Key Points
-
Introgression is the permanent integration of genes into a new genome through several generations of hybridization and backcrossing.
-
Like all genes, transgenes are subject to new selective pressures when they are transferred to another genome through hybridization. It is the new selective regime that determines whether or not transgenes will be stably introgressed.
-
The introgression of transgenes from sorghum to johnsongrass should be considered as a high risk. Crops of medium introgression risk are alfalfa, wheat, canola and sunflower.
-
Transgenes that confer herbicide tolerance will be intrinsically subject to high selective pressure in many agricultural environments, but selectively neutral outside cultivation. By contrast, other 'fitness enhancing' transgenes will have lower selective pressure in agricultural fields and variable selective advantage in nature.
-
Linkage disequilibrium (LD) is expected to be an important genomic characteristic with regards to the introgression of transgenes. If a transgene is integrated into a genomic region that is not normally subject to introgression, it would not be expected to introgress.
-
Strategies to decrease or prevent transgene introgression can manipulate selection using LD through transgene placement or engineering tandem transgenes. Biotechnological tools such as transplastomics, gene-use restriction technologies and male sterility can also be developed to decrease introgression in certain crops.
-
The risks of introgression for crops and transgenes should be assessed on a case-by-case basis. Ultimately, ecological and agronomical risk is determined more by transgene performance in a specific genomic location in a plant than by introgression per se.
Abstract
Transgenes engineered into annual crops could be unintentionally introduced into the genomes of their free-living wild relatives. The fear is that these transgenes might persist in the environment and have negative ecological consequences. Are some crops or transgenic traits of more concern than others? Are there natural genetic barriers to minimize gene escape? Can the genetic transformation process be exploited to produce new barriers to gene flow? Questions abound, but luckily so do answers.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Quist, D. & Chapela, I. H. Transgenic DNA introgressed into traditional maize landraces in Oaxaca, Mexico. Nature 414, 541–543 (2001).
Editorial note. Nature 416, 600 (2002).
Rieseberg, L. H. & Wendel, J. F. in Hybrid Zones and the Evolutionary Process (ed. Harrison, R. G.) 70–109 (Oxford Univ. Press, New York, 1993). This extensive review showed how the use of molecular markers has transformed our understanding of introgression.
Metz, M. & Fütterer, J. Suspect evidence of transgenic contamination. Nature 416, 600–601 (2002).
Stebbins, G. L. The inviability, weakness, and sterility of interspecific hybrids. Adv. Genet. 9, 147–215 (1958).
Anderson, E. Introgressive Hybridization (Wiley, New York, 1949). This book was the first to recognize the evolutionary implications of introgressive hybridization, develop methods for its analysis and provide experimental studies of introgression.
Heiser, C. B. Introgression re-examined. Bot. Rev. 39, 347–366 (1973).
Grant, V. Plant Speciation 2nd edn (Columbia Univ. Press, New York, 1981).
Abbott, R. J. Plant invasions, interspecific hybridization and the evolution of plant taxa. Trends Ecol. Evol. 7, 401–405 (1992).
Ellstrand, N. C. & Schierenbeck, K. Hybridization as a stimulus for the evolution of invasiveness in plants? Proc. Natl Acad. Sci. USA 97, 7043–7050 (2000).
Abbott, R. J., James, J. K., Milne, R. I. & Gillies, A. C. M. Plant introductions, hybridization and gene flow. Phil. Trans. R. Soc. Lond. B 358, 1123–1132 (2003).
Rieseberg, L. H. & Brunsfeld, S. J. in Molecular Systematics of Plants (eds Soltis, P. S., Soltis, D. E. & Doyle, J. J.) 151–176 (Chapman and Hall, New York, 1992).
Rieseberg, L. H., Baird, S. J. E. & Gardner, K. A. Hybridization, introgression, and linkage evolution. Plant Mol. Biol. 42, 205–224 (2000). This excellent review makes a convincing case for using molecular map-based approaches for the study of hybridization and introgression.
