Key Points
-
Despite the recommendations of many expert groups, public health surveillance systems have not yet improved to the point where emerging infectious threats can be better anticipated. The Ebola and Zika epidemics are the latest to demonstrate that pathogens often spread undetected for some time before being diagnosed in a population.
-
Next-generation sequencing, particularly the use of portable genomic sequencers, offers an intriguing solution to the diagnosis and surveillance problems — it enables rapid in situ diagnostics through amplicon-based or metagenomics approaches and creates a stream of genomic data that can reveal critical epidemiological aspects of an outbreak or epidemic's dynamics.
-
Genomic epidemiology for rapid outbreak response has demonstrated some early successes in Ebola and Zika, but there are a number of challenges to overcome — some technical and some cultural. Data sharing is one of these, but other ethical and legal issues must be considered.
-
The power of a genomic epidemiology approach could be extended by incorporating concepts from digital disease detection and One Health. By coupling sequencing to an enhanced surveillance and response platform, we could take a more anticipatory approach to outbreak prevention and control.
Abstract
The recent Ebola and Zika epidemics demonstrate the need for the continuous surveillance, rapid diagnosis and real-time tracking of emerging infectious diseases. Fast, affordable sequencing of pathogen genomes — now a staple of the public health microbiology laboratory in well-resourced settings — can affect each of these areas. Coupling genomic diagnostics and epidemiology to innovative digital disease detection platforms raises the possibility of an open, global, digital pathogen surveillance system. When informed by a One Health approach, in which human, animal and environmental health are considered together, such a genomics-based system has profound potential to improve public health in settings lacking robust laboratory capacity.
Similar content being viewed by others
Main
Everything that happens twice will surely happen a third time
Paulo Coelho — The Alchemist
In late 2013 and early 2014, a lethal haemorrhagic fever spread throughout forested Guinea (Guinée forestière), undiagnosed for months. By the time it was reported to be Ebola, the virus had spread to three countries1 and was likely past the point at which case-level control measures, such as isolation and infection control, could have contained the nascent outbreak. In 2015, a new dengue-like illness was implicated in a dramatic increase in Brazil's microcephaly cases; one year later, analyses revealed that the Zika virus had been sweeping through the Americas, unnoticed by existing surveillance systems, since late 2013 (Refs 2,3,4).
Although public health surveillance systems have evolved to meet the changing needs of our global population, we continue to dramatically underestimate our vulnerability to pathogens, both old and new5. Indeed, the recent events in West Africa and Brazil highlight the gaps in existing infectious disease surveillance systems, particularly when dealing with novel pathogens or pathogens whose geographic range has extended into a new region. Despite the lessons learned from previous outbreaks6, such as the severe acute respiratory syndrome (SARS) epidemic in 2002–2003 and the 2009 influenza pandemic — particularly the need for enhanced national surveillance and diagnostic capacity — infectious threats continue to surprise and sometimes overwhelm the global health response.
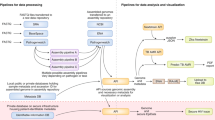
The cost of these epidemics demands that we take action: with fewer than 30,000 cases, the Ebola outbreak ultimately resulted in over 11,000 deaths, left nearly 10,000 children without parents7 and caused cumulative gross domestic product losses of more than 10%8. As with prior crises, in the wake of Ebola, multiple commissions have offered suggestions for essential reforms8,9. Most focus on systems-level change, such as funding research and development or creating a centralized pandemic preparedness and response agency. However, they also call for enhanced molecular diagnostic and surveillance capacity coupled to data-sharing frameworks. This hints at an emerging paradigm for rapid outbreak response, one that employs new tools for pathogen genome sequencing and epidemiological analysis (Fig. 1) and that can be deployed anywhere. In this model, portable, in-country genomic diagnostics are targeted to key settings for routine human, animal and environmental surveillance or rapidly deployed to a setting with a nascent outbreak. Within our increasingly digital landscape, wherein a clinical sample can be transformed into a stream of data for rapid analysis and dissemination in a matter of hours, we face a tremendous opportunity to more proactively respond to disease events. However, the potential benefits of such a system are not guaranteed, and many obstacles remain.
Portable genome sequencing technology and digital epidemiology platforms form the foundation for both real-time pathogen and disease surveillance systems and outbreak response efforts, all of which exist within the One Health context, in which surveillance, outbreak detection and response span the human, animal and environmental health domains.
Here, we review recent advances in genomics-informed outbreak response, including the role of real-time sequencing in both diagnostics and epidemiology. We outline the opportunities for integrating sequencing with the One Health and digital epidemiology fields, and we examine the ethical, legal and social issues that must be addressed if we are to move towards an era of genomics-informed pathogen surveillance.
Genomics in rapid-response diagnostics
Next-generation sequencing (NGS) platforms have recently moved from proof-of-concept studies to routine use in the clinical microbiology laboratory10. Most NGS services rely on bench-top instruments and sequencing from culture. However, when trying to proactively detect emerging infections or in many rapid outbreak responses, the aetiological agent behind a cluster is often unknown. Even if the agent is known, both the limited culture capacity in a field laboratory and the need for diagnostic turnaround times in hours, not days, preclude sequencing from culture. Sequencing directly from a sample using a portable sequencing platform is therefore more relevant in the field. Similarly, the need for a sequencer that can withstand being shipped and operated under rough field conditions, coupled with the need for rapid turnaround, make small, portable sequencers an attractive option.
Clinical metagenomics. With its untargeted approach to sequencing, clinical metagenomics can cross disciplines in a way that clinical microbiology struggles to — identifying viral, bacterial, fungal and other eukaryotic pathogens in a single assay11 and coupling pathogen detection to pathogen discovery. Given the current high cost of the technique — conservatively estimated at several thousand dollars — it is most often used when dealing with potentially lethal infections that fail the conventional diagnostic paradigm, such as the recent diagnosis of an unusual case of meningoencephalitis caused by the amoeboid parasite Balamuthia mandrillaris12 or the diagnosis and treatment of neuroleptospirosis in a critically unwell teenager13. In the latter case, despite a high index of suspicion for infection, Leptospira santarosai was not detected by culture or PCR, as the diagnostic primer sequences were eventually found to be a poor match to the genome of the pathogen. Intravenous antibiotic therapy resulted in rapid recovery. In such an example, the costs are easily justified, particularly when offset against the cost of a stay in an intensive treatment unit. However, routine diagnostic metagenomics is currently limited to a handful of clinical research laboratories worldwide; it is therefore regarded as a 'test of last resort' and kept in reserve for vexing diagnostic conundrums.
Substantial practical challenges hinder the adoption of metagenomics for diagnostics (Fig. 2) (reviewed in depth in Ref. 11). Chief among these is analytic sensitivity, which depends on pathogen factors (for example, genome size, ease of lysis and life cycle); analytic factors (for example, the completeness of reference databases and the potential to mistake a target for a close genetic relative); and sample factors (for example, pathogen abundance within a sample and contaminating background DNA). As an example of a problematic sample, during Zika surveillance, attempts to perform untargeted metagenomics sequencing on blood yielded few, or in some cases zero, reads owing to low viral titres14. Target-enrichment technologies (reviewed in Ref. 15) such as bait probes can be employed, but even these were unsuccessful at recovering whole Zika genomes, necessitating PCR enrichment14. In addition to sensitivity, universal pathogen detection through clinical metagenomics is complicated by specificity issues arising from misclassification or contaminated reagents, the challenge of reproducing results from a complex clinical workflow, nucleic acid stability under varying assay conditions, ever-changing bioinformatics workflows and cost.
