Key Points
-
In most bilateral animals, the Hox-cluster genes encode a group of conserved transcription factors that are responsible for defining axial identity on the anterior–posterior (A–P) axis. They do this by activating or repressing genes in a variety of tissues throughout development.
-
Hox proteins directly regulate high-level 'executive' genes such as transcription factors and intercellular signaling molecules, as well as 'realizator' genes that affect cell adhesion, number, shape and growth.
-
The ability of different Hox proteins to select specific target enhancers is partly attributable to cooperative binding with Extradenticle (EXD)–PBX (Pre-B-cell homeobox) and Homothorax (HTH)–MEIS proteins. On other targets, Hox proteins act through monomer binding sites, with the aid of other unknown cofactors. Teashirt and Disco are good candidates for additional Hox cofactors.
-
Hox-response elements often possess EXD-binding and HTH-binding sites in addition to Hox-protein-binding sites, but otherwise they have little in common, showing great variation in size, conservation and number of Hox sites required for proper function.
-
In silico searches for new direct Hox-regulated enhancers have so far been unsuccessful, indicating that we still only have a primitive understanding of Hox-dependent transcriptional regulation.
-
The evolution of Hox gene expression patterns and protein sequences has probably contributed significantly to the current morphological diversity that is seen in bilateria.
-
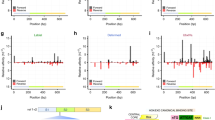
The expression patterns of Hox proteins might be modulated by several well-conserved microRNAs, including miR-10, miR-iab-4 and miR-196, which are expressed in Hox-like patterns along the A–P axis of animal embryos.
Abstract
With their power to shape animal morphology, few genes have captured the imagination of biologists as the evolutionarily conserved members of the Hox clusters have done. Recent research has provided new insight into how Hox proteins cause morphological diversity at the organismal and evolutionary levels. Furthermore, an expanding collection of sequences that are directly regulated by Hox proteins provides information on the specificity of target-gene activation, which might allow the successful prediction of novel Hox-response genes. Finally, the recent discovery of microRNA genes within the Hox gene clusters indicates yet another level of control by Hox genes in development and evolution.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Lewis, E. B. A gene complex controlling segmentation in Drosophila. Nature 276, 565–570 (1978).
Kaufman, T. C., Seeger, M. A. & Olsen, G. Molecular and genetic organization of the antennapedia gene complex of Drosophila melanogaster. Adv. Genet. 27, 309–362 (1990).
McGinnis, W. & Krumlauf, R. Homeobox genes and axial patterning. Cell 68, 283–302 (1992).
Beeman, R. W., Stuart, J. J., Haas, M. S. & Denell, R. E. Genetic analysis of the homeotic gene complex (HOM-C) in the beetle Tribolium castaneum. Dev. Biol. 133, 196–209 (1989).
Krumlauf, R. Hox genes in vertebrate development. Cell 78, 191–201 (1994).
Bienz, M. Homeotic genes and positional signalling in the Drosophila viscera. Trends Genet. 10, 22–26 (1994).
Zákány, J. & Duboule, D. Hox genes in digit development and evolution. Cell Tissue Res. 296, 19–25 (1999).
Arenas-Mena, C., Cameron, A. R. & Davidson, E. H. Spatial expression of Hox cluster genes in the ontogeny of a sea urchin. Development 127, 4631–4643 (2000).
Arenas-Mena, C., Martinez, P., Cameron, R. A. & Davidson, E. H. Expression of the Hox gene complex in the indirect development of a sea urchin. Proc. Natl Acad. Sci. USA 95, 13062–13067 (1998).
Ishii, M. et al. Hbox1 and Hbox7 are involved in pattern formation in sea urchin embryos. Dev. Growth Differ. 41, 241–252 (1999).
Finnerty, J. R., Pang, K., Burton, P., Paulson, D. & Martindale, M. Q. Origins of bilateral symmetry: Hox and dpp expression in a sea anemone. Science 304, 1335–1337 (2004). Using Hox and dpp expression patterns in sea anemone as evidence, the authors argue that bilateral symmetry was the basal state before the evolutionarily divergence of cnidarians from the ancestors of triploblastic animals such as chordates, arthropods and molluscs.
