution (Academic, Burlington, 2006).
Gerhart, J. & Kirschner, M. Cells, Embryos, and Evolution: Toward a Cellular and Developmental Understanding of Phenotypic Variation and Evolutionary Adaptability (Blackwell Science, Malden, 1997).
Ruvkun, G., Wightman, B., Burglin, T. & Arasu, P. Dominant gain-of-function mutations that lead to misregulation of the C. elegans heterochronic gene lin-14, and the evolutionary implications of dominant mutations in pattern-formation genes. Dev. Suppl. 1, 47–54 (1991).
Pastinen, T. et al. A survey of genetic and epigenetic variation affecting human gene expression. Physiol. Genomics 16, 184–193 (2003).
Ronald, J., Brem, R. B., Whittle, J. & Kruglyak, L. Local regulatory variation in Saccharomyces cerevisiae. PLoS Genet. 1, e25 (2005).
Wittkopp, P. J., Haerum, B. K. & Clark, A. G. Evolutionary changes in cis and trans gene regulation. Nature 430, 85–88 (2004).
Li, W. H. Molecular Evolution (Sinauer Associates, Sunderland, 1997). References 13–15 measured the relative contributions of mutations in cis and trans to differences in transcription.
Force, A. et al. Preservation of duplicate genes by complementary, degenerative mutations. Genetics 151, 1531–1545 (1999).
Gompel, N. & Carroll, S. B. Genetic mechanisms and constraints governing the evolution of correlated traits in drosophilid flies. Nature 424, 931–935 (2003).
Hollocher, H., Hatcher, J. L. & Dyreson, E. G. Genetic and developmental analysis of abdominal pigmentation differences across species in the Drosophila dunni subgroup. Evolution 54, 2057–2071 (2000).
Kopp, A., Duncan, I., Godt, D. & Carroll, S. B. Genetic control and evolution of sexually dimorphic characters in Drosophila. Nature 408, 553–559 (2000).
Kopp, A. & True, J. R. Evolution of male sexual characters in the Oriental Drosophila melanogaster species group. Evol. Dev. 4, 278–291 (2002).
Hovemann, B. Tissue specific expression of the ebony gene. J. Neurogenet. 7, 128–128 (1991).
Hovemann, B. T. et al. The Drosophila ebony gene is closely related to microbial peptide synthetases and shows specific cuticle and nervous system expression. Gene 221, 1–9 (1998).
True, J. R. et al. Drosophila tan encodes a novel hydrolase required in pigmentation and vision. PLoS Genet. 1, e63 (2005).
Walter, M. F. et al. Temporal and spatial expression of the yellow gene in correlation with cuticle formation and dopa decarboxylase activity in Drosophila development. Dev. Biol. 147, 32–45 (1991).
Wittkopp, P. J., True, J. R. & Carroll, S. B. Reciprocal functions of the Drosophila yellow and ebony proteins in the development and evolution of pigment patterns. Development 129, 1849–1858 (2002).
Gompel, N., Prud'homme, B., Wittkopp, P. J., Kassner, V. A. & Carroll, S. B. Chance caught on the wing: cis-regulatory evolution and the origin of pigment patterns in Drosophila. Nature 433, 481–487 (2005).
Jeong, S., Rokas, A. & Carroll, S. B. Regulation of body pigmentation by the Abdominal-B Hox protein and its gain and loss in Drosophila evolution. Cell 125, 1387–1399 (2006). This study illustrates the power of using a combination of detailed molecular genetic analyses with an in-depth sampling of related species.
Prud'homme, B. et al. Repeated morphological evolution through cis-regulatory changes in a pleiotropic gene. Nature 440, 1050–1053 (2006). References 26 and 28 illustrate a remarkable case of similar genetic bases for a parallel change in wing pigmentation in flies.
Wittkopp, P. J., Vaccaro, K. & Carroll, S. B. Evolution of yellow gene regulation and pigmentation in Drosophila. Curr. Biol. 12, 1547–1556 (2002).
Beverley, S. M. & Wilson, A. C. Molecular evolution in Drosophila and the higher Diptera II. A time scale for fly evolution. J. Mol. Evol. 21, 1–13 (1984).
