Abstract
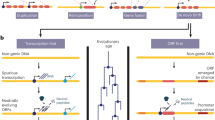
The study of de novo protein-coding genes is maturing from the ad hoc reporting of individual cases to the systematic analysis of extensive genomic data from several species. We identify three key challenges for this emerging field: understanding how best to identify de novo genes, how they arise and why they spread. We highlight the intellectual challenges of understanding how a de novo gene becomes integrated into pre-existing functions and becomes essential. We suggest that, as with protein sequence evolution, antagonistic co-evolution may be key to de novo gene evolution, particularly for new essential genes and new cancer-associated genes.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Change history
27 July 2016
In Table 1 of the original version of this article the gene name NCYM was incorrectly written as NYCM. This has now been corrected. The editors apologize for this error.
References
Levine, M. T., Jones, C. D., Kern, A. D., Lindfors, H. A. & Begun, D. J. Novel genes derived from noncoding DNA in Drosophila melanogaster are frequently X-linked and exhibit testis-biased expression. Proc. Natl Acad. Sci. USA 103, 9935–9939 (2006).
Begun, D. J., Lindfors, H. A., Thompson, M. E. & Holloway, A. K. Recently evolved genes identified from Drosophila yakuba and D. erecta accessory gland expressed sequence tags. Genetics 172, 1675–1681 (2006).
Xiao, W. et al. A rice gene of de novo origin negatively regulates pathogen-induced defense response. PLoS ONE 4, e4603 (2009).
Knowles, D. G. & McLysaght, A. Recent de novo origin of human protein-coding genes. Genome Res. 19, 1752–1759 (2009).
Li, L. et al. Identification of the novel protein QQS as a component of the starch metabolic network in Arabidopsis leaves. Plant J. 58, 485–498 (2009).
Cai, J., Zhao, R., Jiang, H. & Wang, W. De novo origination of a new protein-coding gene in Saccharomyces cerevisiae. Genetics 179, 487–496 (2008).
Zhou, Q. & Wang, W. On the origin and evolution of new genes — a genomic and experimental perspective. J. Genet. Genom. 35, 639–648 (2008).
Toll-Riera, M. et al. Origin of primate orphan genes: a comparative genomics approach. Mol. Biol. Evol. 26, 603–612 (2009).
Wu, D.-D., Irwin, D. M. & Zhang, Y.-P. De novo origin of human protein-coding genes. PLoS Genet. 7, e1002379 (2011).
Tautz, D. & Domazet-Loso, T. The evolutionary origin of orphan genes. Nat. Rev. Genet. 12, 692–702 (2011).
McLysaght, A. & Guerzoni, D. New genes from non-coding sequence: the role of de novo protein-coding genes in eukaryotic evolutionary innovation. Phil. Trans. R. Soc. B 370, 20140332 (2015).
Schlötterer, C. Genes from scratch — the evolutionary fate of de novo genes. Trends Genet. 31, 215–219 (2015).
Guerzoni, D. & McLysaght, A. De novo genes arise at a slow but steady rate along the primate lineage and have been subject to incomplete lineage sorting. Genome Biol. Evol. 8, 1222–1232 (2016).
Domazet-Loso, T., Brajkovic´, J. & Tautz, D. A phylostratigraphy approach to uncover the genomic history of major adaptations in metazoan lineages. Trends Genet. 23, 533–539 (2007).
Wolfe, K. Evolutionary genomics: yeasts accelerate beyond BLAST. Curr. Biol. 14, R392–R394 (2004).
Elhaik, E., Sabath, N. & Graur, D. The “inverse relationship between evolutionary rate and age of mammalian genes” is an artifact of increased genetic distance with rate of evolution and time of divergence. Mol. Biol. Evol. 23, 1–3 (2006).
Moyers, B. A. & Zhang, J. Phylostratigraphic bias creates spurious patterns of genome evolution. Mol. Biol. Evol. 32, 258–267 (2015).
Moyers, B. A. & Zhang, J. Evaluating phylostratigraphic evidence for widespread de novo gene birth in genome evolution. Mol. Biol. Evol. 33, 1245–1256 (2016).
Carvunis, A.-R. et al. Proto-genes and de novo gene birth. Nature 487, 370–374 (2012).
Neme, R. & Tautz, D. Phylogenetic patterns of emergence of new genes support a model of frequent de novo evolution. BMC Genomics 14, 117 (2013).
Alba, M. M. & Castresana, J. On homology searches by protein Blast and the characterization of the age of genes. BMC Evol. Biol. 7, 53 (2007).