Castelli-Gair, J. & Akam, M. How the Hox gene Ultrabithorax specifies two different segments: the significance of spatial and temporal regulation within metameres. Development 121, 2973–2982 (1995).
Salser, S. & Kenyon, C. A C. elegans Hox gene switches on, off, on and off again to regulate proliferation, differentiation and morphogenesis. Development 122, 1651–1661 (1996).
Mann, R. S. & Chan, S. K. Extra specificity from extradenticle: the partnership between HOX and PBX/EXD homeodomain proteins. Trends Genet. 12, 258–262 (1996).
Chang, C. -P. et al. Pbx proteins display hexapeptide-dependent cooperative DNA binding with a subset of Hox proteins. Genes Dev. 9, 663–674 (1995).
Van Auken, K. et al. Roles of the Homothorax/Meis/Prep homolog UNC-62 and the Exd/Pbx homologs CEH-20 and CEH-40 in C. elegans embryogenesis. Development 129, 5255–5268 (2002).
Mann, R. & Affolter, M. Hox proteins meet more partners. Curr. Opin. Genet. Dev. 8, 423–429 (1998).
Kuziora, M. A. & McGinnis, W. Autoregulation of a Drosophila homeotic selector gene. Cell 55, 477–485 (1988).
Pöpperl, H. et al. Segmental expression of Hoxb-1 is controlled by a highly conserved autoregulatory loop dependent on exd/pbx. Cell 81, 1031–1042 (1995).
Gould, A., Morrison, A., Sproat, G., White, R. A. H. & Krumlauf, R. Positive cross-regulation and enhancer sharing: two mechanisms for specifying overlapping Hox expression patterns. Genes Dev. 11, 900–913 (1997).
Henderson, K. D. & Andrew, D. J. Regulation and function of Scr, exd, and hth in the Drosophila salivary gland. Developmental Biology 217, 362–374 (2000).
Azpiazu, N. & Morata, G. Functional and regulatory interactions between Hox and extradenticle genes. Genes Dev. 12, 261–273 (1998).
Weatherbee, S. D., Halder, G., Kim, J., Hudson, A. & Carroll, S. Ultrabithorax regulates genes at several levels of the wing-patterning hierarchy to shape the development of the Drosophila haltere. Genes Dev. 12, 1474–1482 (1998).
Lei, H., Wang, H., Juan, A. H. & Ruddle, F. H. The identification of Hoxc8 target genes. Proc. Natl Acad. Sci. USA 102, 2420–2424 (2005).
Williams, T. M. et al. Candidate downstream regulated genes of HOX group 13 transcription factors with and without monomeric DNA binding capability. Dev. Biol. 279, 462–480 (2005).
Cobb, J. & Duboule, D. Comparative analysis of genes downstream of the Hoxd cluster in developing digits and external genitalia. Development 132, 3055–3067 (2005). The authors use mouse microarrays to identify several genes that are regulated by the Hoxd cluster in both limb and genital appendage primordia.
Capovilla, M. & Botas, J. Functional dominance among Hox genes: repression dominates activation in the regulation of dpp. Development 125, 4949–4957 (1998).
Vachon, G. et al. Homeotic genes of the bithorax complex repress limb development in the abdomen of the Drosophila embryo through the target gene Distal-less. Cell 71, 437–450 (1992).
Liu, J. & Fire, A. Overlapping roles of two Hox genes and the exd ortholog ceh-20 in diversification of the C. elegans postembryonic mesoderm. Development 127, 5179–5190 (2000).
Garcia-Bellido, A. Homeotic and atavic mutations in insects. Am. Zool. 17, 613–629 (1977).
Yokouchi, Y. et al. Misexpression of Hoxa-13 induces cartilage homeotic transformation and changes cell adhesiveness in chick limb buds. Genes Dev. 9, 2509–2522 (1995).
Stadler, H. S., Higgins, K. M. & Capecchi, M. R. Loss of Eph-receptor expression correlates with loss of cell adhesion and chondrogenic capacity in Hoxa13 mutant limbs. Development 128, 4177–4188 (2001).