Vieira, J., Vieira, C. P., Hartl, D. L. & Lozovskaya, E. R. A framework physical map of Drosophila virilis based on P1 clones: applications in genome evolution. Chromosoma 106, 99–107 (1997).
Wittkopp, P. J., Williams, B. L., Selegue, J. E. & Carroll, S. B. Drosophila pigmentation evolution: divergent genotypes underlying convergent phenotypes. Proc. Natl Acad. Sci. USA 100, 1808–1813 (2003). This study demonstrated that parallel morphological changes do not always have a similar underlying genetic basis.
Simpson, P., Woehl, R. & Usui, K. The development and evolution of bristle patterns in Diptera. Development 126, 1349–1364 (1999).
Sucena, E. & Stern, D. L. Divergence of larval morphology between Drosophila sechellia and its sibling species caused by cis-regulatory evolution of ovo/shaven-baby. Proc. Natl Acad. Sci. USA 97, 4530–4534 (2000).
Skaer, N. & Simpson, P. Genetic analysis of bristle loss in hybrids between Drosophila melanogaster and D. simulans provides evidence for divergence of cis-regulatory sequences in the achaete–scute gene complex. Dev. Biol. 221, 148–167 (2000).
Sucena, E., Delon, I., Jones, I., Payre, F. & Stern, D. L. Regulatory evolution of shavenbaby/ovo underlies multiple cases of morphological parallelism. Nature 424, 935–938 (2003).
Peichel, C. L. et al. The genetic architecture of divergence between threespine stickleback species. Nature 414, 901–905 (2001).
Bell, M. A. & Foster, S. A. (eds) The Evolutionary Biology Of The Threespine Stickleback (Oxford Univ. Press, New York, 1994).
Cresko, W. A. et al. Parallel genetic basis for repeated evolution of armor loss in Alaskan threespine stickleback populations. Proc. Natl Acad. Sci. USA 101, 6050–6055 (2004).
Shapiro, M. D. et al. Genetic and developmental basis of evolutionary pelvic reduction in threespine sticklebacks. Nature 428, 717–723 (2004). This study used a combination of genetic mapping and expression analyses to demonstrate that cis -regulatory mutation(s) constitute the primary genetic basis for loss of pelvic armour in sticklebacks.
Shang, J., Luo, Y. & Clayton, D. A. Backfoot is a novel homeobox gene expressed in the mesenchyme of developing hind limb. Dev. Dyn. 209, 242–253 (1997).
Marcil, A., Dumontier, E., Chamberland, M., Camper, S. A. & Drouin, J. Pitx1 and Pitx2 are required for development of hindlimb buds. Development 130, 45–55 (2003).
McLennan, D. A. & Mattern, M. Y. The phylogeny of the Gasterosteidae: combining behavioral and morphological data sets. Cladistics 17, 11–27 (2001).
Bell, M. A., Baumgartner, J. V. & Olson, E. C. Patterns of temporal change in single morphological characters of a miocene stickleback fish. Paleobiology 11, 258–271 (1985).
Shapiro, M. D., Bell, M. A. & Kingsley, D. M. Parallel origins of pelvic reduction in vertebrates. Proc. Natl Acad. Sci. USA 103, 3753–3758 (2006).
Tanaka, M. et al. Developmental genetic basis for the evolution of pelvic fin loss in the pufferfish Takifugu rubripes. Dev. Biol. 281, 227–239 (2005).
Carroll, S. B. Genetics and the making of Homo sapiens. Nature 422, 849–857 (2003).
Vallender, E. J. & Lahn, B. T. Positive selection on the human genome. Hum. Mol. Genet. 13, R245–R254 (2004).
Chaudhuri, A. et al. Expression of the Duffy antigen in K562 cells. Evidence that it is the human erythrocyte chemokine receptor. J. Biol. Chem. 269, 7835–7838 (1994).
Horuk, R. et al. A receptor for the malarial parasite Plasmodium vivax: the erythrocyte chemokine receptor. Science 261, 1182–1184 (1993).
Tournamille, C. et al. Sequence, evolution and ligand binding properties of mammalian Duffy antigen/receptor for chemokines. Immunogenetics 55, 682–694 (2004).
Pogo, A. O. & Chaudhuri, A. The Duffy protein: a malarial and chemokine receptor. Semin. Hematol. 37, 122–129 (2000).