Harrison, R. G. Hybrid zones: windows on evolutionary process. Oxford Surv. Evol. Biol. 7, 69–128 (1990).
Burke, J. M., Voss, T. J. & Arnold, M. L. Genetic interactions and natural selection in Louisiana iris hybrids. Evolution 52, 1304–1310 (1998).
Rieseberg, L. H., Whitton, J. & Gardner, K. Hybrid zones and the genetic architecture of a barrier to gene flow between two wild sunflower species. Genetics 152, 713–727 (1999).
Barton, N. H. & Hewitt, G. M. Analysis of hybrid zones. Ann. Rev. Ecol. Syst. 16, 113–148 (1985).
Kalloo, G. & Chowdhury, J. B. Distant Hybridization of Crop Plants (Springer, Heidelberg, 1992).
Zamir, D. Improving plant breeding with exotic genetic libraries. Nature Rev. Genet. 2, 983–989 (2001).
Tanksley, S. D. & McCouch, S. R. Seed banks and molecular maps: unlocking genetic potential from the wild. Science 227, 1063–1066 (1997).
Harlan, J. R. Crops and Man (Crop Science Soc. of America, Madison, Wisconsin, 1975).
Simmonds, N. W. Principles of Crop Improvement (Longman, New York, 1979).
Baker, H. G. in The Genetics of Colonizing Species (eds Baker, H. G. & Stebbins, G. L.) 147–168 (Academic, New York, 1965). In this paper, weediness traits are described and compared with domestication traits.
Baker, H. G. The evolution of weeds. Ann. Rev. Ecol. Syst. 5, 1–24 (1974).
Charlesworth, D., Charlesworth, B. & Morgan, M. T. The pattern of neutral molecular variation under the background selection model. Genetics 141, 1619–1632 (1995).
Cummings, M. P. & Clegg, M. T. Nucleotide sequence diversity at the alcohol dehydrogenase 1 locus in wild barley (Hordeum vulgare ssp. spontaneum): an evaluation of the background hypothesis. Proc. Natl Acad. Sci. USA 95, 5637–5642 (1998).
Remington, D. L. et al. Structure of linkage disequilibrium and phenotypic associations in the maize genome. Proc. Natl Acad. Sci. USA 98, 11479–11484 (2001).
Haygood, R., Ives, A. R. & Andow, D. A. Consequences of recurrent gene flow from crops to wild relatives. Proc. R. Soc. Lond. B 25 July 2003 (doi: 10.1098/rspb.2003).
Ellstrand, N. C., Prentice, H. C. & Hancock, J. F. Gene flow and introgression from domesticated plants into their wild relatives. Ann. Rev. Ecol. Syst. 30, 539–563 (1999).
Raybould, A. & Gray, A. J. Genetically modified crops and hybridization with wild relatives: a UK perspective. J. Appl. Ecol. 30, 199–219 (1993). This much cited review was one of the first to ascribe gene-flow risks for transgenic crops.
Hancock, J. F. A framework for assessing the risk of transgenic crops. Bioscience 53, 512–519 (2003).
Gressel, J. & Rotteveel, A. W. Genetic and ecological risks from biotechnologically-derived herbicide-resistant crops: decision trees for risk assessment. Plant Breed. Rev. 18, 251–303 (2000).
Shoemaker, R. C. et al. Genome duplication in soybean (Glycine subgenus soja). Genetics 144, 329–338 (1996).
Concibido, V. C. et al. Introgression of a quantitative trait locus for yield from Glycine soja into commercial soybean cultivars. Theor. Appl. Genet. 106, 575–582 (2003).
Leroy, A. R., Fehr, W. R. & Cianzio, S. R. Introgression of genes for small seed size from Glycine soja into G. max. Crop Sci. 31, 693–697 (1991).
Simpson, C. E. Use of wild Arachis species/introgression of genes into A. hypogaea L. Peanut Sci. 28, 114–116 (2001).
Garcia, G. M., Stalker, H. T. & Kochert, G. Introgression analysis of an interspecific hybrid population in peanuts (Arachis hypogaea L.) using RFLP and RAPD markers. Genome 38, 166–176 (1995).