Poliakov, A., Cotrina, M. & Wilkinson, D. G. Diverse roles of eph receptors and ephrins in the regulation of cell migration and tissue assembly. Dev. Cell 7, 465–480 (2004).
Chen, J. & Ruley, H. E. An enhancer element in the EphA2 (Eck) gene sufficient for rhombomere-specific expression is activated by HOXA1 and HOXB1 homeobox proteins. J. Biol. Chem. 273, 24670–24675 (1998).
Bruhl, T. et al. Homeobox A9 transcriptionally regulates the EphB4 receptor to modulate endothelial cell migration and tube formation. Circ. Res. 94, 743–751 (2004).
Bromleigh, V. C. & Freedman, L. P. p21 is a transcriptional target of HOXA10 in differentiating myelomonocytic cells. Genes Dev. 14, 2581–2586 (2000).
Magli, M. C., Largman, C. & Lawrence, H. J. Effects of HOX homeobox genes in blood cell differentiation. J. Cell. Physiol. 173, 168–177 (1997).
Thorsteinsdottir, U. et al. Overexpression of HOXA10 in murine hematopoietic cells perturbs both myeloid and lymphoid differentiation and leads to acute myeloid leukemia. Mol. Cell. Biol. 17, 495–505 (1997).
Lohmann, I., McGinnis, N., Bodmer, M. & McGinnis, W. The Drosophila Hox gene Deformed sculpts head morphology via direct regulation of the apoptosis activator reaper. Cell 110, 457–466 (2002).
Bello, B. C., Hirth, F. & Gould, A. P. A pulse of the Drosophila Hox protein Abdominal-Aschedules the end of neural proliferation via neuroblast apoptosis. Neuron 37, 209–219 (2003).
Salser, S. J. & Kenyon, C. Activation of a C. elegans Antennapedia homologue in migrating cells controls their direction of migration. Nature 355, 255–258 (1992).
Clark, S. G., Chisholm, A. D. & Horvitz, H. R. Control of cell fates in the central body region of C. elegans by the homeobox gene lin-39. Cell 74, 43–55 (1993).
Wang, B. B. et al. A homeotic gene cluster patterns the anteroposterior body axis of C. elegans. Cell 74, 29–42 (1993).
Sun, B., Hursh, D. A., Jackson, D. & Beachy, P. A. Ultrabithorax protein is necessary but not sufficient for full activation of decapentaplegic expression in the visceral mesoderm. EMBO J. 14, 520–535 (1995).
Haerry, T. & Gehring, W. A conserved cluster of homeodomain binding sites in the mouse Hoxa-4 intron functions in Drosophila embryos as an enhancer that is directly regulated by Ultrabithorax. Dev. Biol. 186, 1–15 (1997).
Capovilla, M., Kambris, Z. & Botas, J. Direct regulation of the muscle-identity gene apterous by a Hox protein in the somatic mesoderm. Development 128, 1221–1230 (2001).
Schier A. F. & Gehring W. J. Direct homeodomain-DNA interaction in the autoregulation of the fushi tarazu gene. Nature 356, 804–807 (1992).
Zaffran, S., Kuchler, A., Lee, H. H. & Frasch, M. biniou (FoxF), a central component in a regulatory network controlling visceral mesoderm development and midgut morphogenesis in Drosophila. Genes Dev. 15, 2900–2915 (2001).
Zeng, C., Pinsonneault, J., Gellon, G., McGinnis, N. & McGinnis, W. Deformed protein binding sites and cofactor binding sites are required for the function of a small segment-specific regulatory element in Drosophila embryos. EMBO J. 13, 2362–2377 (1994).
Lou, L., Bergson, C. & McGinnis, W. Deformed expression in the Drosophila central nervous system is controlled by an autoactivated intronic enhancer. Nucleic Acids Res. 23, 3481–3487 (1995).
Galant, R., Walsh, C. M. & Carroll, S. B. Hox repression of a target gene: extradenticle-independent, additive action through multiple monomer binding sites. Development 129, 3115–3126 (2002).