Alba, M. M. & Castresana, J. Inverse relationship between evolutionary rate and age of mammalian genes. Mol. Biol. Evol. 22, 598–606 (2005).
Domazet-Loso, T. & Tautz, D. An ancient evolutionary origin of genes associated with human genetic diseases. Mol. Biol. Evol. 25, 2699–2707 (2008).
Smith, N. G. C. & Eyre-Walker, A. Human disease genes: patterns and predictions. Gene 318, 169–175 (2003).
ENCODE Project Consortium. An integrated encyclopedia of DNA elements in the human genome. Nature 489, 57–74 (2012).
Hurst, L. D. Open questions: a logic (or lack thereof) of genome organization. BMC Biol. 11, 58 (2013).
Graur, D. et al. On the immortality of television sets: 'function' in the human genome according to the evolution-free gospel of ENCODE. Genome Biol. Evol. 5, 578–590 (2013).
Doolittle, W. F. Is junk DNA bunk? A critique of ENCODE. Proc. Natl Acad. Sci. USA 110, 5294–5300 (2013).
Jaillon, O. et al. Translational control of intron splicing in eukaryotes. Nature 451, 359–362 (2008).
Cusack, B. P., Arndt, P. F., Duret, L. & Roest Crollius, H. Preventing dangerous nonsense: selection for robustness to transcriptional error in human genes. PLoS Genet. 7, e1002276 (2011).
Dewey, C. N., Rogozin, I. B. & Koonin, E. V. Compensatory relationship between splice sites and exonic splicing signals depending on the length of vertebrate introns. BMC Genomics 7, 311 (2006).
Schüler, A., Ghanbarian, A. T. & Hurst, L. D. Purifying selection on splice-related motifs, not expression level nor RNA folding, explains nearly all constraint on human lincRNAs. Mol. Biol. Evol. 31, 3164–3183 (2014).
Ruiz-Orera, J., Messeguer, X., Subirana, J. A. & Alba, M. M. Long non-coding RNAs as a source of new peptides. eLife 3, e03523 (2014).
Chen, J.-Y. et al. Emergence, retention and selection: a trilogy of origination for functional de novo proteins from ancestral lncRNAs in primates. PLoS Genet. 11, e1005391 (2015).
Zhao, L., Saelao, P., Jones, C. D. & Begun, D. J. Origin and spread of de novo genes in Drosophila melanogaster populations. Science 343, 769–772 (2014).
Galtier, N., Duret, L., Glémin, S. & Ranwez, V. GC-biased gene conversion promotes the fixation of deleterious amino acid changes in primates. Trends Genet. 25, 1–5 (2009).
Blomen, V. A. et al. Gene essentiality and synthetic lethality in haploid human cells. Science 350, 1092–1096 (2015).
Wang, T. et al. Identification and characterization of essential genes in the human genome. Science 350, 1096–1101 (2015).
Wang, J. et al. Primate-specific endogenous retrovirus-driven transcription defines naive-like stem cells. Nature 516, 405–409 (2014).
Lavialle, C. et al. Paleovirology of 'syncytins', retroviral env genes exapted for a role in placentation. Phil. Trans. R. Soc. B 368, 20120507 (2013).
Li, D., Yan, Z., Lu, L., Jiang, H. & Wang, W. Pleiotropy of the de novo-originated gene MDF1. Sci. Rep. 4, 7280 (2014).
Li, D. et al. A de novo originated gene depresses budding yeast mating pathway and is repressed by the protein encoded by its antisense strand. Cell Res. 20, 408–420 (2010).
Ghysen, A. Debatable issues. Interview with L Wolpert and A García-Bellido. Int. J. Dev. Biol. 42, 511–518 (1998).
Tautz, D. A genetic uncertainty problem. Trends Genet. 16, 475–477 (2000).
Chalfin, L. et al. Mapping ecologically relevant social behaviours by gene knockout in wild mice. Nat. Commun. 5, 4569 (2014).
Xu, J. & Zhang, J. Are human translated pseudogenes functional? Mol. Biol. Evol. 33, 755–760 (2016).
Chen, S., Zhang, Y. E. & Long, M. New genes in Drosophila quickly become essential. Science 330, 1682–1685 (2010).
Bird, A. P. Gene number, noise reduction and biological complexity. Trends Genet. 11, 94–100 (1995).
Hurst, L. D. Evolutionary genomics and the reach of selection. J. Biol. 8, 12 (2009).
Prestridge, D. S. & Burks, C. The density of transcriptional elements in promoter and non-promoter sequences. Hum. Mol. Genet. 2, 1449–1453 (1993).