Beebe, S., Toro, C. O., González, A. V., Chacón, M. I. & Debouck, D. G. Wild-weed–crop complexes of common bean (Phaseolus vulgaris L., Fabaceae) in the Andes of Peru and Colombia, and their implications for conservation and breeding. Genet. Res. Crop Evol. 44, 73–91 (1997).
Johns, M. A. et al. Gene pool classification of common bean landraces from Chile based on RAPD and morphological data. Crop Sci. 37, 605–613 (1997).
Doebley, J. Molecular evidence for gene flow among Zea species. Bioscience 40, 443–448 (1990).
Doebley, J. F. Molecular evidence and the evolution of maize. Econ. Bot. 44 (Suppl.), 6–27 (1990).
Matsuoka, Y. et al. A single domestication for maize shown by multilocus microsatellite genotyping. Proc. Natl Acad. Sci USA 99, 6080–6084 (2002).
Doebley, J., Goodman, M. M. & Stuber, C. W. Patterns of isozyme variation between maize and Mexican annual teosinte. Econ. Bot. 41, 234–246 (1987).
Majumder, N. D., Ram, T. & Sharma, A. C. Cytological and morphological variation in hybrid swarms and introgressed population of interspecific hybrids (Oryza rufipogon Griff. x Oryza sativa L.) and its impact on evolution of intermediate types. Euphytica 94, 295–302 (1997).
Suh, H. S., Sato, Y. I. & Morishima, H. Genetic characterization of weedy rice (Oryza sativa L.) based on morpho-physiology, isozymes and RAPD markers. Theor. Appl. Genet. 94, 316–321 (1997).
Akimoto, M., Shimamoto, Y. & Morishima, H. The extinction of genetic resources of Asian wild rice, Oryza rufipogon Griff.: a case study in Thailand. Gen. Res. Crop Evol. 46, 419–425 (1999).
Song, Z., Baorong, L., Zhu, Y. & Chen, J. Pollen competition between cultivated and wild rice species Oryza sativa and O. rufipogon. New Phytol. 153, 289–296 (2002).
Song, Z. P., Lu, B., Zhu, Y. G. & Chen, J. K. Gene flow from cultivated rice to the wild species Oryza rufipogon under experimental field conditions. New Phytol. 157, 657–665 (2003).
Kanazawa, A., Akimoto, M., Morishima, H. & Shimamoto, Y. Inter- and intra-specific distribution of Stowaway transposable elements in AA-genome species of wild rice. Theor. Appl. Genet. 101, 327–335 (2000).
Jenczewski, E., Prosperi, J. M. & Ronfort, J. Differentiation between natural and cultivated populations of Medicago sativa (Leguminosae) from Spain: analysis with random amplified polymorphic DNA (RAPD) markers and comparison to allozymes. Mol. Ecol. 8, 1317–1330 (1999).
Jenczewski, E., Prosperi, J. M. & Ronfort, J. Evidence for gene flow between wild and cultivated Medicago sativa (Leguminosae) based on allozyme markers and quantitative traits. Amer. J. Bot. 86, 677–687 (1999).
Bartsch, D. et al. Impact of gene flow from cultivated beet on genetic diversity of wild sea beet populations. Mol. Ecol. 8, 1733–1741 (1999).
Arnaud, J -F., Viard, F., Delescluse, M & Cuguen, J. Evidence for gene flow via seed dispersal from crop to wild relatives in Beta vulgaris (Chenopodiaceae): consequences for the release of genetically modified crop species with weedy lineages. Proc. R. Soc. Lond. B 270, 1565–1571 (2003).
Seefeldt, S. S., Zemetra, R., Young, F. L. & Jones, S. S. Production of herbicide-resistant jointed goatgrass (Aegilops cylindrica) x wheat (Triticum aestivum) hybrids in the field by natural hybridization. Weed Sci. 46, 632–634 (1998).