A mobile medical unit deploying a portable clinical metagenomics platform has been established at the epicentre of an infectious disease outbreak, but the team faces challenges throughout the diagnostic process and epidemiological response. For example, in the case of Zika virus, samples, such as blood, with low viral titres, a small genome of <11 kb and transient viraemia120 combine to complicate detection of viral nucleic acid by use of a strictly metagenomic approach. Furthermore, obtaining a sufficient amount of viral nucleic acids for genome sequencing beyond simple diagnostics requires a tiling PCR and amplicon sequencing approach14. Other challenges include, for example, access to a reliable Internet connection, the ability to collect sample metadata and translating genomic findings into real-time, actionable recommendations.
Given these issues, could metagenomics replace conventional microbiological and molecular tests for infection? Recent studies have used metagenomics in common presentations, including sepsis16, pneumonia17, urinary tract infections18 and eye infections19. These have generally yielded promising results, albeit typically at a lower sensitivity than conventional tests and at a much greater cost. Despite these problems, two factors will drive sequencing to eventually become routine clinical practice. First, the ever-decreasing cost of sequencing coupled with the potential for cost savings achieved by using a single diagnostic modality versus tens or hundreds of different diagnostic assays — each potentially requiring specific instrumentation, reagents, validation and labour — is attractive from a laboratory operations perspective. Second, and perhaps most compelling, is the additional information afforded by genomics, including the ability to predict virulence or drug resistance phenotypes, the ability to detect polymicrobial infections and phylogenetic reconstruction for outbreak analysis.
Novel technologies: portable sequencing. Given that outbreaks of emerging infectious diseases (EIDs) most often occur in settings with minimal laboratory capacity, where routine culture and bench-top sequencing are simply not feasible, the need for a portable diagnostic platform capable of in situ clinical metagenomics and outbreak surveillance is evident. A trend towards smaller and less expensive bench-top sequencing instruments was seen with the 454 Genome Sequencer Junior system (which has since been discontinued), the Ion Torrent Personal Genome Machine (PGM) system and the Illumina MiSeq system, which were released in close succession20. Each of these instruments costs <$150,000 and puts NGS capability into the hands of smaller laboratories, including clinical settings. In 2014, the MinION from Oxford Nanopore Technologies was released to early access users21, heralding the potential for highly portable 'lab-in-a-suitcase' sequencing. The MinION is pocket-sized and is controlled and powered through a laptop USB connection. It is provided under a model whereby the hardware is free but the consumer pays a premium for the reagent and flow cell consumables. Compared with bench-top instruments, the absence of a rolling service contract or regular engineer visits makes it theoretically possible to scale this platform out to potentially unlimited numbers of laboratories. Importantly, the MinION has been used in field situations, including in diagnostic tent laboratories during the Ebola epidemic22,23 and in a roving bus-based mobile laboratory in Brazil as part of the ZiBRA project3,24. Others have taken the MinION to more extreme environments where even the smallest traditional bench-top sequencer could not go, including the Arctic25 and Antarctic26, a deep mine27 and zero gravity aboard the reduced-gravity aircraft (nicknamed the 'Vomit Comet')28 and the International Space Station29.
However, this technology is not yet a panacea; remaining challenges include high DNA or RNA input requirements (currently hundreds of nanograms), which often necessitate PCR-based amplification approaches; a flow cell cost of $500, keeping the cost per sample high despite multiplexing approaches; and high error rates, which require that genomes are sequenced to high coverage for single nucleotide polymorphism-based analysis and analysed at the signal level. Moreover, although the long reads produced by the MinION overcome a number of challenges in assembling eukaryotic microbial pathogen genomes, such as the presence of discrete chromosomes or long repetitive regions, the upstream nucleic acid extraction steps required to obtain genomic DNA vary across microbial domains and might necessitate reagents and equipment far less portable than the MinION.
Genomic epidemiology
From transmission to epidemic dynamics. Genomics is capable of informing not just pathogen diagnostics but also epidemiology. Pathogen sequencing has been used for decades to understand transmission in viral outbreaks, from early studies of hantavirus in the United States of America30 to human immunodeficiency virus (HIV) in the United Kingdom31; more recently, the approach has been successfully extended to include bacterial pathogens (reviewed in Ref. 32) and has come to be known as genomic epidemiology, a term encompassing everything from population dynamics to the reconstruction of individual transmission events within outbreaks32. Most transmission-focused investigations to date have been retrospective, with only a subset unfolding in real time, as cases are diagnosed33,34,35,36,37.
In transmission-focused investigations, genetic variants are used to identify person-to-person transmission events (Fig. 3), either through manual interpretation of the variants shared between outbreak cases38 or via model-based approaches39, with the result being a transmission network. Epidemic investigations are very different — only a subset of the epidemic cases are sequenced. Thus, the goal is to use the population structure of the pathogen to understand the overall dynamics of the epidemic. Here, phylodynamic approaches are used to infer epidemiological parameters of interest.
Genomic approaches to identifying transmission events typically involve four steps. In the first step, outbreak isolates, and often non-outbreak control isolates, are sequenced and their genomes either assembled de novo or mapped against a reference genome. Next, the genomic differences between the sequences are identified — depending on the pathogen and the scale of the outbreak, these may include features such as genetic variants, insertions and deletions or the presence or absence of specific genes or mobile genetic elements. In the third step, these features are examined to infer the relationships between the isolates from whence they came — a variant common to a subset of isolates, for example, suggests that those cases are epidemiologically linked. Finally, the genomic evidence for epidemiological linkages is reviewed in the context of known epidemiological information, such as social contact between two cases or a common location or other exposure. Recently, automated methods for inferring potential epidemiological linkages from genomic data alone have been developed, greatly facilitating large-scale genomic epidemiological investigations121.
First conceptualized in 2004 by Grenfell et al. as a union of “immunodynamics, epidemiology, and evolutionary biology” (Ref. 40), phylodynamics captures both epidemiological and evolutionary information from measurably evolving pathogens — those viruses and bacteria for which high mutation rates and/or a range of sampling dates contribute to a meaningful amount of genetic variation between sequences41,42 — in other words, enough genetic diversity to be able to infer an evolutionary history for a pathogen of interest, even if that history is only over the short time frame of an outbreak or epidemic. This is possible for most pathogens, particularly single-stranded DNA viruses, RNA viruses and many bacterial species42,43, but there are certain species for which the lack of a strict molecular clock and/or frequent recombination complicate both phylodynamics studies and attempts to infer transmission events42.
Phylodynamics relies on tools such as Bayesian evolutionary analysis sampling trees (BEAST)44, in which sequence data are used to build a time-labelled phylogenetic tree using a specific evolutionary process as a guide — often variations on a theme of coalescent theory45. From the tree, one can infer epidemiological parameters, including the basic reproductive number R0 (Ref. 46). While the insights that can be gained from genomic data alone are exciting, the utility of phylodynamic approaches is greatly extended when additional data are integrated into the models (reviewed in Ref. 47).
Genomic epidemiology in action: Ebola. The many genomic epidemiology studies from the Ebola outbreak (reviewed in Ref. 48) used bench-top and portable sequencing platforms to reveal outbreak-level events and epidemic-level trends. Real-time analyses published around the peak of the epidemic suggested the following: the outbreak probably arose from a single introduction into humans and not repeated zoonotic introductions49,50; sexual transmission had a previously unrecognized role in maintaining transmission chains51; and survivor transmission — another unrecognized phenomenon — contributed to disease flare-ups later in the outbreak52. The first sequencing efforts, all of which had an effect on the epidemiological response in real time, unfolded months into the epidemic. Had they been deployed earlier, we can only speculate as to their potential impact. Arguably, the most compelling use of early sequencing would have been to provide a definitive Ebola diagnosis in this previously unaffected region of West Africa. However, even after the outbreak was underway, sequencing could have benefited the public health response. For example, ruling out bush meat as a source of repeated viral introductions could have changed public health messaging campaigns from avoiding bush meat to the importance of hygiene and safe funeral practices53, potentially averting some cases. Portable sequencing and phylodynamic approaches are currently being deployed in the ongoing Zika epidemic; whether the real-time reporting of genomic findings is able to alter the course of a vector-borne epidemic remains to be seen.
