Key Points
-
The origins of domestic livestock have long been controversial and have presented a powerful challenge to archeozoologists. Molecular analysis is now helping to unravel when and where livestock domestication occurred.
-
The use of mitochondrial DNA sequencing has revealed a surprisingly complex history of domestication, with numerous mitochondrial lineages being the rule in sequences from the DNA of modern and ancient livestock samples.
-
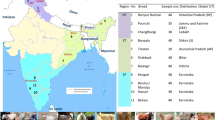
One pattern that emerges in several species (cattle, sheep and pigs, for example) is that they are the result of apparently independent domestication events in the Middle/Near east and east Asia. Africa also seems to have been an important region for cattle domestication.
-
Other molecular markers, such as microsatellites, can give more insight into the recent demographic history of domestic breeds, such as the effects of population bottlenecks, selection, genetic drift and inbreeding, and introgression. The influence of these different demographic events is extremely strong in some breeds.
-
Modern-day patterns of genetic diversity in livestock therefore only partially reflect this complex origin, owing to subsequent translocations by humans and the effects of introgression between different DNA lineages. This makes interpreting the genetic diversity in modern livestock extremely challenging.
-
Conservation of livestock diversity must account for the complex histories of the livestock breeds now under threat of extinction across the globe. A range of molecular markers, especially including those that reflect their evolution, recent demographics and economic potential should be applied in combination to provide optimal data for livestock management and conservation.
Abstract
A series of recent genetic studies has revealed the remarkably complex picture of domestication in both New World and Old World livestock. By comparing mitochondrial and nuclear DNA sequences of modern breeds with their potential wild and domestic ancestors, we have gained new insights into the timing and location of domestication events that produced the farm animals of today. The real surprise has been the high number of domestication events and the diverse locations in which they took place — factors which could radically change our approach to conserving livestock biodiversity resources in the future.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Salamini, F., Özkan, H., Brandolini, A., Schäfer-Pregl, R. & Martin, W. Genetics and geography of wild cereal domestication in the near east. Nature Rev. Genet. 3, 429–441 (2002).
MacHugh, D. E. & Bradley, D. G. Goats buck the trend. Proc. Natl Acad. Sci. USA 98, 5382–5384 (2001).
Goldstein, D. B. & Chikhi, L. Human migrations and population structure: what we know and why it matters. Annu. Rev. Genomics Hum. Genet. 3, 129–152 (2002).
Savolainen, P., Zhang, Y. P., Lu, J., Lundeberg, J. & Leitner, T. Genetic evidence for an East Asian origin of domestic dogs. Science 298, 1610–1613 (2002).
Rosenberg, N. A. et al. Empirical evaluation of genetic clustering methods using multilocus genotypes from 20 chicken breeds, Genetics 159, 699–713 (2001).
Food and Agriculture Organization of the United Nations. Secondary Guidelines for the Development of National Farm Animal Genetic Resources Management: Management of Small Populations at Risk. (Food and Agriculture Organization, Rome, 1998).
Froufe, E., Magyary, I., Lehoczky, I. & Weiss, S. MtDNA sequence data supports an Asian ancestry and single introduction of the common carp into the Danube basin. J. Fish Biol. 61, 301–304 (2002).
Blumler, M. A. Independent inventionism and recent genetic evidence on plant domestication. Econ. Bot. 46, 98–111 (1992).
Sherratt, A. Climatic cycles and behavioural revolutions: the emergence of modern humans and the beginning of farming. Antiquity 71, 271–287 (1997).
Kealhofer, L. Changing perceptions of risk: the development of agro-ecosystems in Southeast Asia. Am. Anthropol. 104, 178–194 (2002).
Diamond, J. Evolution, consequences and future of plant and animal domestication. Nature 418, 700–707 (2002). This review summarizes and synthesizes the different and complementary kinds of data (genetic, languistic, archaeozoological and so on) that are being used to infer the origins and spread of domestic agricultural plants and animals.
Richerson, P. J., Boyd, R. & Bettinger, R. L. Was agriculture impossible during the Pleistocene but mandatory during the Holocene? A climate change hypothesis. Am. Antiq. 66, 387–411 (2001).
Jansen, T. et al. Mitochondrial DNA and the origins of the domestic horse. Proc. Natl Acad. Sci. USA 99, 10905–10910 (2002).
Vila, C. et al. Widespread origins of domestic horse lineages. Science 291, 474–477 (2001).