Hadley, T. J. & Peiper, S. C. From malaria to chemokine receptor: the emerging physiologic role of the Duffy blood group antigen. Blood 89, 3077–3091 (1997).
Miller, L. H., Mason, S. J., Clyde, D. F. & McGinniss, M. H. The resistance factor to Plasmodium vivax in blacks. The Duffy-blood-group genotype, FyFy. N. Engl. J. Med. 295, 302–304 (1976).
Chaudhuri, A., Polyakova, J., Zbrzezna, V. & Pogo, A. O. The coding sequence of Duffy blood group gene in humans and simians: restriction fragment length polymorphism, antibody and malarial parasite specificities, and expression in nonerythroid tissues in Duffy-negative individuals. Blood 85, 615–621 (1995).
Peiper, S. C. et al. The Duffy antigen/receptor for chemokines (DARC) is expressed in endothelial cells of Duffy negative individuals who lack the erythrocyte receptor. J. Exp. Med. 181, 1311–1317 (1995).
Iwamoto, S., Li, J., Sugimoto, N., Okuda, H. & Kajii, E. Characterization of the Duffy gene promoter: evidence for tissue-specific abolishment of expression in Fy(a-b-) of black individuals. Biochem. Biophys. Res. Commun. 222, 852–859 (1996).
Tournamille, C., Colin, Y., Cartron, J. P. & Le Van Kim, C. Disruption of a GATA motif in the Duffy gene promoter abolishes erythroid gene expression in Duffy-negative individuals. Nature Genet. 10, 224–228 (1995).
Hamblin, M. T. & Di Rienzo, A. Detection of the signature of natural selection in humans: evidence from the Duffy blood group locus. Am. J. Hum. Genet. 66, 1669–1679 (2000). References 58 and 59 provided evidence regarding the molecular and evolutionary mechanisms, respectively, for adaptive evolution of a regulatory SNP that provides protection from malarial infection.
Hamblin, M. T., Thompson, E. E. & Di Rienzo, A. Complex signatures of natural selection at the Duffy blood group locus. Am. J. Hum. Genet. 70, 369–383 (2002).
Swallow, D. M. Genetics of lactase persistence and lactose intolerance. Annu. Rev. Genet. 37, 197–219 (2003).
Bersaglieri, T. et al. Genetic signatures of strong recent positive selection at the lactase gene. Am. J. Hum. Genet. 74, 1111–1120 (2004).
Olds, L. C. & Sibley, E. Lactase persistence DNA variant enhances lactase promoter activity in vitro: functional role as a cis regulatory element. Hum. Mol. Genet. 12, 2333–2340 (2003).
Tishkoff, S. A. et al. Convergent adaptation of human lactase persistence in Africa and Europe. Nature Genet. 39, 31–40 (2007). This study demonstrated a nearly parallel genetic basis for an important physiological adaptation during recent human evolution.
Fisher, S. E. & Marcus, G. F. The eloquent ape: genes, brains and the evolution of language. Nature Rev. Genet. 7, 9–20 (2006).
Sikela, J. M. The jewels of our genome: the search for the genomic changes underlying the evolutionarily unique capacities of the human brain. PLoS Genet. 2, e80 (2006).
Enard, W. et al. Intra- and interspecific variation in primate gene expression patterns. Science 296, 340–343 (2002). The authors demonstrated an elevated rate of change in the expression of genes within the brain on the branch leading to humans relative to that leading to chimpanzees.
Caceres, M. et al. Elevated gene expression levels distinguish human from non-human primate brains. Proc. Natl Acad. Sci. USA 100, 13030–13035 (2003).
Khaitovich, P. et al. Parallel patterns of evolution in the genomes and transcriptomes of humans and chimpanzees. Science 309, 1850–1854 (2005).
Khaitovich, P. et al. Regional patterns of gene expression in human and chimpanzee brains. Genome Res. 14, 1462–1473 (2004).
Rockman, M. V. et al. Ancient and recent positive selection transformed opioid cis-regulation in humans. PLoS Biol. 3, e387 (2005). This study found evidence of positive selection on specific cis -regulatory mutations at a locus encoding a neuropeptide with roles in human cognition.