Appel, B. & Sakonju, S. Cell-type-specific mechanisms of transcriptional repression by the homeotic gene products UBX and ABD-A in Drosophila embryos. EMBO J. 12, 1099–1109 (1993).
Grieder, N. C., Marty, T., Ryoo, H. D., Mann, R. S. & Affolter, M. Synergistic activation of a Drosophila enhancer by HOM/EXD and DPP signaling. EMBO J. 16, 7402–7410 (1997).
Ebner, A., Cabernard, C., Affolter, M. & Merabet, S. Recognition of distinct target sites by a unique Labial/Extradenticle/Homothorax complex. Development 132, 1591–1600 (2005). In this paper, an enhancer regulated by Labial (LAB) and Extradenticle (EXD) through an unusual binding site is serendipitously found near an apparently functionless consensus LAB–EXD binding site that was identified in silico.
Fasano, L. et al. The gene teashirt is required for the development of Drosophila embryonic trunk segments and encodes a protein with widely spaced zinc finger motifs. Cell 64, 63–79 (1991).
de Zulueta, P., Alexandre, E., Jacq, B. & Kerridge, S. Homeotic complex and teashirt genes co-operate to establish trunk segmental identities in Drosophila. Development 120, 2287–2296 (1994).
Mahaffey, J. P., Griswold, C. M. & Cao, Q. The Drosophila genes disconnected and disco-related are redundant with respect to larval head development and accumulation of mRNAs from Deformed target genes. Genetics 157, 225–236 (2001).
Robertson, L. K., Bowling, D. B., Mahaffey, J. P., Imiolczyk, B. & Mahaffey, J. W. An interactive network of zinc-finger proteins contributes to regionalization of the Drosophila embryo and establishes the domains of HOM-C-protein function. Development 131, 2781–2789 (2004). This paper provides strong genetic evidence that supports the involvement of Disco/Disco-related and Teashirt as regionalizing factors that are required by Hox proteins for axial specification.
Chan, S. K., Popperl, H., Krumlauf, R. & Mann, R. S. An extradenticle-induced conformational change in a HOX protein overcomes an inhibitory function of the conserved hexapeptide motif. EMBO J. 15, 2476–2487 (1996).
Chan, S. K., Ryoo, H. D., Gould, A., Krumlauf, R. & Mann, R. S. Switching the in vivo specificity of a minimal Hox-responsive element. Development 124, 2007–2014 (1996).
Ryoo, H. D. & Mann, R. S. The control of trunk Hox specificity and activity by Extradenticle. Genes Dev. 13, 1704–1716 (1999).
Carroll, S. B., Grenier, J. K. & Weatherbee, S. D. in From DNA to Diversity (ed. Carroll, S.) 1–214 (Blackwell Science, London, 2005).
Averof, M. & Patel, N. H. Crustacean appendage evolution associated with changes in Hox gene expression. Nature 388, 682–686 (1997).
Stern, D. L. A role of Ultrabithorax in morphological differences between Drosophila species. Nature 396, 463–466 (1998).
Hsia, C. C. & McGinnis, W. Evolution of transcription factor function. Curr. Opin. Genet. Dev. 13, 199–206 (2003).
Ronshaugen, M., McGinnis, N. & McGinnis, W. Hox protein mutation and macroevolution of the insect body plan. Nature 415, 914–917 (2002).
Galant, R. & Carroll, S. B. Evolution of a transcriptional repression domain in an insect Hox protein. Nature 415, 910–913 (2002).
Hughes, C. L. & Kaufman, T. C. Exploring the myriapod body plan: expression patterns of the ten Hox genes in a centipede. Development 129, 1225–1238 (2002). In this article, all ten centipede Hox genes are cloned and the expression patterns are analysed by in situ hybridizations on embryos, inviting comparisons with other invertebrate Hox expression patterns.
Telford, M. J. Evidence for the derivation of the Drosophila fushi tarazu gene from a Hox gene orthologous to lophotrochozoan Lox5. Curr. Biol. 10, 349–352 (2000).