Balasse, M., Bocherens, H., Tresset, A., Mariotti, A. & Vigne, J. D. Emergence of dairy production in the Neolithic? Contribution of isotopic analysis of cattle archaeological bones. Comptes Rendus Acad. Sci. Ser. II. 325, 1005–1010 (1997).
Alvard, M. S. & Kuznar, L. Deferred harvests: the transition from hunting to animal husbandry. Am. Anthropol. 103, 295–311 (2001).
Zeder, M. A. & Hesse, B. The initial domestication of goats (Capra hircus) in the Zagros mountains 10,000 years ago. Science 287, 2254–2257 (2000). This paper uses numerous precise carbon dates and a shift in demographic profiles (age and sex of harvested males) from goat fossil material to provide some of the oldest and best evidence for goat domestication along the border of Iraq and Iran.
Mannion, A. M. Domestication and the origins of agriculture: an appraisal. Phys. Geog. 23, 37–56 (1999).
Karp, A. et al. Molecular technologies for biodiversity evaluation: opportunities and challenges. Nature Biotech. 15, 625–628 (1997).
Sunnucks, P. Efficient genetic markers for population biology Trends Ecol. Evol. 15, 199–203 (2000).
Loftus, R. T., MacHugh, D. E., Bradley, D. G., Sharp, P. M. & Cunningham, P. Evidence for two independent domestications of cattle. Proc. Natl Acad. Sci. USA 91, 2757–2761 (1994). This paper was the first in a series of investigations of domestic animals that pointed towards numerous geographically separated domestications.
MacHugh, D. E., Loftus, R. T., Bradley, D. G., Sharp, P. M. & Cunningham, P. Microsatellite DNA variation within and among European cattle breeds. Proc. R. Soc. Lond. B 256, 25–31 (1994).
Handt, O., Meyer, S. & von Haeseler, A. Compilation of human mtDNA control region sequences. Nucleic Acids Res. 26, 126–129 (1998).
Luikart, G. et al. Multiple maternal origins and weak phylogeographic structure in domestic goats. Proc. Natl Acad. Sci. USA 98, 5927–5932 (2001). This study detected extremely high mtDNA diversity and surprisingly little intercontinental differentiation in goats as compared to cattle and sheep. It identified three potential origins (that is, three divergent mtDNA lineages), including one from south or east Asia, and suggested an emerging pattern of East–West dual domestications in farm animals.
Avise, J. C. Molecular Markers, Natural History and Evolution (Kluwer Academic, Boston, MA, 1993).
MacHugh, D. E., Shriver, M. D., Loftus, R. T., Cunningham, P. & Bradley, D. G. Microsatellite DNA variation and the evolution, domestication and phylogeography of taurine and zebu cattle (Bos taurus and Bos indicus). Genetics 146, 1071–1086 (1997). The novelty of this work was the uncoupling of paternal and maternal lineages in domestic population genetic history. African cattle populations show substantial Bos indicus ancestry in their Y chromosomes and autosomes, but lack any mtDNA variants from that taxon.
Buntjer, J. B., Otsen, M., Nijman, I. J., Kuiper, M. T. R. & Lenstra, J. A. Phylogeny of bovine species based on AFLP fingerprinting. Heredity 88, 46–51 (2002).
Ajmone Marsan, P. et al. Genetic distances within and across cattle breeds as indicated by biallelic AFLP markers. Anim. Genet. 33, 280–286 (2002).
Bruford, M. W. & Wayne, R. K. Microsatellites and their application to population genetics. Curr. Opin. Genet. Dev. 3, 939–943 (1993).
Diez-Tascón, C., Littlejohn, R. P., Almeida, P. A. R. & Crawford, A. M. Genetic variation within the Merino sheep breed: analysis of closely related populations using microsatellites. Anim. Genet. 31, 243–251 (2000).
Kadwell, M. et al. Genetic analysis reveals the wild ancestors of the llama and alpaca. Proc. R. Soc. Lond. B 268, 2575–2584 (2001). This study integrated mtDNA and microsatellite analyses to elucidate the origins of South American livestock despite their recurrent hybridization.
Matsuoka, Y. et al. A single domestication for maize shown by multilocus microsatellite genotyping. Proc. Natl Acad. Sci. USA 99, 6080–6084 (2002).
Cornuet, J. M., Piry, S., Luikart, G., Estoup, A. & Solignac, M. New methods employing multilocus genotypes to select or exclude populations as origins of individuals. Genetics 153, 1989–2000 (1999).