Wang, Z., Zemetra, R. S., Hansen, J. & Mallory-Smith, C. A. The fertility of wheat x jointed goatgrass hybrid and its backcross progenies. Weed Sci. 49, 340–345 (2001).
Morrison, L. A., Crémieux, L. C. & Mallory-Smith, C. A. Infestations of jointed goatgrass (Aegilops cylindrica) and its hybrids with wheat in Oregon wheat fields. Weed Sci. 50, 737–747 (2002).
Hansen, L. B., Siegismund, H. R. & Jørgensen, R. B. Introgression between oilseed rape (Brassica napus L.) and its weedy relative B. rapa L. in a natural population. Gen. Res. Crop Evol. 48, 621–627 (2001).
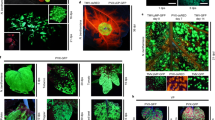
Halfhill, M. D., Richards, H. A., Mabon, S. A. & Stewart, C. N. Expression of GFP and Bt transgenes in Brassica napus and hybridization with Brassica rapa. Theor. Appl. Genet. 103, 659–667 (2001). In this study, an in vivo monitoring system was used to observe introgression and gene expression through the visual marker green fluorescent protein (GFP)
Halfhill, M. D., Millwood, R. J., Weissinger, A. K., Warwick, S. I. & Stewart, C. N. Additive transgene expression and genetic introgression in multiple GFP transgenic crop x weed hybrid generations. Theor. Appl. Genet. (DOI 10.1007/s00122-003-1397-7).
Warwick, S. I. et al. Hybridization between Brassica napus L. and its wild relatives: B. rapa L., Raphanus raphanistrum L., Sinapis arvensis L., and Erucastrum gallicum (Willd.) O. E. Schulz. Theor. Appl. Genet. 107, 528–539 (2003). The first evidence of transgene escape to a wild relative from a commercially released GM crop.
Scott, S. E. & Wilkinson, M. J. Transgene risk is low. Nature 393, 320 (1998).
Linder, C., Taha, I., Seiler, G., Snow, A. & Rieseberg, L. Long-term introgression of crop genes into wild sunflower populations. Theor. Appl. Genet. 96, 339–347 (1998).
Whitton, J., Wolf, D. E., Arias, D. M., Snow, A. A. & Rieseberg, L. H. The persistence of cultivar alleles in wild populations of sunflowers five generations after hybridization. Theor. Appl. Genet. 95, 33–40 (1997).
Rieseberg, L. H., Kim, M. J. & Seiler, G. J. Introgression between the cultivated sunflower and a sympatric wild relative, Helianthus petiolaris (Asteraceae). Int. J. Plant Sci. 160, 102–108 (1999).
Renganayaki, K., Amirthadevaranthinam, A. & Sadasivam, S. Species relationship and hybrid identification in sorghum using RAPD, protein and isozyme techniques. J. Genet. Breed. 54, 117–124 (2000).
Paterson, A. H., Schertz, K. F., Lin, Y. R., Liu, S -C. & Chang, Y -L. The weediness of wild plants: molecular analysis of genes influencing dispersal and persistence of Johnsongrass, Sorghum halepense (L.) Pers. Proc. Natl Acad. Sci. USA 92, 6127–6131 (1995). This paper describes an early approach to the analysis of the genomic basis of weediness.
Darmency, H. The impact of hybrids between genetically modified crop plants and their related species: introgression and weediness. Mol. Ecol. 3, 37–40 (1994).
Warwick, S. I., Beckie, H. J. & Small, E. Transgenic crops: new weed problems for Canada? Phytoprotection 80, 71–84 (1999).
Snow, A. A. & Palma, P. M. Commercialization of transgenic plants: potential ecological risks. Bioscience 47, 86–96 (1997).
Muir, W. M. & Howard, R. D. Fitness components and ecological risk of transgenic release: a model using Japanese medaka (Oryzias latipes) Am. Nat. 158, 1–16 (1999).
Cummings, C. L. et al. Fecundity selection in an experimental sunflower crop–wild system: how well do ecological data predict crop allele persistence. Ecol. Appl. 12, 1661–1671 (2002).