Retrospective phylodynamic investigations are also useful for pandemic preparedness planning. A recent analysis of 1,610 Ebola virus genomes — approximately 5% of all cases — reconstructs the movement of the virus across West Africa and reveals drivers for its spread1. The authors deduce that Ebola importation was more likely to occur between regions of a country than across international borders and that both population size and distance to a nearby large urban centre were associated with local expansion of the virus. These findings may affect decision-making around border closures in future Ebola outbreaks and point to the need to develop surveillance, diagnostic and treatment capacity in urban centres.
The role of the environment
In deploying genomics for surveillance, diagnostics and epidemiological investigation, a key question remains: where? Many regions lack the diagnostic laboratory capacity to carry out basic surveillance, but continuous genomic surveillance in all of these settings would be impossible. Numerous projects have attempted to describe the pool of geographic hot spots and candidate pathogens from which the next epidemic or pandemic will arise. Determining these factors is key to predicting and preventing spillover events (Fig. 4), but huge gaps in our understanding of disease ecology remain. Woolhouse et al. describe 1,399 human pathogens, of which 87 — mostly viral — have emerged since 1980 (Ref. 54). Jones et al. extend this to include 335 new EIDs since 1940 (Ref. 55). They report an increasing number of events each decade, generally located in hot spots defined by specific environmental, ecological and socio-economic characteristics.
In spillover, a pathogen previously restricted to animals gradually begins to move into the human population. During stage one (pre-emergence), as a result of changing demographics and/or land use, a pathogen undergoes a population expansion, extends its host range or moves into a new geographic region. During stage two (localized emergence), contact with animals or animal products results in spillover of the pathogen from its natural reservoir(s) into humans but with little to no onward person-to-person transmission. During stage three (pandemic emergence), the pathogen is able to sustain long transmission chains, that is, a series of disease transmission events, such as a sequential series of person-to-person transmissions, and its movement across borders is facilitated by human travel patterns65.
Most EIDs are zoonotic in origin, with the highest risk of spillover in regions with high wildlife diversity that have experienced recent demographic change and/or recent increases in farming activity55. A global biogeographic analysis of human infectious disease further supports the use of biodiversity as a proxy for EID hot spots56, and reviews focused on systems-level, rather than ecological, factors identify the breakdown of local public health systems as drivers of outbreaks, suggesting that surveillance ought to be targeted to settings where biodiversity and changing demographics meet inadequate sanitation and hygiene, lack of a public health infrastructure for delivering interventions and no or limited resources for control of zoonoses and vector-borne diseases57.
These analyses provide a shortlist of regions, including parts of eastern and southeastern Asia, India and equatorial Africa, on which genomic and other surveillance activities should be focused55,58. Within these regions, sewer systems and wastewater treatment plants could be important foci for sample collection, providing a single point of entry to biological readouts from an entire community. Indeed, proof-of-concept metagenomics studies have revealed the presence of antibiotic resistance genes59, human-specific viruses60 and other pathogens of interest in this readily accessible sample type. Other recent surveys offer insight into what such systems might need to look for. In 2013, Rosenberg et al. reported that viruses dominate the list of agents newly recognized to cause disease in humans61. Most were zoonotic in origin, and over one-quarter had been detected in non-human species many years before being identified as human pathogens. A later review reiterates this observation, noting that recent agents of concern — Ebola, Zika and chikungunya — had been identified decades before they achieved pandemic magnitude62. As a result of NGS technology, the pace of novel virus discovery is accelerating, with recent large-scale studies revealing 184 new viruses sampled from macaque faeces in a single geographic location63 and 1,445 new viruses discovered from RNA transcriptomic analyses of multiple invertebrate species64. However, understanding which of these new entities might pose a threat requires a new approach.
One Health. The emergence of a zoonotic pathogen proceeds in stages65 (Fig. 4); in an effort to better anticipate these transitions and more proactively respond to emerging threats, the One Health movement was launched in 2004. Recognizing that human, domestic animal and wildlife health and disease are linked to each other and that changing land-use patterns contribute to disease spread, One Health aims to develop systems-minded, forward-thinking approaches to disease surveillance, control and prevention66. By investing in infrastructure for human and animal health surveillance, committing to timely information sharing and establishing collaborations across multiple sectors and disciplines, the goal of the One Health community is an integrated system incorporating human, animal and environmental surveillance — a goal in which genomics can have an important role.
The One Health approach has been implemented through the PREDICT project, which is part of the Emerging Pandemic Threats (EPT) programme of the US Agency for International Development (USAID). PREDICT explores the spillover of selected viral zoonoses from particular wildlife taxa67, and early efforts have focused on developing non-invasive sampling techniques for wildlife68, estimating the breadth of mammalian viral diversity across nine viral families and at least 320,000 undiscovered species69 and demonstrating that viral community diversity is at least a partially deterministic process, suggesting that forecasting community changes, which potentially signal spillover, is a possibility63. Although the goal of using integrated surveillance information to predict an outbreak is still many years away, One Health studies are already leveraging the tools and techniques of genomic epidemiology to understand current outbreaks.
Combining genomic data with data streams from enhanced One Health surveillance platforms presents an opportunity to detect the population expansions and/or cross-species transmissions that may precede a human health event. For example, genome sequences from a raccoon-associated variant of rabies virus (RRV), when paired with fine-scale geographic information and data from Canadian and US wildlife rabies vaccination programmes, demonstrated that multiple cross-border incursions were responsible for the expansion of RRV into Canada and sustained outbreaks in several provinces70; this finding led to renewed concern about and action against rabies on the part of public health authorities71. One of the first studies coupling detailed wildlife and livestock movement data with phylodynamic analysis of a bacterial pathogen revealed that cross-species jumps from an elk reservoir were the source of increasing rates of Brucella abortus infections in nearby livestock72; as the most common zoonosis of humans, brucellosis control programmes will benefit substantially from this sort of One Health approach73.
This model, in which diagnostic testing in reference laboratories triggers genomic follow-up, represents an effective near-term solution for integrating genomics into One Health surveillance efforts as the community explores solutions to the many challenges facing in situ clinical metagenomics surveillance of animal populations (reviewed in Ref. 74). Initial forays into this area have been successful; for example, metagenomics analysis of human diarrhoeal specimens and stools from nearby pigs revealed potential zoonotic transmission of rotavirus75. However, metagenomic sequencing across a range of animal species and environments yields more questions than answers. What is an early signal of pathogen emergence versus background microbial noise65? Which emerging agents are capable of crossing the species barrier and causing human disease74? What degree of sampling is required to capture potential spillovers67? Ultimately, a more efficient use of metagenomics in a One Health surveillance strategy might be scanning for zoonotic 'jumps' in selected sentinel human populations rather than a sweeping animal surveillance strategy62, with sentinels chosen according to EID hotspot maps and other factors65 and interesting genomic signals triggering follow-up sequencing in the relevant animal reservoirs. By combining genomic data generated through these targeted surveillance efforts with phylodynamic approaches, it will be possible to take simple presence or absence signals and derive useful epidemiological insights: signals of population expansion; evidence of transmission within and between animal reservoirs and humans; and epidemiological analysis of a pathogen's early expansion.
Digital epidemiology
Most modern surveillance systems use human, animal, environmental and other data76 to carry out disease-specific surveillance, in which a single disease is monitored through one or more data streams, such as positive laboratory test results or reportable communicable disease notifications. Despite marked advances over the preceding decades, testimony from multiple expert groups has repeatedly emphasized the need for improved surveillance capacity8,77, including the use of syndromic surveillance, a more pathogen-agnostic approach aimed at early detection of emerging disease78,79. Syndromic surveillance systems might leverage unique data streams such as school or employee absenteeism, grocery store or pharmacy purchases of specific items or calls to a nursing hotline as signals of illness in a population. Increasingly, digital streams are being used as an input to these systems, be they participatory epidemiology projects such as Flu Near You80, the automated analysis of trending words or phrases on social media sites, such as Twitter81,82, or Internet search queries83,84,85.