Hoekstra, H. E. & Coyne, J. A. The locus of evolution: evo devo and the genetics of adaptation. Evolution 61, 995–1016 (2007).
Begun, D. J., Lindfors, H. A., Kern, A. D. & Jones, C. D. Evidence for de novo evolution of testis-expressed genes in the Drosophila yakuba/Drosophila erecta clade. Genetics 176, 1131–1137 (2007).
Ebisuya, M., Yamamoto, T., Nakajima, M. & Nishida, E. Ripples from neighbouring transcription. Nat. Cell Biol. 10, 1106–1113 (2008).
Siepel, A. Darwinian alchemy: human genes from noncoding DNA. Genome Res. 19, 1693–1695 (2009).
Murphy, D. N. & McLysaght, A. De novo origin of protein-coding genes in murine rodents. PLoS ONE 7, e48650 (2012).
Gotea, V., Petrykowska, H. M. & Elnitski, L. Bidirectional promoters as important drivers for the emergence of species-specific transcripts. PLoS ONE 8, e57323 (2013).
Wu, X. & Sharp, P. A. Divergent transcription: a driving force for new gene origination? Cell 155, 990–996 (2013).
Akiva, P. et al. Transcription-mediated gene fusion in the human genome. Genome Res. 16, 30–36 (2006).
Parra, G. et al. Tandem chimerism as a means to increase protein complexity in the human genome. Genome Res. 16, 37–44 (2006).
Nacu, S. et al. Deep RNA sequencing analysis of readthrough gene fusions in human prostate adenocarcinoma and reference samples. BMC Med. Genom. 4, 11 (2011).
Ruiz-Orera, J. et al. Origins of de novo genes in human and chimpanzee. PLoS Genet. 11, e1005721 (2015).
Neme, R. & Tautz, D. Fast turnover of genome transcription across evolutionary time exposes entire non-coding DNA to de novo gene emergence. eLife 5, e09977 (2016).
Necsulea, A. & Kaessmann, H. Evolutionary dynamics of coding and non-coding transcriptomes. Nat. Rev. Genet. 15, 734–748 (2014).
Warnecke, T., Huang, Y., Przytycka, T. M. & Hurst, L. D. Unique cost dynamics elucidate the role of frameshifting errors in promoting translational robustness. Genome Biol. Evol. 2, 636–645 (2010).
Lercher, M. J., Urrutia, A. O., Pavlícek, A. & Hurst, L. D. A unification of mosaic structures in the human genome. Hum. Mol. Genet. 12, 2411–2415 (2003).
Wang, J. et al. Sequence features and chromatin structure around the genomic regions bound by 119 human transcription factors. Genome Res. 22, 1798–1812 (2012).
Wang, T. et al. Species-specific endogenous retroviruses shape the transcriptional network of the human tumor suppressor protein p53. 104, 18613–18618 (2007).
Gotea, V. & Makałowski, W. Do transposable elements really contribute to proteomes? Trends Genet. 22, 260–267 (2006).
Thornburg, B. G., Gotea, V. & Makałowski, W. Transposable elements as a significant source of transcription regulating signals. Gene 365, 104–110 (2006).
Göke, J. et al. Dynamic transcription of distinct classes of endogenous retroviral elements marks specific populations of early human embryonic cells. Cell Stem Cell 16, 135–141 (2015).
Denli, A. M. et al. Primate-specific ORF0 contributes to retrotransposon-mediated diversity. Cell 163, 583–593 (2015).
Wang, Y. et al. Endogenous miRNA sponge lincRNA-RoR regulates Oct4, Nanog, and Sox2 in human embryonic stem cell self-renewal. Dev. Cell 25, 69–80 (2013).
Galagan, J. E., & Selker, E. U. RIP: the evolutionary cost of genome defense. Trends Genet. 20, 417–413 (2004).
Xie, C. et al. Hominoid-specific de novo protein-coding genes originating from long non-coding RNAs. PLoS Genet. 8, e1002942 (2012).
Palmieri, N., Kosiol, C. & Schlötterer, C. The life cycle of Drosophila orphan genes. eLife 3, e01311 (2014).
Neme, R. & Tautz, D. Evolution: dynamics of de novo gene emergence. Curr. Biol. 24, R238–R240 (2014).
Kamijyo, A., Yura, K. & Ogura, A. Distinct evolutionary rate in the eye field transcription factors found by estimation of ancestral protein structure. Gene 555, 73–79 (2015).
Vaquerizas, J. M., Kummerfeld, S. K., Teichmann, S. A. & Luscombe, N. M. A census of human transcription factors: function, expression and evolution. Nat. Rev. Genet. 10, 252–263 (2009).