Rodgers, R. J. & Cooper, S. J. (eds) Endorphins, Opiates, and Behavioural Processes (Wiley, New York, 1988).
Wagner, J. J., Terman, G. W. & Chavkin, C. Endogenous dynorphins inhibit excitatory neurotransmission and block LTP induction in the hippocampus. Nature 363, 451–454 (1993).
Hurd, Y. L. Subjects with major depression or bipolar disorder show reduction of prodynorphin mRNA expression in discrete nuclei of the amygdaloid complex. Mol. Psychiatry 7, 75–81 (2002).
Peckys, D. & Hurd, Y. L. Prodynorphin and k opioid receptor mRNA expression in the cingulate and prefrontal cortices of subjects diagnosed with schizophrenia or affective disorders. Brain Res. Bull. 55, 619–624 (2001).
Stogmann, E. et al. A functional polymorphisms in the prodynorphin gene promoter is associated with temporal lobe epilepsy. Ann. Neurol. 51, 260–263 (2002).
Ventriglia, M. et al. Allelic variation in the human prodynorphin gene promoter and schizophrenia. Neuropsychobiology 46, 17–21 (2002).
Nachman, M. W., Hoekstra, H. E. & D'Agostino, S. L. The genetic basis of adaptive melanism in pocket mice. Proc. Natl Acad. Sci. USA 100, 5268–5273 (2003).
Theron, E., Hawkins, K., Bermingham, E., Ricklefs, R. E. & Mundy, N. I. The molecular basis of an avian plumage polymorphism in the wild: a melanocortin-1-receptor point mutation is perfectly associated with the melanic plumage morph of the bananaquit, Coereba flaveola. Curr. Biol. 11, 550–557 (2001).
Kopp, A. Basal relationships in the Drosophila melanogaster species group. Mol. Phylogenet. Evol. 39, 787–798 (2006).
Enattah, N. S. et al. Identification of a variant associated with adult-type hypolactasia. Nature Genet. 30, 233–237 (2002).
Bachner-Melman, R. et al. AVPR1a and SLC6A4 gene polymorphisms are associated with creative dance performance. PLoS Genet. 1, e42 (2005).
Hammock, E. A. & Young, L. J. Microsatellite instability generates diversity in brain and sociobehavioral traits. Science 308, 1630–1634 (2005).
Daborn, P. J. et al. A single p450 allele associated with insecticide resistance in Drosophila. Science 297, 2253–2256 (2002).
Lerman, D. N. & Feder, M. E. Naturally occurring transposable elements disrupt hsp70 promoter function in Drosophila melanogaster. Mol. Biol. Evol. 22, 776–783 (2005).
Lerman, D. N., Michalak, P., Helin, A. B., Bettencourt, B. R. & Feder, M. E. Modification of heat-shock gene expression in Drosophila melanogaster populations via transposable elements. Mol. Biol. Evol. 20, 135–144 (2003).
Enoch, M. A. et al. 5-HT2A promoter polymorphism –1438G/A, anorexia nervosa, and obsessive-compulsive disorder. Lancet 351, 1785–1786 (1998).
Moraes, M. O. et al. Interleukin-10 promoter single-nucleotide polymorphisms as markers for disease susceptibility and disease severity in leprosy. Genes Immun. 5, 592–595 (2004).
Shin, H. D. et al. Genetic restriction of HIV-1 pathogenesis to AIDS by promoter alleles of IL10. Proc. Natl Acad. Sci. USA 97, 14467–14472 (2000).
He, G. et al. Interleukin-10 –1082 promoter polymorphism is associated with schizophrenia in a Han Chinese sib-pair study. Neurosci. Lett. 394, 1–4 (2006).
Crawford, D. L., Segal, J. A. & Barnett, J. L. Evolutionary analysis of TATA-less proximal promoter function. Mol. Biol. Evol. 16, 194–207 (1999).
Caspi, A. et al. Role of genotype in the cycle of violence in maltreated children. Science 297, 851–854 (2002).
Kim-Cohen, J. et al. MAOA, maltreatment, and gene–environment interaction predicting children's mental health: new evidence and a meta-analysis. Mol. Psychiatry 11, 903–913 (2006).