This new approach to surveillance is known as digital epidemiology and is also referred to as digital disease detection86. In digital epidemiology, information is first retrieved from a range of sources, including digital media, newswires, official reports and crowdsourcing; second, translated and processed, which includes extracting disease events and ensuring reports are not duplicated; third, analysed for trends; and fourth, disseminated to the community through media, including websites, email lists and mobile alerts87. At least 50 digital epidemiology platforms are currently operating88, and their flexible nature and cost-effective, real-time reporting make them effective tools for gathering epidemic intelligence, particularly in settings lacking traditional disease surveillance systems.
Modelling drivers of infectious-disease emergence. The fields of One Health and digital epidemiology are increasingly overlapping. In the PREDICT consortium, the HealthMap system89 and local media surveillance were combined to identify 307 health events in five countries over a 16-week period90. PREDICT also suggested a role for digital epidemiology in not just event detection but also the identification of changing EID drivers. EIDs are driven by multiple factors, many of which have digital outputs and represent novel sources of surveillance data91. For example, human movement can be revealed by mobile phone data or by the patterns of lighted cities at night, hunting data collected by states can reveal interactions between humans and wildlife, and social media and digital news sources can reveal early signals of famine, war and other social unrest. A major challenge is that the number of digital data sets available for each driver varies substantially, from hundreds for surveying land use changes — many based on remote sensing data92 — to mere handfuls around social inequalities and human susceptibility to infection, with most data biased towards North America and Europe.
The digital and genomic epidemiology domains are also starting to overlap. In the Ebola outbreak, digital epidemiology revealed that drivers of infection risk included settings where households lacked a radio, with high rainfall and with urban land cover93, echoing the evidence from a genomic study suggesting that sites at which urban and rural populations mix contribute to disease1. During the Zika epidemic, Majumder et al. used HealthMap and Google Trends to estimate the basic reproductive number R0 to be 1.42–3.8394; phylodynamic estimates from Brazilian genomic data gave similar ranges (1.29–3.85)3, indicating that both types of data streams can be leveraged in calculating epidemiological parameters that help shape the public health response.
A digital pathogen surveillance era
Recent reports have called for the integration of genomic data with digital epidemiology streams92,95. When informed by a One Health approach, the epidemiological potential of this digital pathogen surveillance system is profound. Imagine parallel networks of portable pathogen sequencers deployed to laboratories and communities in EID hot spots — regions that are traditionally underserved with respect to laboratory and surveillance capacity — and processing samples collected from targeted sentinel wildlife species, insect vectors and humans (Fig. 5). Samples would be pooled for routine surveillance — either through targeted diagnostics or, if the issue of analytical sensitivity can be overcome, through metagenomics — with a full genomic work-up of individual samples should a pathogenic signal be detected. At the same time, existing Internet-based platforms such as HealthMap and new local participatory epidemiology efforts would be collecting data to both identify potential hotspot regions and detect EID events, enabling both prospective and rapid-response deployment of additional sequencers. Genome sequencing data coupled with rich metadata would then be released in real time to web-based platforms, such as Virological for collaborative analysis and Nextstrain for analysis and visualization96. These sites — already used in the Ebola and Zika responses — would act as the nexus for a global network of interested parties contributing to real-time phylodynamic and epidemiological analyses and looking for signals of spillover, pathogen population expansion and sustained human-to-human transmission. Results would be immediately shared with the One Health frontline — epidemiologists, veterinarians and community health workers — who would then implement evidence-based interventions to mitigate further spread.
It is 2027, and our planet's changing climate and land-use patterns have meant that new emerging infectious diseases (EIDs) are spilling over into humans from wildlife reservoirs with increasing frequency. Building off EID hotspot maps developed in 2008 (Ref. 55), a global public health consortium has implemented an online surveillance tool that scans the digital output of citizens, news organizations and governments in those regions, including data from local retailers on key health-related products, such as tissues and over-the-counter cold remedies. In one such region, the syndromic surveillance system reports higher-than-average sales of a common medication used to relieve fever. Spatial analysis of the data from the pharmacies in the region suggests that the trend is unique to a particular district; a follow-up geographic information system (GIS) analysis using satellite data reveals that this area borders a forest and is increasingly being used for the commercial production of bat guano. An alert is triggered, and the field response team meets with citizens in the area. Nasopharyngeal swabs are taken from humans and livestock with fever as well as from guano and bat tissue collected in the area. The samples are immediately analysed using a portable DNA sequencer coupled to a smartphone. An app on the phone reports the clinical metagenomic results in real time, revealing that in many of the ill humans and animals, a novel coronavirus makes up the bulk of the microbial nucleic acid fraction. The sequencing data are immediately uploaded to a public repository as they are generated, tagged with metadata about the host, sample type and location and stored according to a pathogen surveillance ontology. The data release triggers an announcement via social media of a novel sequence, and within minutes, interested virologists have created a shared online workspace and open lab notebook to collect their analyses of the new pathogen.
The pathway to such a reality is not without its roadblocks. Apart from technical and implementation challenges, a series of larger concerns surrounds the rollout of genomics-based rapid outbreak response, ranging from the uptake of a new, disruptive technology to effecting systems-level change on a global scale.
Ethical, legal and social issues. Sequencing-based diagnostics, particularly clinical metagenomics approaches, are still straddling the boundary between research and clinical use. In this realm, uncertainty is a certainty, be it uncertainty inherent to the technology itself or informational uncertainty, such as how accurate, complete and reliable results actually are97. Early adopters of genomics in the academic domain are used to uncertainty, often acknowledging and appraising it, but routine clinical use requires meeting the evidentiary thresholds mandated by a range of stakeholders, from regulators to the laboratories implementing new sequencing-based tests. Decision criteria that influence whether a new genomic test is adopted include the ability of the assay to differentiate pathogens from commensals, the correlation of pathogen presence with disease, the sensitivity and specificity of the test, its reproducibility and robustness across sample types and settings and a cost comparable to that of existing platforms98.
Validation — defining the conditions needed to obtain reliable results from an assay, evaluating the performance of the assay under said conditions and specifying how the results should be interpreted, including outlining limitations99 — is also critical. Much can be learned from the domain of microbial forensics, where sequencing is playing a large part100. Budowle et al. review validation considerations for NGS101, noting that this technology requires validating sample preparation protocols, including extraction, enrichment and library preparation steps, sequencing protocols, and downstream bioinformatics analyses, including alignment and assembly, variant calling, the underlying reference databases and software tools and the interpretation of the data. Complete validation of a sequencing assay may not always be possible, particularly for emerging pathogens. Therefore, just as the West African Ebola virus outbreak triggered a review of the ethical context for trialling new therapeutics and vaccines102, the scale-up of NGS in emerging epidemics will engender similar conversations. Rather than wait for this to happen, an anticipatory approach is best, outlining the exceptional circumstances under which unvalidated approaches might be used, selecting the appropriate approach and examining the benefits of a potentially untested approach in light of individual and societal interests.
If the social landscape surrounding the introduction of a new technology is not considered, prior experience suggests that the road to implementation will be difficult, with hurdles ranging from public mistrust to moratoria on research103. The enthusiasm of the scientific community for new technology must not lead to inflated claims of clinical utility and poor downstream decisions around the deployment of that technology. Howard et al. outline several principles for successfully integrating genomics into the public health system, and as we pilot digital pathogen surveillance, the community would do well to keep many of them in mind: ensuring that the instruments and processes used are reliable and that reporting is standardized and readily interpretable by end users; that the technology is used to address important health problems; that the advantages of the approach outweigh the disadvantages; and that economic evaluation suggests savings to the health care system and society104. It is also important to reconsider the role of the diagnostic reference laboratory in the new genomic landscape. As their mandates expand to include enhanced surveillance and closer collaboration with field epidemiologists, laboratory directors will face new challenges, from managing exploratory work alongside routine clinical care to hiring a new sort of technologist, one with basic genomics and epidemiology training.