Maudet, C., Luikart, G. & Taberlet, P. Genetic diversity and assignment tests among seven French cattle breeds based on microsatellite DNA analysis. J. Anim. Sci. 80, 942–950 (2002).
Luikart, G. & Cornuet, J. -M. Empirical evaluation of a test for detecting recent historical population bottlenecks. Conserv. Biol. 12, 228–237 (1998).
Stanley, H. F., Kadwell, M. & Wheeler, J. C. Molecular evolution of the family Camelidae: a mitochondrial study. Proc. R. Soc. Lond. B 256, 1–6 (1994).
Wheeler, J. C. Evolution and present situation of the South American Camelidae. Biol. J. Linn. Soc. 54, 271–295 (1995).
Bradley, D. G., MacHugh, D. E., Cunningham, P. & Loftus, R. T. Mitochondrial diversity and the origins of African and European cattle. Proc. Natl Acad. Sci. USA 93, 5131–5135 (1996).
Loftus, R. T. et al. Mitochondrial genetic variation in European, African and Indian cattle populations. Anim. Genet. 25, 265–271 (1994).
Perkins, D., Jr. Fauna of Çatal Hüyük: evidence for early cattle domestication in Anatolia. Science 164, 177–178 (1969).
Vernesi, C. et al. Genetic characterization of the body attributed to the evangelist Luke. Proc. Natl Acad. Sci. USA 98, 13460–13463 (2001).
Troy, C. S. et al. Genetic evidence for Near-Eastern origins of European cattle. Nature 410, 1088–1091 (2001). Ancient and modern mtDNA phylogeography in European cattle indicates a derived origin from the Near East and supports a different history for African cattle.
Leonard, J. A. et al. Ancient DNA evidence for Old World origin of New World dogs. Science 298, 1613–1616 (2002).
Lister, A. M. et al. Ancient and modern DNA from a variety of sources in a study of horse domestication. Anc. Biomol. 2, 267–280 (1998).
Hiendleder, S., Mainz, K., Plante, Y. & Lewalski, H. Analysis of mitochondrial DNA indicates that domestic sheep are derived from two ancestral maternal sources: no evidence for contributions from urial and argali sheep. J. Hered. 89, 113–120 (1998).
Hiendleder, S., Kaupe, B., Wassmuth, R. & Janke, A. Molecular analysis of wild and domestic sheep questions current nomenclature and provides evidence for domestication from two different subspecies. Proc. R. Soc. Lond. B 269, 893–904 (2002).
Lau, C. H. et al. Genetic diversity of Asian water buffalo (Bubalus bubalis): mitochondrial D-loop and cytochrome b sequence variation. Anim. Genet. 29, 253–264 (1998).
Tanaka, K. et al. Phylogenetic relationship among all living species of the genus Bubalus based on DNA sequences of the cytochrome b gene. Biochem. Genet. 34, 443–452 (1996).
Guiffra, E. et al. The origin of the domestic pig: independent domestication and subsequent introgression. Genetics 154, 1785–1791 (2000). A clear illustration of the separation of the eastern and western clades of domestic pig, including comparisons of both mtDNA and nuclear gene sequences.
Kijas, J. M. H. & Andersson, L. A phylogenetic study of the origin of the domestic pig estimated from the near-complete mtDNA genome. J. Mol. Evol. 52, 302–308 (2001).
Watanobe, T. et al. Prehistoric introduction of domestic pigs onto the Okinawa islands: ancient mitochondrial DNA evidence. J. Mol. Evol. 55, 222–231 (2002).
Townsend, S. J. Genetic diversity and domestication in sheep (Ovis). Thesis, Univ. East Anglia (2000).
Kim, K. I., Lee, J. H., Lee, S. S. & Yang, Y. H. Phylogenetic relationships of northeast Asian cattle to other cattle populations determined using mitochondrial DNA D-loop sequence polymorphism. Biochem. Genet. 41, 91–98 (2003).
Kikkawa, Y. et al. Phylogenies using mtDNA and SRY provide evidence for male-mediated introgression in Asian domestic cattle. Anim. Genet. 34, 96–101 (2003).
Miretti, M. M., Pereira, H. A., Poli, M. A., Contel, E. P. B. & Ferro, J. A. African-derived mitochondria in South American native cattle breeds (Bos taurus): evidence of a new taurine mitochondrial lineage. J. Hered. 93, 323–330 (2002).