Doebley, J. in Molecular Sytematics of Plants (eds Soltis, P. S., Soltis, D. E. & Doyle, J. J.) 202–222 (Chapman and Hall, New York, 1992).
Burke, J. M., Tang, S., Knapp, S. J. & Rieseberg, L. H. Genetic analysis of sunflower domestication. Genetics 161, 1257–1267 (2002).
Ingram, J. The separation distance required to ensure cross-pollination is below specified limits in non-seed crops of sugar beet, maize and oilseed rape. Plant Var. and Seeds 13, 181–199 (2000).
Morris, W. F., Kareiva, P. M. & Raymer, P. L. Do barren zones and pollen traps reduce gene escape from transgenic crops? Ecol. Appl. 4, 157–165 (1994).
Nordborg, M. et al. The extent of linkage disequilibrium in Arabidopsis thaliana. Nature Genet. 30, 190–193 (2002).
Jiang, C. X. et al. Multilocus interactions restrict gene introgression in interspecific populations of polyploid Gossypium (cotton). Evolution 54, 798–814 (2000).
Halfhill, M. D., Millwood, R. J., Raymer, P. L. & Stewart, C. N. Bt-transgenic oilseed rape hybridization with its weedy relative, Brassica rapa. Environmental Biosafety Res. 1, 19–28 (2002).
Metz, P. L. J., Jacobsen, E., Nap, J. P., Pereira, A. & Stiekema, W. J. The impact on biosafety of the phosphinothricin-tolerance transgene in inter-specific B. rapa x B. napus hybrids and their successive backcrosses. Theor. Appl. Genet. 95, 442–450 (1997).
Gressel, J. Tandem constructs: preventing the rise of superweeds. Trends Biotech. 17, 361–366 (1999). This was the first account to recognize the power of using transgene linkage as a means to prevent or decrease introgression.
Daniell, H., Datta, R., Varma, S., Gray, S. & Lee, S. B. Containment of herbicide resistance through genetic engineering of the chloroplast genome. Nature Biotechnol. 16, 345–348 (1998).
Corriveau, J. P. & Coleman, A. W. Rapid screening method to detect potential biparental inheritance of plastid DNA and results for over 200 angiosperm species. Am. J. Bot. 75, 1443–1458 (1988).
Hagemann, R. in Cell Organelles (ed. Hermann, R. G.) 65–96 (Springer, Berlin, 1992).
Reboud, X. & Zeyl, C. Organelle inheritance in plants. Heredity 72, 132–140 (1994).
Chamberlain, D. & Stewart, C. N. Transplastomics and transgene escape. Nature Biotechnol. 17, 330–331 (1999).
Huang, C. Y., Ayliffe, M. A. & Timmis, J. N. Direct measurement of the transfer rate of chloroplast DNA into the nucleus. Nature 422, 72–76 (2003).
DeBlock, M. & Debrouwer, D. Engineered fertility control in transgenic Brassica napus L.: histochemical analysis of anther development. Planta 189, 218–225 (1993).
Schmuelling, T., Roehrig, H., Pilz, S., Walden, R. & Schell, J. Restoration of fertility by antisense RNA in genetically engineered male sterile tobacco plants. Mol. Gen. Genet. 237, 385–394 (1993).
Keenan, R. J. & Stemmer, W. P. C. Nontransgenic crops from transgenic plants. Nature Biotechnol. 20, 215–216 (2002).
Masood, E. Monsanto set to back down over “terminator” gene? Nature 396, 503 (1998).
Kuvshinov, V., Koivu, K., Kanerva, A. & Pehu, E. Molecular control of transgene escape from genetically modified plants. Plant Sci. 160, 517–522 (2001).
Schernthaner, J. P., Fabijanski, S. F., Arnison, P. G., Racicot, M. & Robert, L. S. Control of seed germination in transgenic plants based on the segregation of a two-component genetic system. Proc. Natl Acad. Sci. USA 100, 6855–6859 (2003).