The ethical, social and legal implications of digital pathogen surveillance are an emerging area of research (reviewed in Ref. 105). Chief among the issues that Geller et al. identify is the tension that exists when a new technology has the power to identify a problem but there is limited or no capacity to address the issue. Balancing the benefits and harms to both individuals and populations is challenging when the predictive insight offered by a genomic technology is variable — for example, using genomics to identify an individual as a 'super spreader' has important implications for quarantine and isolation, but that label may be predicated on a tenuous prediction. The problem is further compounded by the fact that many infectious disease diagnoses carry with them a certain amount of stigma and that an individual's right to privacy might be superseded by the need to protect the larger population105.
Data sharing and integration. A critical need for successful digital pathogen surveillance is the capacity for rapid, barrier-free data sharing, and arguments for such sharing are frequently rehashed after outbreaks and epidemics. Genomic epidemiology was born largely in the academic sphere, with early papers coming from laboratories with extensive histories in microbial genomics and bioinformatics. For this community, open access to genome sequences, software and, more recently, publications has tended to be the rule rather than the exception. Indeed, a 2004 National Research Council report described “the culture of genomics” as “unique in its evolution into a global web of tools and information” (Ref. 106). The same report includes a series of recommendations on access to pathogen genome data, including the statement that “rapid, unrestricted public access to primary genome sequence data, annotations of genome data, genome databases, and Internet-based tools for genome analysis should be encouraged” (Ref. 106).
As genomics has moved into the domain of clinical and public health practice, the notion of free and immediate access to genomic surveillance data has encountered several barriers: the siloing of critical metadata across multiple public health databases with no interoperability; balancing openness and transparency with patient privacy and safety; variable data quality, particularly in resource-limited settings; concerns over data reuse by third parties; a lack of standards and ontologies to capture metadata; and career advancement disincentives to releasing data107,108,109. Despite these challenges, the spirit of open access and open data remains strong in the community, with over 40 public health leaders from around the world recently signing a joint statement on data sharing for public health surveillance110. The Ebola and Zika responses in particular highlight the role of real-time sharing of data and samples, be it through the use of chat groups and a LabKey server to disseminate Zika data111 or GitHub to share Ebola data112.
In the wake of Ebola, Yozwiak et al.113 and Chretien et al.114 outline additional issues facing data sharing, from differing cultures and academic norms to complicated consent procedures and technical limitations. They note that we as a community must agree on standards and practices promoting cooperation — a conversation that could begin by examining how the Global Alliance for Genomics and Health (GA4GH) framework for responsible sharing of genomic and health-related data (Box 1) could be adapted for the digital pathogen surveillance community.
The future: the sequencing singularity?
Transformative change to public and global health is profoundly difficult. Complicating the existence of a rapid, open, transparent response is the fact that no matter the setting, there are often conflicting interests at work. In an outbreak scenario, conflict may result from governments wishing to keep an outbreak quiet and/or from the tension between lower-income and middle-income countries with few resources for generating and using data and the researchers or response teams from better-resourced settings115. Indeed, the conflicting values in outbreak responses meet the definition of a 'wicked' problem, where issues resist simple resolution and span multiple jurisdictions and where each stakeholder has a different perspective on the solution. Even the International Health Regulations (IHR), which ostensibly provide a legal instrument for global health security, fail to effect a basic surveillance and outbreak response. As of the most recent self-reporting, only 30% of the 196 member countries of the IHR are in compliance, meeting the prescribed minimum public health core capacities5. In these settings, digital pathogen surveillance must be within the purview of the larger global health community and its diverse group of non-state actors rather than being solely the responsibility of nations themselves116. This raises an important issue: if nations are willing to cede a certain amount of surveillance and diagnostic control to the global health community, the notion of reciprocity suggests that they should derive some corresponding local benefit. The 'trickle-down' effects of global genomic surveillance have yet to be fully articulated, but they are likely to be realized first in the zoonotic domain, where global surveillance efforts will feed back into improved animal health at a local level, in turn benefiting local farmers.
Outbreaks occur at the intersection of risk perception, governance, policy and economics117, and outbreak response is often based on political instinct rather than data5. Building a resilient and responsive public health system is therefore more than just enhancing surveillance and coupling it to novel technology — it is about engagement, trust, cooperation and building local capacity8, as well as a focus on pandemic prevention through development rather than pandemic response via disaster relief mechanisms57. Expert panels convened by Harvard and the London School of Hygiene and Tropical Medicine9 and by the National Academy of Medicine8 have called for a central pandemic preparedness and response agency and also underscored the need for deeper partnerships between formal and informal surveillance, epidemiology and academic and public health networks5. More recently, evolutionary biologist Michael Worobey wrote: “Systematic pathogen surveillance is within our grasp, but is still undervalued and underfunded relative to the magnitude of the threat” (Ref. 118). If we are to achieve the sequencing singularity — the moment at which pathogen, environmental and digital data streams are integrated into a global surveillance system — we require a community united behind a vision in which public health and the attendant data belong to the public and behind the idea that we are a better, healthier society when the public is able to access and benefit from the data being collected about us and the pathogens we share the planet with.
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
References
Dudas, G. et al. Virus genomes reveal factors that spread and sustained the Ebola epidemic. Nature 544, 309–315 (2017). This large, retrospective genomic analysis of the Ebola outbreak demonstrates how phylodynamic approaches can provide important insight into the epidemiology of the outbreak.
Faria, N. R. et al. Zika virus in the Americas: Early epidemiological and genetic findings. Science 352, 345–349 (2016). This work is the first to leverage genome sequences generated early in the Zika outbreak to provide a real-time glimpse into the spread of the virus.
Faria, N. R. et al. Establishment and cryptic transmission of Zika virus in Brazil and the Americas. Nature 546, 406–410 (2017).
Grubaugh, N. D. et al. Genomic epidemiology reveals multiple introductions of Zika virus into the United States. Nature 546, 401–405 (2017). This paper is the first to use a genomic approach to track the entry of Zika into the USA.
Gostin, L. O. Our shared vulnerability to dangerous pathogens. Med. Law Rev. 25, 185–199 (2017).
Braden, C. R., Dowell, S. F., Jernigan, D. B. & Hughes, J. M. Progress in global surveillance and response capacity 10 years after severe acute respiratory syndrome. Emerg. Infect. Dis. 19, 864–869 (2013).
Evans, D. K. & Popova, A. West African Ebola crisis and orphans. Lancet 385, 945–946 (2015).
Commission on a Global Health Risk Framework for the Future. The Neglected Dimension of Global Security: A Framework to Counter Infectious Disease Crises (National Academies Press (US), 2016). In the wake of the Ebola crisis, the Commission on a Global Health Risk Framework for the Future presented this report to describe the institutional, policy and financial framework needed for public health preparedness.
Moon, S. et al. Will Ebola change the game? Ten essential reforms before the next pandemic. The report of the Harvard-LSHTM Independent Panel on the Global Response to Ebola. Lancet 386, 2204–2221 (2015).
Deurenberg, R. H. et al. Application of next generation sequencing in clinical microbiology and infection prevention. J. Biotechnol. 243, 16–24 (2017).
Schlaberg, R. et al. Validation of metagenomic next-generation sequencing tests for universal pathogen detection. Arch. Pathol. Lab. Med. 141, 776–786 (2017). This report, from the American Society for Microbiology and the College of American Pathologists, provides a comprehensive overview of clinical metagenomics and the associated validation challenges.
Wilson, M. R. et al. Diagnosing Balamuthia mandrillaris encephalitis with metagenomic deep sequencing. Ann. Neurol. 78, 722–730 (2015).