Hanotte, O. et al. African pastoralism: genetic imprints of origins and migrations. Science 296, 336–339 (2002). This applied the synthetic map approach of Cavalli-Sforza and colleagues to cattle diversity on a well-sampled continent and uncovered separate and interpretable levels of genetic variation.
Wood, N. J. & Phua, S. H. Variation in the control region sequence of the sheep mitochondrial genome. Anim. Genet. 27, 25–33 (1996).
Porter, V. Goats of the World (Farming Press, Ipswich, UK, 1996).
Yerxat, M. Y. Application of wild goats in cashmere breeding. Small Ruminant Res. 15, 287–291 (1995).
Moritz, C. Applications of mitochondrial DNA analysis in conservation — a critical review. Mol. Ecol. 3, 401–411 (1994).
Weitzman, S. 'On diversity'. Quart. J. Econ. 107, 363–405 (1992).
Hanotte, O. et al. Geographic distribution and frequency of a taurine Bos taurus and an indicine Bos indicus Y specific allele amongst sub-Saharan African cattle breeds. Mol. Ecol. 9, 387–396 (2000).
Heaton, M. P. et al. Selection and use of SNP markers for animal identification and paternity analysis in US beef cattle. Mamm. Genome 13, 272–281 (2002).
Irwin, D. M., Kocher, T. D. & Wilson, A. C. Evolution of the cytochrome-b gene of mammals. J. Mol. Evol. 32, 128–144 (1991).
Avise, J. C. Phylogeography: The History and Formation of Species (Harvard Univ. Press, Harvard, 2000).
Bandelt, H. J., Forster, P., Sykes, B. C. & Richards, M. B. Mitochondrial portraits of human populations using median networks. Genetics 141, 743–753 (1995).
Bandelt, H. J., Forster, P. & Röhl, A. Median joining networks for inferring intraspecific phylogenies. Mol. Biol. Evol. 16, 37–48 (1999).
Rogers, A. R. & Harpending, H. Population growth makes waves in the distribution of pairwise genetic differences. Mol. Biol. Evol. 9, 552–569 (1992).
Loftus, R. T. et al. A microsatellite survey of cattle from a centre of origin: the Near East. Mol. Ecol. 8, 2015–2022 (1999).
Steinborn, R. et al. Coexistence of Bos taurus and Bos indicus mitochondrial DNAs in nuclear transfer-derived somatic cattle clones. Genetics 162, 823–829 (2002).
Watanobe, T. et al. Genetic relationship and distribution of the Japanese wild boar (Sus scrofa leucomystax) and Ryukyu wild boar (Sus scrofa riukiuanus) analysed by mitochondrial DNA. Mol. Ecol. 8, 1509–1512 (1999).
Brown, W. M., George, M. and Wilson, A. C. Rapid evolution of animal mitochondrial DNA. Proc. Natl Acad. Sci. USA 76, 1967–1971 (1979).
Horai, S., Hayasaka, K., Kondo, R., Tsugane, K. & Takahata, N. Recent African origin of modern humans revealed by complete sequences of hominid mitochondrial DNAs. Proc. Natl Acad. Sci. USA 92, 532–536 (1995).
IUCN/SSC Caprinae Specialist Group. Wild Sheep and Goats and their Relatives (Shackleton, D. M., ed.) (Gland, Switzerland and Cambridge, UK, 1997).
Acknowledgements
M.W.B. would like to acknowledge funding from the UK Ministry of Agriculture, Fisheries and Food and the European Commission (ECONOGENE project). S. Townsend, K. Byrne, L. Chikhi, G.M. Hewitt, L. Alderson, The Rare Breeds Survival Trust, J. Wheeler, R, Rosadio, M. Kadwell and M. Fernandez have all contributed significantly to M.W.B.'s livestock research. G.L. was supported by grants from NATO, the German Federal Agency for Nature Conservation (Scientific Authority to CITES), the French CNRS/INSERM sequencing programme and the European Commission (ECONOGENE project).
Author information
Authors and Affiliations
Corresponding author
Related links
Glossary
- DOMESTICATION
-
The process of genetically adapting an animal or plant to better suit the needs of human beings (for example, breeding cattle for milk production).
- ANTHROPOLOGY
-
The study of humans and non-human primates, which includes the comparative study of societies and cultures and the science of human zoology and evolution.
- ARCHAEOBIOLOGY
-
The study of non-human animal, plant and microbial remains in archaeological sites.