Heyn, F. W. Analysis of unreduced gametes in the Brassiceae by crosses between species and ploidy levels. Z. Pflanzenzüchtg. 78, 13–30 (1977).
Ramachandran, S., Buntin, G. D., All, J. N., Raymer, P. L. & Stewart, C. N. Intraspecific competition of an insect-resistant transgenic canola in seed mixtures. Agron. J. 92, 368–374 (2000).
Burke, J. M. & Rieseberg, L. H. Fitness effects of transgenic disease resistance in sunflowers. Science 300, 1250 (2003). This is the most comprehensive study, so far, of the consequences of the introgression of a fitness-associated transgene into a wild relative of a crop. This showed that a disease-resistance transgene would not increase the fitness of a wild plant.
Arnold, M. L. Natural hybridization as an evolutionary process. Ann. Rev. Ecol. Syst. 23, 237–261 (1992).
Arnold, M. L. & Bennett, B. D. in Hybrid Zones and the Evolutionary Process (ed. Harrison, R. G.) 115–139 (Oxford Univ. Press, New York, 1993).
Arnold, M. L. Iris nelsonii (Iridaceae): origin and genetic composition of a homoploid hybrid species. Amer. J. Bot. 80, 577–583 (1993).
Bennett, B. D. & Grace, J. B. Shade tolerance and its effect on the segregation of two species of Louisiana Iris and their hybrids. Amer. J. Bot. 77, 100–107 (1990).
Cruzan, M. B. & Arnold, M. L. Assortative mating and natural selection in an Iris hybrid zone. Evolution 48, 1946–1958 (1994).
Saxena, D. & Stotzky, G. Bt corn has a higher lignin content than non-Bt corn. Am. J. Bot. 88, 1704–1706 (2001).
Snow, A. A. et al. A Bt transgene reduces herbivory and enhances fecundity in wild sunflowers. Ecol. Appl. 13, 279–286 (2003).
Hoag, H. Tougher rules aim to prevent gene flow into crops. Nature 422, 103 (2003).
United States Department of Agriculture. USDA announces actions regarding Plant Protection Act violations involving Prodigene Inc. News Release No. 4098.02 [online], (cited 05 Sep. 2003), <http://www.usde.gov/news/releases/2002/12/0498.htm> (2002).
N, U. Genomic-analysis in Brassica with special reference to the experimental formation of B. napus and peculiar mode of fertilization. Jpn. J. Bot. 7, 389–452 (1935). This impressive deductive investigation of the polyploid genomic relationships among Brassicas was made long before the development of molecular tools. The author used the word 'genomic' 60 years before the modern genomics revolution and his conclusions have stood the test of time.
Aldrich, P. R. & Doebley, J. Restriction fragment length variation in the nuclear and chloroplast genomes of cultivated and wild Sorghum bicolor. Theor. Appl. Genet. 85, 293–302 (1992).
Gómez, M. I. et al. Tetraploid nature of Sorghum bicolor (L.) Moench. J. Heredity 89, 188–190 (1998).
Santoni, S. & Berville, A. Evidence for gene exchanges between sugar beet (Beta vulgaris L.) and wild beets: consequences for transgenic sugar beets. Plant Mol. Biol. 20, 578–580 (1992).
Desplanque, B. et al. Genetic diversity and gene flow between wild, cultivated and weedy forms of Beta vulgaris L. (Chenopodiaceae) assessed by RFLP and microsatellite markers. Theor. Appl. Genet. 98, 1194–1201 (1999).
Acknowledgements
We are grateful for support from the United States Department of Agriculture Boitechnology Risk Assessment Grants Program and the University of Tenessee Institute of Agriculture.
Author information
Authors and Affiliations
Corresponding author
Glossary
- INTROGRESSION
-
The permanent incorporation of genes from one set of differentiated populations (species, subspecies, races and so on) into another.
- LANDRACES
-
A crop cultivar that evolved with and has been genetically improved by traditional agriculturalists, but has not been directly influenced by modern breeding practices.