Beyzade, S. et al. Influences of matrix metalloproteinase-3 gene variation on extent of coronary atherosclerosis and risk of myocardial infarction. J. Am. Coll. Cardiol. 41, 2130–2137 (2003).
Ye, S. et al. Progression of coronary atherosclerosis is associated with a common genetic variant of the human stromelysin-1 promoter which results in reduced gene expression. J. Biol. Chem. 271, 13055–13060 (1996).
Marcellini, S. & Simpson, P. Two or four bristles: functional evolution of an enhancer of scute in drosophilidae. PLoS Biol. 4, e386 (2006).
Hariri, A. R. et al. Serotonin transporter genetic variation and the response of the human amygdala. Science 297, 400–403 (2002).
Trefilov, A., Berard, J., Krawczak, M. & Schmidtke, J. Natal dispersal in rhesus macaques is related to serotonin transporter gene promoter variation. Behav. Genet. 30, 295–301 (2000).
Clark, R. M., Wagler, T. N., Quijada, P. & Doebley, J. A distant upstream enhancer at the maize domestication gene tb1 has pleiotropic effects on plant and inflorescent architecture. Nature Genet. 38, 594–597 (2006).
Wang, R. L., Stec, A., Hey, J., Lukens, L. & Doebley, J. The limits of selection during maize domestication. Nature 398, 236–239 (1999).
Stern, D. L. A role of Ultrabithorax in morphological differences between Drosophila species. Nature 396, 463–466 (1998).
Drapeau, M. D., Cyran, S. A., Viering, M. M., Geyer, P. K. & Long, A. D. A cis-regulatory sequence within the yellow locus of Drosophila melanogaster required for normal male mating success. Genetics 172, 1009–1030 (2006).
Knight, J. C. Regulatory polymorphisms underlying complex disease traits. J. Mol. Med. 83, 97–109 (2005).
Rockman, M. V. & Wray, G. A. Abundant raw material for cis-regulatory evolution in humans. Mol. Biol. Evol. 19, 1991–2004 (2002).
Acknowledgements
Thanks to C. Babbitt, D. Garfield, J. Tung, and three anonymous reviewers for helpful comments. G.A.W.'s research is supported by the US National Science Foundation and the Institute for Genome Sciences & Policy at Duke University.
Author information
Authors and Affiliations
Ethics declarations
Competing interests
The author declares no competing financial interests.
Related links
Glossary
- Cis-regulatory region
-
A segment of DNA that regulates transcription; such segments typically lie immediately 5′ of the start site of transcription, but are often discontinuous, and individual segments can reside within introns, 5′ and 3′ UTRs, or tens of kilobases on either side of the gene they regulate.
- Clade
-
A group of species that share a unique common ancestor.
- Co-dominant
-
A mutation that has an additive phenotypic impact, and is therefore apparent in heterozygotes.
- Pleiotropy
-
The ability of a gene or mutation to alter more than one trait.
- Functional trade-off
-
For many traits, improving one aspect of function might incur a cost in some other aspect of function.
- Crypsis
-
Concealment from predators, usually through shape and colouration of the integument.
- Candidate gene
-
A gene that seems likely, on the basis of its function or a prior association study, to contain a mutation or mutations that underlie a phenotypic trait of interest.
- Trans
-
Located far away from the gene of interest; in practical terms, anywhere in the genome except nearby.
- Macrochaete
-
The largest bristles on flies; their function is mechanosensory.
- Pastoralism
-
The practice of tending domesticated animals for the milk they produce.
Rights and permissions
About this article
Cite this article
Wray, G. The evolutionary significance of cis-regulatory mutations. Nat Rev Genet 8, 206–216 (2007). https://doi.org/10.1038/nrg2063
Issue Date:
DOI: https://doi.org/10.1038/nrg2063
This article is cited by
-
Decoding enhancer complexity with machine learning and high-throughput discovery
Genome Biology (2023)
-
Leveraging massively parallel reporter assays for evolutionary questions
Genome Biology (2023)
-
The embryology, metamorphosis, and muscle development of Schizocardium karankawa sp. nov. (Enteropneusta) from the Gulf of Mexico
EvoDevo (2023)
-
The evolution and mutational robustness of chromatin accessibility in Drosophila
Genome Biology (2023)
-
Robustness and innovation in synthetic genotype networks
Nature Communications (2023)