Mouchel-Vielh, E., Blin, M., Rigolot, C. & Deutsch, J. S. Expression of a homologue of the fushi tarazu (ftz) gene in a cirripede crustacean. Evol. Dev. 4, 76–85 (2002).
Brown, S. J., Hilgenfeld, R. B. & Denell, R. E. The beetle Tribolium castaneum has a fushi tarazu homolog expressed in stripes during segmentation. Proc. Natl Acad. Sci. USA 91, 12922–12926 (1994).
Dawes, R., Dawson, I., Falciani, F., Tear, G. & Akam, M. Dax, a locust Hox gene related to fushi-tarazu but showing no pair-rule expression. Development 120, 1561–1572 (1994).
Lohr, U., Yussa, M. & Pick, L. Drosophila fushi tarazu: a gene on the border of homeotic function. Curr. Biol. 11, 1403–1412 (2001).
Lohr, U. & Pick, L. Cofactor-interaction motifs and the cooption of a homeotic Hox protein into the segmentation pathway of Drosophila melanogaster. Curr. Biol. 15, 643–649 (2005). This paper shows that the Hox-complex gene fushi tarazu (ftz ) can be switched between segmentation and homeotic functions by inserting or deleting alternate cofactor-interaction domains that are found in different ftz orthologues.
Pasquinelli, A. E., Hunter, S. & Bracht, J. MicroRNAs: a developing story. Curr. Opin. Genet. Dev. 15, 200–205 (2005).
Bartel, D. MicroRNAs: genomics, biogenesis, mechanism, and function. Cell 116, 281–297 (2004).
Brend, T., Gilthorpe, J., Summerbell, D. & Rigby, P. W. Multiple levels of transcriptional and post-transcriptional regulation are required to define the domain of Hoxb4 expression. Development 130, 2717–2728 (2003).
Nelson, C. E. et al. Analysis of Hox gene expression in the chick limb bud. Development 122, 1449–1466 (1996).
Abzhanov, A. & Kaufman, T. C. Novel regulation of the homeotic gene Scr associated with a crustacean leg-to-maxilliped appendage transformation. Development 126, 1121–1128 (1999).
Lewis, B., Shih, I., Jones-Rhoades, M., Bartel, D. & Burge, C. Prediction of mammalian microRNA targets. Cell 115, 787–798 (2003).
Enright, A. et al. MicroRNA targets in Drosophila. Genome Biol. 5, R1 (2003).
Yekta, S., Shih, I. & Bartel, D. MicroRNA-directed cleavage of HOXB8 mRNA. Science 304, 594–596 (2004). This article provides evidence for the cleavage of several mammalian Hox genes by a conserved microRNA that is located within the Hox clusters.
Mansfield, J. H. et al. MicroRNA-responsive 'sensor' transgenes uncover Hox-like and other developmentally regulated patterns of vertebrate microRNA expression. Nature Genet. 36, 1079–1083 (2004).
Brennecke, J., Hipfner, D. R., Stark, A., Russell, R. B. & Cohen, S. M. bantam encodes a developmentally regulated microRNA that controls cell proliferation and regulates the proapoptotic gene hid in Drosophila. Cell 113, 25–36 (2003).
Kosman, D. et al. Multiplex detection of RNA expression in Drosophila embryos. Science 305, 846 (2004).
Valoczi, A. et al. Sensitive and specific detection of microRNAs by northern blot analysis using LNA-modified oligonucleotide probes. Nucleic Acids Res. 32, e175 (2004).
Wienholds, E. et al. MicroRNA expression in zebrafish embryonic development. Science 309, 310–311 (2005). The authors use microarrays and LNA oligonucleotides to examine the expression of 115 conserved microRNAs in zebrafish. They find highly diverse expression patterns, suggesting wide-ranging developmental control.
Duboule, D. Vertebrate Hox gene regulation: clustering and/or colinearity? Curr. Opin. Genet. Dev. 8, 514–518 (1998).
Duboule, D. & Deschamps, J. Colinearity loops out. Dev. Cell 6, 738–740 (2004).