- F1 HYBRIDIZATION
-
The initial cross between parent plants of different varieties, subspecies, species or genera.
- BACKCROSSES
-
The mating of an individual with its parent, or with an individual of the same genotype as its parent, to follow the inheritance of alleles and phenotypes.
- GENE FLOW
-
The dispersal of genes, in both gametes and zygotes, in and between breeding populations.
- BC1 AND BC2 HYBRID
-
The offspring of a cross between a hybrid and one of the recurrent parent species or varieties. The subscript number represents the number of generations that have been crossed in this fashion.
- HYBRID ZONES
-
(Hybrid swarms). Areas in which hybrid plants backcross to the parents and cross with themselves, so there is a continuous intergradation of forms in the population.
- ISOZYME
-
Different forms of the same enzyme (synonymous with allozymes), which were used as some of the first biochemically-based genetic markers.
- ALLOPATRIC
-
Occurring in geographically separate areas.
- LINKAGE DISEQUILIBRIUM
-
(LD). A statistical measure of the non-independence of alleles. Departure from the predicted frequencies of multiple locus gamete types, assuming that all alleles are randomly associated.
- CLINE
-
A variational trend in space that is found in a poulation, or a series of populations, of a species.
- GENETIC LINKAGE MAP
-
A linear map of the relative positions of genes along a chromosome. Distances are established by linkage analysis, which determines the frequency at which two gene loci become separated during chromosomal recombination.
- ELITE CROP VARIETIES
-
Agronomically desirable crop varieties that are widely used, adapted to local environments, perform well under intensive agricultural practices and are typically the product of intensive breeding.
- SEED SHATTERING
-
Seeds dispersing from their fruits before harvest.
- EXOTIC CROP VARIETIES
-
A variety that is from outside a breeding region or has traits that are uncommon to the prevalent crop variety.
- FITNESS
-
The potential evolutionary success of a genotype, which is defined as the reproductive success or the proportion of genes that an individual leaves in the gene pool of a population. The individuals with the greatest fitness leave the largest numbers of offspring.
- WIDE CROSSES
-
Hybridization between differentiated taxa.
- MIGRATIONAL MELTDOWN
-
The theoretical state of gene flow that leads to the fixation of a 'bad' gene and the subsequent reduction of population size.
- ECOTYPE
-
A genetic variety of a single species that is adapted for local ecological conditions.
- HOMEOLOGOUS CHROMOSOME
-
A partially homologous chromosome, which usually indicates some original ancestral homology.
- POLYPLOID
-
The genomic state of having three or more sets of homologous chromosomes (for example, tetraploid organisms, which have four sets of chromosomes).
- FUNCTIONAL TRANSLATIONAL FUSION
-
The in-frame chimaera of two or more genes that gives rise to a single chimeric protein.
- GENE USE-RESTRICTION TECHNOLOGY
-
(GURT). A biotechnological tool that controls embryo viability whereby the addition of a recoverable blocking sequence prevents some essential physiological function in a host plant and can be inducibly removed to recover viability.
Rights and permissions
About this article
Cite this article
Stewart, C., Halfhill, M. & Warwick, S. Transgene introgression from genetically modified crops to their wild relatives. Nat Rev Genet 4, 806–817 (2003). https://doi.org/10.1038/nrg1179
Issue Date:
DOI: https://doi.org/10.1038/nrg1179
This article is cited by
-
Agronomic and phenotypic plant traits as indicators for environmental risks of genetically modified plants
Environmental Sciences Europe (2024)
-
Unravelling the genetic potential of untapped crop wild genetic resources for crop improvement
Conservation Genetics Resources (2022)
-
Genome-edited Camelina sativa with a unique fatty acid content and its potential impact on ecosystems
Environmental Sciences Europe (2021)
-
Incorporating the field border effect to reduce the predicted uncertainty of pollen dispersal model in Asia
Scientific Reports (2021)
-
Risk assessment of genetically engineered plants that can persist and propagate in the environment
Environmental Sciences Europe (2020)