Wilson, M. R. et al. Actionable diagnosis of neuroleptospirosis by next-generation sequencing. N. Engl. J. Med. 370, 2408–2417 (2014).
Quick, J. et al. Multiplex PCR method for MinION and Illumina sequencing of Zika and other virus genomes directly from clinical samples. Nat. Protoc. 12, 1261–1276 (2017).
Houldcroft, C. J., Beale, M. A. & Breuer, J. Clinical and biological insights from viral genome sequencing. Nat. Rev. Microbiol. 15, 183–192 (2017).
Grumaz, S. et al. Next-generation sequencing diagnostics of bacteremia in septic patients. Genome Med. 8, 73 (2016).
Pendleton, K. M. et al. Rapid pathogen identification in bacterial pneumonia using real-time metagenomics. Am. J. Respir. Crit. Care Med. http://dx.doi.org/10.1164/rccm.201703-0537LE (2017).
Schmidt, K. et al. Identification of bacterial pathogens and antimicrobial resistance directly from clinical urines by nanopore-based metagenomic sequencing. J. Antimicrob. Chemother. 72, 104–114 (2017).
Doan, T. et al. Illuminating uveitis: metagenomic deep sequencing identifies common and rare pathogens. Genome Med. 8, 90 (2016).
Loman, N. J. et al. Performance comparison of benchtop high-throughput sequencing platforms. Nat. Biotechnol. 30, 434–439 (2012).
Jain, M., Olsen, H. E., Paten, B. & Akeson, M. The Oxford Nanopore MinION: delivery of nanopore sequencing to the genomics community. Genome Biol. 17, 239 (2016).
Quick, J. et al. Real-time, portable genome sequencing for Ebola surveillance. Nature 530, 228–232 (2016).
Hoenen, T. et al. Nanopore sequencing as a rapidly deployable ebola outbreak tool. Emerg. Infect. Dis. 22, 331–334 (2016).
Faria, N. R. et al. Mobile real-time surveillance of Zika virus in Brazil. Genome Med. 8, 97 (2016).
Edwards, A., Debbonaire, A. R., Sattler, B., Mur, L. A. & Hodson, A. J. Extreme metagenomics using nanopore DNA sequencing: a field report from Svalbard, 78 N. bioRxiv http://dx.doi.org/10.1101/073965 (2016).
Johnson, S. S., Zaikova, E., Goerlitz, D. S., Bai, Y. & Tighe, S. W. Real-time DNA sequencing in the Antarctic Dry Valleys ising the Oxford Nanopore sequencer. J. Biomol. Tech. 28, 2–7 (2017).
Edwards, A. et al. Deep sequencing: intra-terrestrial metagenomics illustrates the potential of off-grid Nanopore DNA sequencing. bioRxiv http://dx.doi.org/10.1101/133413 (2017).
McIntyre, A. B. R. et al. Nanopore sequencing in microgravity. npj Microgravity 2, 16035 (2016).
Castro-Wallace, S. L. et al. Nanopore DNA sequencing and genome assembly on the International Space Station. bioRxiv http://dx.doi.org/10.1101/077651 (2016).
Nichol, S. T. et al. Genetic identification of a hantavirus associated with an outbreak of acute respiratory illness. Science 262, 914–917 (1993).
Holmes, E. C. et al. The molecular epidemiology of human immunodeficiency virus type 1 in Edinburgh. J. Infect. Dis. 171, 45–53 (1995).
Popovich, K. J. & Snitkin, E. S. Whole genome sequencing — implications for infection prevention and outbreak investigations. Curr. Infect. Dis. Rep. 19, 15 (2017).
Joensen, K. G. et al. Real-time whole-genome sequencing for routine typing, surveillance, and outbreak detection of verotoxigenic Escherichia coli. J. Clin. Microbiol. 52, 1501–1510 (2014).
Graham, R. M. A., Doyle, C. J. & Jennison, A. V. Real-time investigation of a Legionella pneumophila outbreak using whole genome sequencing. Epidemiol. Infect. 142, 2347–2351 (2014).
Inns, T. et al. A multi-country Salmonella enteritidis phage type 14b outbreak associated with eggs from a German producer: 'near real-time' application of whole genome sequencing and food chain investigations, United Kingdom, May to September 2014. Eurosurveillance 20, 21098 (2015).
Quick, J. et al. Rapid draft sequencing and real-time nanopore sequencing in a hospital outbreak of Salmonella. Genome Biol. 16, 114 (2015).
Jackson, B. R. et al. Implementation of nationwide real-time whole-genome sequencing to enhance listeriosis outbreak detection and investigation. Clin. Infect. Dis. 63, 380–386 (2016).
Guthrie, J. L. & Gardy, J. L. A brief primer on genomic epidemiology: lessons learned from Mycobacterium tuberculosis. Ann. NY Acad. Sci. 1388, 59–77 (2017).
Didelot, X., Fraser, C., Gardy, J. & Colijn, C. Genomic infectious disease epidemiology in partially sampled and ongoing outbreaks. Mol. Biol. Evol. 34, 997–1007 (2017).
Grenfell, B. T. et al. Unifying the epidemiological and evolutionary dynamics of pathogens. Science 303, 327–332 (2004). This paper introduces the concept of phylodynamics, which has since become a key tool in the population genomics and epidemiology toolboxes.
Drummond, A. J., Pybus, O. G., Rambaut, A., Forsberg, R. & Rodrigo, A. G. Measurably evolving populations. Trends Ecol. Evol. 18, 481–488 (2003).
Duchêne, S. et al. Genome-scale rates of evolutionary change in bacteria. Microb. Genom. 2, e000094 (2016). This paper is the first to demonstrate the degree to which the concept of a measurably evolving population can be applied to bacteria.
Norström, M. M., Karlsson, A. C. & Salemi, M. Towards a new paradigm linking virus molecular evolution and pathogenesis: experimental design and phylodynamic inference. New Microbiol. 35, 101–111 (2012).
Drummond, A. J., Suchard, M. A., Xie, D. & Rambaut, A. Bayesian phylogenetics with BEAUti and the BEAST 1.7. Mol. Biol. Evol. 29, 1969–1973 (2012). This paper describes BEAST, a frequently used toolkit for phylogenetics and phylodynamic reconstructions.
Popinga, A., Vaughan, T., Stadler, T. & Drummond, A. J. Inferring epidemiological dynamics with Bayesian coalescent inference: the merits of deterministic and stochastic models. Genetics 199, 595–607 (2015).
Pybus, O. G. et al. The epidemic behavior of the hepatitis C virus. Science 292, 2323–2325 (2001).
Baele, G., Suchard, M. A., Rambaut, A. & Lemey, P. Emerging concepts of data integration in pathogen phylodynamics. Syst. Biol. 66, e47–e65 (2017).
Holmes, E. C., Dudas, G., Rambaut, A. & Andersen, K. G. The evolution of Ebola virus: insights from the 2013–2016 epidemic. Nature 538, 193–200 (2016).
Baize, S. et al. Emergence of Zaire Ebola virus disease in Guinea. N. Engl. J. Med. 371, 1418–1425 (2014).
Gire, S. K. et al. Genomic surveillance elucidates Ebola virus origin and transmission during the 2014 outbreak. Science 345, 1369–1372 (2014). This is the first genomics paper to come out of the 2014 Ebola outbreak.
Mate, S. E. et al. Molecular evidence of sexual transmission of Ebola virus. N. Engl. J. Med. 373, 2448–2454 (2015).
Blackley, D. J. et al. Reduced evolutionary rate in reemerged Ebola virus transmission chains. Sci. Adv. 2, e1600378 (2016).
Seytre, B. The wanderings of the communication on the Ebola virus disease. Bull. Soc. Pathol. Exot. 109, 314–323 (2016).
Woolhouse, M. & Gaunt, E. Ecological origins of novel human pathogens. Crit. Rev. Microbiol. 33, 231–242 (2007).