Ringrose, L. & Paro, R. Epigenetic regulation of cellular memory by the Polycomb and Trithorax group proteins. Annu. Rev. Genet. 38, 413–443 (2004).
Allen, E. et al. Evolution of microRNA genes by inverted duplication of target gene sequences in Arabidopsis thaliana. Nature Genet. 36, 1282–1290 (2004).
Popperl, H. et al. Segmental expression of Hoxb-1 is controlled by a highly conserved autoregulatory loop dependent upon Exd/Pbx. Cell 81, 1031–1042 (1995).
Gebelein, B., Culi, J., Ryoo, H. D., Zhang, W. & Mann, R. S. Specificity of Distalless repression and limb primordia development by abdominal Hox proteins. Dev. Cell 3, 487–498 (2002).
Gebelein, B., McKay, D. J. & Mann, R. S. Direct integration of Hox and segmentation gene inputs during Drosophila development. Nature 431, 653–659 (2004). A careful dissection of a DNA element that is repressed by abdominal Hox proteins reveals a multiprotein repressive complex that integrates Hox and segmentation protein inputs.
Capovilla, M., Brandt, M. & Botas, J. Direct regulation of decapentaplegic by Ultrabithorax and its role in Drosophila midgut morphogenesis. Cell 76, 461–475 (1994).
Peifer, M. & Wieschaus, E. Mutations in the Drosophila gene extradenticle affect the way specific homeo domain proteins regulate segmental identity. Genes Dev. 4, 1209–1223 (1990).
Rauskolb, C., Smith, K., Peifer, M. & Wieschaus, E. extradenticle determines segmental identities throughout Drosophila development. Development 121, 3663–3673 (1995).
Pederson, J. A. et al. Regulation by homeoproteins: a comparison of Deformed-responsive elements. Genetics 156, 667–686 (2000).
Andrew, D. J., Horner, M. A., Petitt, M. G., Smolik, S. M. & Scott, M. P. Setting limits on homeotic gene function: restraint of Sex combs reduced activity by teashirt and other homeotic genes. EMBO J. 13, 1132–1144 (1994).
Hersh, B. M. & Carroll, S. B. Direct regulation of knot gene expression by Ultrabithorax and the evolution of cis-regulatory elements in Drosophila. Development 132, 1567–1577 (2005).
Safaei R. A target of the HoxB5 gene from the mouse nervous system. Brain Res. Dev. Brain Res. 100, 5–12 (1997).
Maconochie M. K. et al. Cross-regulation in the mouse HoxB complex: the expression of Hoxb2 in rhombomere 4 is regulated by Hoxb1. Genes Dev. 11, 1885–1895 (1997).
Serpente P. et al. Direct crossregulation between retinoic acid receptor β and Hox genes during hindbrain segmentation. Development 132, 503–513 (2005).
Shi, X., Bai, S., Li, L. & Cao X. Hoxa-9 represses transforming growth factor-β-induced osteopontin gene transcription. J. Biol. Chem. 276, 850–855 (2001).
Houghton L. & Rosenthal N. Regulation of a muscle-specific transgene by persistent expression of Hox genes in postnatal murine limb muscle. Dev. Dyn. 216, 385–397 (1999).
Lampe X., Picard J. J. & Rezsohazy R. The Hoxa2 enhancer 2 contains a critical Hoxa2 responsive regulatory element. Biochem. Biophys. Res. Commun. 316, 898–902 (2004).
Graba Y. et al. DWnt-4, a novel Drosophila Wnt gene acts downstream of homeotic complex genes in the visceral mesoderm. Development 121, 209–218 (1995).
Kremser T. et al. Expression of the β3 tubulin gene (βTub60D) in the visceral mesoderm of Drosophila is dependent on a complex enhancer that binds Tinman and UBX. Dev. Biol. 216, 327–339 (1999).
Zhou B., Bagri A. & Beckendorf S. K. Salivary gland determination in Drosophila: a salivary-specific, fork head enhancer integrates spatial pattern and allows fork head autoregulation. Dev. Biol. 237, 54–67 (2001).