- ANIMAL GENETIC RESOURCES
-
(AnGR). Genetic diversity, either characterized or as yet uncharacterized, that is found in economically important animals, plants and microbes. This does not indicate that these species are necessarily domesticated.
- CAMELID
-
A mammal of the Camelidae family comprising camels, the llama and its relatives (for example, the alpaca).
- ARCHAEOZOOLOGY
-
The study of non-human animal remains in archaeological sites.
- FERTILE CRESCENT
-
A region that spans modern-day Israel, Jordan, Lebanon and western Syria, into southeast Turkey and, along the Tigris and Euphrates rivers, into Iraq and the western flanks of Iran.
- HAPLOTYPES
-
Genetic loci which co-segregate or are in linkage disequilibrium (LD). As mitochondrial DNA is a small extranuclear molecule, which does not undergo recombination, all markers on this genome are effectively linked as a single haplotype. At present, LD mapping is being used effectively in several species, including humans and livestock, to identify regions of the nuclear genome that have undergone intense episodes of selection.
- MOLECULAR CLOCK
-
The principle that any gene or protein has a near-constant long-term rate of evolution in all branches of a clade, which means that the amount of sequence divergence between two sequences will be proportional to the amount of time elapsed since their shared ancestor existed.
- AMPLIFIED FRAGMENT LENGTH POLYMORPHISM
-
(AFLP). PCR-linker-generated multifragment profiles (DNA fingerprints) that are predominantly inherited in a dominant fashion, but that have recently proved useful as tools for genetic diversity estimation and in genome mapping projects.
- BOTTLENECKS
-
Episodes of demographic contraction (small population size) that might result in reduced genetic variation and loss of viability of populations in future generations in the absence of immigration of new genetic material.
- PARSIMONY
-
As applied to phylogenetic reconstruction, a criterion for estimating historical changes by minimizing the number of substitution events that are required to explain how one DNA sequence evolves into another.
- RESTRICTION FRAGMENT LENGTH POLYMORPHISM
-
(RFLP). A fragment length variant in DNA sequences that is generated through the gain or loss of a restriction site owing to a DNA substitution.
- MAXIMUM-LIKELIHOOD
-
A method that selects the phylogenetic tree that has the highest probability of explaining the sequence data, under a specific model of substitution (changes in the nucleotide or amino-acid sequence).
- NEIGHBOUR-JOINING
-
A distance-based molecular phylogenetic method that involves the sequential addition of taxa and the minimization of branch lengths, but does not assume a molecular clock.
- BOOTSTRAP ANALYSIS
-
A type of statistical analysis to test the reliability of certain branches in an evolutionary tree. The bootstrap proceeds by re-sampling the original data, with replacement, to create a series of bootstrap samples of the same size as the original data. The bootstrap value of a node is the percentage of times that a node is present in the set of trees that is constructed from the new data sets.
- COALESCENCE
-
The joining of genetic lineages to common ancestors when they are traced backwards in time.
- INTROGRESSION
-
The transfer or genetic material from one species to another by hybridization and repeated backcrossing.
- EVOLUTIONARILY SIGNIFICANT UNIT
-
Populations that share a common ancestor, but that have been demographically independent for long enough that they no longer share mitochondrial DNA haplotypes.
- MANAGEMENT UNIT
-
Populations that have significant allele or haplotype frequency differences at mitochondrial DNA and nuclear DNA.
Rights and permissions
About this article
Cite this article
Bruford, M., Bradley, D. & Luikart, G. DNA markers reveal the complexity of livestock domestication. Nat Rev Genet 4, 900–910 (2003). https://doi.org/10.1038/nrg1203
Issue Date:
DOI: https://doi.org/10.1038/nrg1203
This article is cited by
-
Mitogenome sequences of domestic cats demonstrate lineage expansions and dynamic mutation processes in a mitochondrial minisatellite
BMC Genomics (2023)
-
Managing genomic diversity in conservation programs of Chinese domestic chickens
Genetics Selection Evolution (2023)
-
Universal mtDNA fragment for Cervidae barcoding species identification using phylogeny and preliminary analysis of machine learning approach
Scientific Reports (2023)
-
The genomic landscape of mammal domestication might be orchestrated by selected transcription factors regulating brain and craniofacial development
Development Genes and Evolution (2023)
-
Excessive replacement changes drive evolution of global sheep prion protein (PRNP) sequences
Heredity (2022)