Jones, K. E. et al. Global trends in emerging infectious diseases. Nature 451, 990–993 (2008). This landmark work surveys the emergence of infectious diseases since 1940 and identifies a number of hot spots for disease emergence.
Murray, K. A. et al. Global biogeography of human infectious diseases. Proc. Natl Acad. Sci. USA 112, 12746–12751 (2015).
Bogich, T. L. et al. Preventing pandemics via international development: a systems approach. PLoS Med. 9, e1001354 (2012).
Daszak, P. A call for 'Smart Surveillance': a lesson learned from H1N1. EcoHealth 6, 1–2 (2009).
Pal, C., Bengtsson-Palme, J., Kristiansson, E. & Larsson, D. G. J. The structure and diversity of human, animal and environmental resistomes. Microbiome 4, 54 (2016).
Furtak, V. et al. Environmental surveillance of viruses by tangential flow filtration and metagenomic reconstruction. Eurosurveillance 21, 30193 (2016).
Rosenberg, R., Johansson, M. A., Powers, A. M. & Miller, B. R. Search strategy has influenced the discovery rate of human viruses. Proc. Natl Acad. Sci. USA 110, 13961–13964 (2013).
Rosenberg, R. Detecting the emergence of novel, zoonotic viruses pathogenic to humans. Cell. Mol. Life Sci. 72, 1115–1125 (2015).
Anthony, S. J. et al. Non-random patterns in viral diversity. Nat. Commun. 6, 8147 (2015).
Shi, M. et al. Redefining the invertebrate RNA virosphere. Nature 540, 539–543 (2016).
Morse, S. S. et al. Prediction and prevention of the next pandemic zoonosis. Lancet 380, 1956–1965 (2012).
Cook, R., Karesh, W. & Osofsky, S. Conference summary: One World, One Health: building interdisciplinary bridges to health in a globalized world. One World, One Health http://www.oneworldonehealth.org/sept2004/owoh_sept04.html (2004).
Kelly, T. R. et al. One Health proof of concept: bringing a transdisciplinary approach to surveillance for zoonotic viruses at the human-wild animal interface. Prev. Vet. Med. 137, 112–118 (2017).
Smiley Evans, T. et al. Optimization of a novel non-invasive oral sampling technique for zoonotic pathogen surveillance in nonhuman primates. PLoS Negl. Trop. Dis. 9, e0003813 (2015).
Anthony, S. J. et al. A strategy to estimate unknown viral diversity in mammals. mBio 4, e00598-13 (2013).
Trewby, H., Nadin-Davis, S. A., Real, L. A. & Biek, R. Processes underlying rabies virus incursions across US-Canada border as revealed by whole-genome phylogeography. Emerg. Infect. Dis. 23, 1454–1461 (2017).
Filejski, C. The changing face of rabies in Canada. Can. Comm. Rep. 42, 118–120 (2016).
Kamath, P. L. et al. Genomics reveals historic and contemporary transmission dynamics of a bacterial disease among wildlife and livestock. Nat. Commun. 7, 11448 (2016).
Godfroid, J. Brucellosis in livestock and wildlife: zoonotic diseases without pandemic potential in need of innovative one health approaches. Arch. Public Health 75, 34 (2017).
Temmam, S., Davoust, B., Berenger, J.-M., Raoult, D. & Desnues, C. Viral metagenomics on animals as a tool for the detection of zoonoses prior to human infection? Int. J. Mol. Sci. 15, 10377–10397 (2014).
Phan, M. V. T. et al. Unbiased whole-genome deep sequencing of human and porcine stool samples reveals circulation of multiple groups of rotaviruses and a putative zoonotic infection. Virus Evol. 2, vew027 (2016).
Abat, C., Chaudet, H., Rolain, J.-M., Colson, P. & Raoult, D. Traditional and syndromic surveillance of infectious diseases and pathogens. Int. J. Infect. Dis. 48, 22–28 (2016).
National Research Council (US) Committee on Achieving Sustainable Global Capacity for Surveillance and Response to Emerging Diseases of Zoonotic Origin. Sustaining Global Surveillance and Response to Emerging Zoonotic Diseases. (National Academies Press (US), 2009).
Mandl, K. D. et al. Implementing syndromic surveillance: a practical guide informed by the early experience. J. Am. Med. Inform. Assoc. 11, 141–150 (2004).
Henning, K. J. What is syndromic surveillance? MMWR Suppl. 53, 5–11 (2004).
Smolinski, M. S. et al. Flu Near You: crowdsourced symptom reporting spanning 2 influenza seasons. Am. J. Public Health 105, 2124–2130 (2015).
Aslam, A. A. et al. The reliability of tweets as a supplementary method of seasonal influenza surveillance. J. Med. Internet Res. 16, e250 (2014).
Paul, M. J., Dredze, M. & Broniatowski, D. Twitter improves influenza forecasting. PLoS Curr. http://dx.doi.org/10.1371/currents.outbreaks.90b9ed0f59bae4ccaa683a39865d9117 (2014).
Hulth, A., Rydevik, G. & Linde, A. Web queries as a source for syndromic surveillance. PLoS ONE 4, e4378 (2009).
Carneiro, H. A. & Mylonakis, E. Google trends: a web-based tool for real-time surveillance of disease outbreaks. Clin. Infect. Dis. 49, 1557–1564 (2009).
Cook, S., Conrad, C., Fowlkes, A. L. & Mohebbi, M. H. Assessing Google flu trends performance in the United States during the 2009 influenza virus A (H1N1) pandemic. PLoS ONE 6, e23610 (2011).
Brownstein, J. S., Freifeld, C. C. & Madoff, L. C. Digital disease detection — harnessing the Web for public health surveillance. N. Engl. J. Med. 360, 2153–2157 (2009). This paper introduces the notion of digital epidemiology to the wider community.
Hartley, D. M. et al. An overview of internet biosurveillance. Clin. Microbiol. Infect. 19, 1006–1013 (2013).
O'Shea, J. Digital disease detection: a systematic review of event-based internet biosurveillance systems. Int. J. Med. Inform. 101, 15–22 (2017).
Brownstein, J. S. & Freifeld, C. C. HealthMap: the development of automated real-time internet surveillance for epidemic intelligence. Eurosurveillance 12, 3322 (2007). HealthMap has become one of the most important digital epidemiology resources; this paper describes how the system works.
Schwind, J. S. et al. Evaluation of local media surveillance for improved disease recognition and monitoring in global hotspot regions. PLoS ONE 9, e110236 (2014).
Olson, S. H. et al. Drivers of emerging infectious disease events as a framework for digital detection. Emerg. Infect. Dis. 21, 1285–1292 (2015).
Flahault, A. et al. Precision global health in the digital age. Swiss Med. Wkly 147, w14423 (2017).
Zinszer, K., Morrison, K., Verma, A. & Brownstein, J. S. Spatial determinants of Ebola virus disease risk for the West African epidemic. PLoS Curr. http://dx.doi.org/10.1371/currents.outbreaks.b494f2c6a396c72ec24cb4142765bb95 (2017).
Majumder, M. S. et al. Utilizing nontraditional data sources for near real-time estimation of transmission dynamics during the 2015–2016 Colombian Zika virus disease outbreak. JMIR Public Health Surveill. 2, e30 (2016).
Khoury, M. J., Iademarco, M. F. & Riley, W. T. Precision public health for the era of precision medicine. Am. J. Prev. Med. 50, 398–401 (2016).
Neher, R. A. & Bedford, T. nextflu: real-time tracking of seasonal influenza virus evolution in humans. Bioinformatics 31, 3546–3548 (2015). This paper describes the nextflu project, which gave rise to the Nextstrain platform, whose approach to analysis and visualization recently earned an international prize for open science.
Newson, A. J., Leonard, S. J., Hall, A. & Gaff, C. L. Known unknowns: building an ethics of uncertainty into genomic medicine. BMC Med. Genom. 9, 57 (2016).