Heuer J. G., Li K. & Kaufman T. C. The Drosophila homeotic target gene centrosomin (cnn) encodes a novel centrosomal protein with leucine zippers and maps to a genomic region required for midgut morphogenesis. Development 121, 3861–3876 (1995).
Chan S. K. et al. Switching the in vivo specificity of a minimal Hox-responsive element. Development 124, 2007–2014 (1997).
Cui, M. & Han, M. Cis regulatory requirements for vulval cell-specific expression of the Caenorhabditis elegans fibroblast growth factor gene egl-17. Dev. Biol. 257, 104–116 (2003).
Crooks G. E., Hon G., Chandonia J. M. & Brenner S. E. WebLogo: a sequence logo generator. Genome Res. 14, 1188–1190 (2004).
Streit, A. et al. Conserved regulation of the Caenorhabditis elegans labial/Hox1 gene ceh-13. Dev. Biol. 242, 96–108 (2002).
Koh, K. et al. Cell fates and fusion in the C. elegans vulval primordium are regulated by the EGL-18 and ELT-6 GATA factors — apparent direct targets of the LIN-39 Hox protein. Development 129, 5171–5180 (2002).
McCormick A., Core N., Kerridge S., Scott M. P. Homeotic response elements are tightly linked to tissue-specific elements in a transcriptional enhancer of the teashirt gene. Development 121, 2799–2812 (1995).
Graba, Y. et al. Homeotic control in Drosophila; the scabrous gene is an in vivo target of Ultrabithorax proteins. EMBO J. 11, 3375–3384 (1992).
Strutt, D. I. & White, R. A. Characterization of T48, a target of homeotic gene regulation in Drosophila embryogenesis. Mech. Dev. 46, 27–39 (1994).
Chauvet, S. et al. dlarp, a new candidate Hox target in Drosophila whose orthologue in mouse is expressed at sites of epithelium/mesenchymal interactions. Dev. Dyn. 218, 401–413 (2000).
Manak J. R., Mathies L. D. & Scott M. P. Regulation of a decapentaplegic midgut enhancer by homeotic proteins. Development 120, 3605–3612 (1994).
Gould A. P. & White R. A. Connectin, a target of homeotic gene control in Drosophila. Development 116, 1163–1174 (1992).
Mastick G. S., McKay R., Oligino T., Donovan K. . & Lopez A. J. Identification of target genes regulated by homeotic proteins in Drosophila melanogaster through genetic selection of Ultrabithorax protein-binding sites in yeast. Genetics 139, 349–363 (1995).
Grienenberger A. et al. TGF-β signaling acts on a Hox response element to confer specificity and diversity to Hox protein function. Development 130, 5445–5455 (2003).
Hooiveld, M. H. et al. Novel interactions between vertebrate Hox genes. Int. J. Dev. Biol. 43, 665–674 (1999).
Morsi El-Kadi A. S., in der Reiden, P., Durston, A. & Morgan, R. The small GTPase Rap1 is an immediate downstream target for Hoxb4 transcriptional regulation. Mech. Dev. 113, 131–139 (2002).
Theokli, C., Morsi El-Kadi, A. S. & Morgan, R. TALE class homeodomain gene Irx5 is an immediate downstream target for Hoxb4 transcriptional regulation. Dev. Dyn. 227, 48–55 (2003).
Morgan, R., Nalliah, A. & Morsi El-Kadi, A. S. FLASH, a component of the FAS-CAPSASE8 apoptotic pathway, is directly regulated by Hoxb4 in the notochord. Dev. Biol. 265, 105–112 (2004).
Acknowledgements
We thank the National Institutes of Health (NIH) and the National Science Foundation (NSF) for funding support, and our colleagues in the Hox and microRNA fields for illuminating conversations.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Related links
Related links
DATABASES
Entrez
SwissProt
FURTHER INFORMATION
Glossary
- HOMEOTIC TRANSFORMATION
-
The transformation of one body region into the likeness of another.
- BILATERIA
-
A phylogenetic subdivision of animals that is characterized by left–right symmetry along the primary body axis at some stage of the life cycle.
- ORAL–ABORAL AXIS
-
The body axis from the mouth to the body surface opposite the mouth, commonly used in animals that have no obvious bilateral symmetry.