Birko, S., Dove, E. S. & Özdemir, V. A. Delphi technology foresight study: mapping social construction of scientific evidence on metagenomics tests for water safety. PLoS ONE 10, e0129706 (2015).
Budowle, B. et al. Criteria for validation of methods in microbial forensics. Appl. Environ. Microbiol. 74, 5599–5607 (2008).
Schmedes, S. E., Sajantila, A. & Budowle, B. Expansion of microbial forensics. J. Clin. Microbiol. 54, 1964–1974 (2016).
Budowle, B. et al. Validation of high throughput sequencing and microbial forensics applications. Investig. Genet. 5, 9 (2014).
Calain, P. The Ebola clinical trials: a precedent for research ethics in disasters. J. Med. Ethics http://dx.doi.org/10.1136/medethics-2016-103474 (2016).
Ishii, T. Germline genome-editing research and its socioethical implications. Trends Mol. Med. 21, 473–481 (2015).
Howard, H. C. et al. The ethical introduction of genome-based information and technologies into public health. Public Health Genomics 16, 100–109 (2013).
Geller, G. et al. Genomics and infectious disease: a call to identify the ethical, legal and social implications for public health and clinical practice. Genome Med. 6, 106 (2014).
National Research Council (US) Committee on Genomics Databases for Bioterrorism Threat Agents. Seeking Security: Pathogens, Open Access, and Genome Databases. (National Academies Press (US), 2004).
Ross, E. Perspectives on data sharing in disease surveillance. Chatham House: The Royal Institute of International Affairs https://www.chathamhouse.org/sites/files/chathamhouse/home/chatham/public_html/sites/default/files/20140430DataSharingDiseaseSurveillanceRoss.pdf (2014).
Sane, J. & Edelstein, M. Overcoming barriers to data sharing in public health: a global perspective. Chatham House: The Royal Institute of International Affairs https://www.chathamhouse.org/sites/files/chathamhouse/field/field_document/20150417OvercomingBarriersDataSharingPublicHealthSaneEdelstein.pdf (2015).
Raza, S. & Luheshi, L. Big data or bust: realizing the microbial genomics revolution. Microb. Genomics http://dx.doi.org/10.1099/mgen.0.000046 (2016).
International Association of Public Health Institutes. Public health surveillance: a call to share data. International Association of Public Health Institutes http://ianphi.org/news/2016/datasharing1.html (2016).
Kallas, E. G. & O'Connor, D. H. Real-time sharing of Zika virus data in an interconnected world. JAMA Pediatr. 170, 633–634 (2016).
Perkel, J. Democratic databases: science on GitHub. Nature 538, 127–128 (2016).
Yozwiak, N. L., Schaffner, S. F. & Sabeti, P. C. Data sharing: make outbreak research open access. Nature 518, 477–479 (2015).
Chretien, J.-P., Rivers, C. M. & Johansson, M. A. Make data sharing routine to prepare for public health emergencies. PLoS Med. 13, e1002109 (2016).
Bull, S. et al. Best practices for ethical sharing of individual-level health research data from low- and middle-income settings. J. Empir. Res. Hum. Res. Ethics 10, 302–313 (2015).
Gostin, L. O. & Mok, E. A. Grand challenges in global health governance. Br. Med. Bull. 90, 7–18 (2009).
Fournier, A., Young, I., Rajic, A., Greig, J. & LeJeune, J. Social and economic aspects of the transmission of pathogenic bacteria between wildlife and food animals: a thematic analysis of published research knowledge. Zoonoses Public Health 62, 417–428 (2015).
Worobey, M. Epidemiology: molecular mapping of Zika spread. Nature 546, 355–357 (2017).
Knoppers, B. M. Framework for responsible sharing of genomic and health-related data. HUGO J. 8, 3 (2014). This document summarizes the GA4GH's statement on data sharing.
Plourde, A. R. & Bloch, E. M. A. Literature review of Zika virus. Emerg. Infect. Dis. 22, 1185–1192 (2016).
Hall, M. D., Woolhouse, M. E. J. & Rambaut, A. Using genomics data to reconstruct transmission trees during disease outbreaks. Rev. Sci. Tech. 35, 287–296 (2016).
Acknowledgements
J.L.G. is funded by the Canada Research Chairs and Michael Smith Foundation for Health Research programmes.
Author information
Authors and Affiliations
Contributions
Both authors contributed equally to all aspects of the article.
Corresponding author
Ethics declarations
Competing interests
J.L.G. declares no competing interests. N.J.L. has received travel expenses and accommodation and an honorarium payment from Oxford Nanopore Technologies to speak at organized symposia. N.J.L. is a member of the Oxford Nanopore MinION Access Programme and has received reagents for nanopore sequencing free of charge.
Related links
Glossary
- Public health surveillance
-
The systematic collection, analysis and dissemination of health-related data to support planning, implementation and evaluation of public health practices and response.
- Outbreaks
-
Outbreaks and epidemics are both defined as increases in the number of cases of a particular disease beyond what is expected in a given setting. In outbreaks, the affected settings are smaller geographic regions; epidemics can span larger areas.
- Pandemic
-
An epidemic that has grown to span multiple countries or continents, often with many affected individuals.
- Cluster
-
A group of epidemiologically related cases defined by their relationship in space and time or via molecular methods.
- Metagenomics
-
The sequencing of genetic material recovered directly from a sample, whether environmental or clinical, permitting the identification of all organisms represented in the sample.
- Bait probes
-
Nucleic acid probes designed to recognize and capture specific DNA sequences, allowing for the enrichment of DNA from a specific organism of interest.
- Emerging infectious diseases
-
(EIDs). Diseases that have recently appeared in a population or that have transitioned from a small number of isolated cases to many cases.
- Transmission
-
The event through which a pathogen is transferred from one entity to another. Transmission can be person-to-person, as in the case of Ebola, vector-to-person, as with Zika, or environment-to-person via routes including food, water and contact with a contaminated object or surface.
- Genomic epidemiology
-
The use of genome sequencing to understand infectious disease transmission and epidemiology. See Fig. 3.
- Basic reproductive number R0
-
The average number of secondary cases of an infectious disease produced by a single infectious case, given a completely susceptible population.
- Zoonotic
-
A term describing infectious diseases that typically exist in an animal reservoir but that can be transmitted to humans.
- Survivor transmission
-
The transmission of an infectious disease, such as Ebola, from a survivor of that disease who has recovered from their symptoms.
- Vector-borne
-
A term describing infectious diseases that are transmitted to humans through contact with a non-human species, particularly those diseases spread through insect bites. An example is the Zika virus, which is carried by mosquitos.
- Hot spots
-
Geographical settings where a variety of factors converge to create the social and environmental conditions that promote disease transmission.
- Spillover
-
The process by which an infectious disease changes from existing exclusively in animals to being able to infect, then transmit between, humans. See Fig. 4.
Rights and permissions
About this article
Cite this article
Gardy, J., Loman, N. Towards a genomics-informed, real-time, global pathogen surveillance system. Nat Rev Genet 19, 9–20 (2018). https://doi.org/10.1038/nrg.2017.88
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrg.2017.88
This article is cited by
-
Tips and tools to obtain and assess mosquito viromes
Archives of Microbiology (2024)
-
Web-Based Tools Validation for Antimicrobial Resistance Prediction: An Empirical Comparative Analysis
SN Computer Science (2024)
-
A machine learning classifier using 33 host immune response mRNAs accurately distinguishes viral and non-viral acute respiratory illnesses in nasal swab samples
Genome Medicine (2023)
-
Global phylodynamics of two relevant aphid-transmitted viruses in cucurbit crops: cucurbit aphid-borne yellows virus and watermelon mosaic virus
Phytopathology Research (2023)
-
Classifying literature mentions of biological pathogens as experimentally studied using natural language processing
Journal of Biomedical Semantics (2023)