- PARALOGUES
-
Genes in the same organism that have evolved from a gene duplication, usually with a subsequent, sometimes subtle, divergence of function.
- RHOMBOMERE
-
Each of seven neuroepithelial segments found in the embryonic hindbrain that adopt distinct molecular and cellular properties, restrictions in cell mixing, and ordered domains of gene expression.
- WEBLOGO
-
A web based application for generating sequence logos — graphical representations of an amino-acid or nucleic-acid motif weight matrix.
- MicroRNAS
-
Small, non-coding RNAs that are components of a large protein complex (RISC) and are involved in repression of protein production from mRNAs that contain sequences with significant complementarity to the microRNA.
- VISCERAL MESODERM
-
A subset of mesodermal cells that surround endodermal tissues such as the gut, also known as splanchnic mesoderm.
- AUTOPODS
-
Distal limb primordia that give rise to structures such as the mouse paw.
- COS7 CELLS
-
A commonly used fibroblast-like cell line that is derived from African green monkey kidneys.
- MYELOMONOCYTIC CELLS
-
Pluripotent cells that differentiate into immune cells such as granulocytes, dendritic cells and monocytes.
- MONOCYTES
-
Immune cells that circulate in the blood stream and engulf foreign invaders. They can migrate into tissues, where they differentiate into macrophages.
- NEUROMERES
-
Segmentally repeated subunits of the developing CNS.
- FOOTPRINTING
-
An in vitro assay that identifies regions of DNA that are protected from digestion by DNaseI, thereby indicating the presence of bound transcription factors.
- IMAGINAL DISC
-
Sacs of cells in larval stages of holometabolous insects that divide and differentiate to form most adult tissues.
- HALTERE
-
A balancing organ that is located on the third thoracic segment in Diptera and is an evolutionary modification of a wing.
- Q NEUROBLAST
-
A C. elegans neural lineage that divides to make QL and QR neuroblasts, which in turn generate identical neural cells on left and right sides of the body.
- CELL AUTONOMOUS
-
If the activity of a gene has effects only in the cells that express it, its function is said to be cell autonomous; if it causes effects in cells other than (or in addition to) those expressing it, its function is cell non-autonomous.
- RNA-INDUCED SILENCING COMPLEX
-
A large protein complex that packages microRNAs or siRNAs, silencing expression of proteins from target mRNAs by endonucleolytic cleavage or other unknown mechanisms, depending on the complementarity of mRNA sequences to the packaged small RNAs.
- GNATHAL APPENDAGES
-
Outgrowths, usually from head segments, that are used to aid feeding.
- RACE CLONES
-
Partial cDNA sequences that are generated from transcripts by the rapid amplification of cDNA ends (RACE) to determine the start and end points of gene transcription.
- LOCKED NUCLEIC ACID OLIGONUCLEOTIDE PROBES
-
Modified nucleic acids that have increased thermal stability relative to DNA or RNA when complexed with complementary DNA or RNA.
- DEUTEROSTOME
-
A major bilaterian subdivision that includes chordates and echinoderms.
- PROTOSTOME
-
One of two major subdivision of bilateria, to which arthropods and molluscs belong.
Rights and permissions
About this article
Cite this article
Pearson, J., Lemons, D. & McGinnis, W. Modulating Hox gene functions during animal body patterning. Nat Rev Genet 6, 893–904 (2005). https://doi.org/10.1038/nrg1726
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrg1726
This article is cited by
-
Single-cell analysis reveals the spatial-temporal expression of genes associated with esophageal malformations
Scientific Reports (2024)
-
Orthodenticle homeobox 2 is transported to lysosomes by nuclear budding vesicles
Nature Communications (2023)
-
Conserved stromal–immune cell circuits secure B cell homeostasis and function
Nature Immunology (2023)
-
Trimeric complexes of Antp-TBP with TFIIEβ or Exd modulate transcriptional activity
Hereditas (2022)
-
Long noncoding RNA HOXC-AS3 interacts with CDK2 to promote proliferation in hepatocellular carcinoma
Biomarker Research (2022)