Hayashi, Y., Sakata, H., Makino, Y., Urabe, I. & Yomo, T. Can an arbitrary sequence evolve towards acquiring a biological function? J. Mol. Evol. 56, 162–168 (2003).
Zhang, W., Landback, P., Gschwend, A. R., Shen, B. & Long, M. New genes drive the evolution of gene interaction networks in the human and mouse genomes. Genome Biol. 16, 202 (2015).
Lercher, M. J. & Pál, C. Integration of horizontally transferred genes into regulatory interaction networks takes many million years. Mol. Biol. Evol. 25, 559–567 (2008).
Batada, N. N., Hurst, L. D. & Tyers, M. Evolutionary and physiological importance of hub proteins. PLoS Comp. Biol. 2, e88 (2006).
Force, A. et al. Preservation of duplicate genes by complementary, degenerative mutations. Genetics 151, 1531–1545 (1999).
Schoorlemmer, J., Pérez-Palacios, R., Climent, M., Guallar, D. & Muniesa, P. Regulation of mouse retroelement MuERV-L/MERVL expression by REX1 and epigenetic control of stem cell potency. Front. Oncol. 4, 14 (2014).
Macfarlan, T. S. et al. Embryonic stem cell potency fluctuates with endogenous retrovirus activity. Nature 487, 57–63 (2012).
Imakawa, K., Nakagawa, S. & Miyazawa, T. Baton pass hypothesis: successive incorporation of unconserved endogenous retroviral genes for placentation during mammalian evolution. Genes Cells 20, 771–788 (2015).
Aakre, C. D. et al. Evolving new protein-protein interaction specificity through promiscuous intermediates. Cell 163, 594–606 (2015).
Esnault, C., Cornelis, G., Heidmann, O. & Heidmann, T. Differential evolutionary fate of an ancestral primate endogenous retrovirus envelope gene, the EnvV syncytin, captured for a function in placentation. PLoS Genet. 9, e1003400 (2013).
Cornelis, G. et al. Retroviral envelope syncytin capture in an ancestrally diverged mammalian clade for placentation in the primitive Afrotherian tenrecs. Proc. Natl Acad. Sci. USA 111, E4332–E4341 (2014).
Cornelis, G. et al. Retroviral envelope gene captures and syncytin exaptation for placentation in marsupials. Proc. Natl Acad. Sci. USA 112, E487–E496 (2015).
Cornelis, G. et al. Captured retroviral envelope syncytin gene associated with the unique placental structure of higher ruminants. Proc. Natl Acad. Sci. USA 110, E828–E837 (2013).
Dupressoir, A., Lavialle, C. & Heidmann, T. From ancestral infectious retroviruses to bona fide cellular genes: role of the captured syncytins in placentation. Placenta 33, 663–671 (2012).
Emera, D. et al. Convergent evolution of endometrial prolactin expression in primates, mice, and elephants through the independent recruitment of transposable elements. Mol. Biol. Evol. 29, 239–247 (2012).
Maston, G. A. & Ruvolo, M. Chorionic gonadotropin has a recent origin within primates and an evolutionary history of selection. Mol. Biol. Evol. 19, 320–335 (2002).
Ross, B. D. et al. Stepwise evolution of essential centromere function in a Drosophila neogene. Science 340, 1211–1214 (2013).
Elliot, M. G. & Crespi, B. J. Phylogenetic evidence for early hemochorial placentation in eutheria. Placenta 30, 949–967 (2009).
Elliot, M. G. & Crespi, B. J. Genetic recapitulation of human pre-eclampsia risk during convergent evolution of reduced placental invasiveness in eutherian mammals. Phil. Trans. R. Soc. B 370, 20140069 (2015).
Izsvák, Z., Wang, J., Singh, M., Mager, D. L. & Hurst, L. D. Pluripotency and the endogenous retrovirus HERVH: conflict or serendipity? Bioessays 38, 109–117 (2016).
Landmann, F., Orsi, G. A., Loppin, B. & Sullivan, W. Wolbachia-mediated cytoplasmic incompatibility is associated with impaired histone deposition in the male pronucleus. PLoS Pathog. 5, e1000343 (2009).
Fine, P. E. On the dynamics of symbiote-dependent cytoplasmic incompatibility in culicine mosquitoes. J. Invertebr. Pathol. 31, 10–18 (1978).
Merrill, C., Bayraktaroglu, L., Kusano, A. & Ganetzky, B. Truncated RanGAP encoded by the Segregation Distorter locus of Drosophila. Science 283, 1742–1745 (1999).
Gerdes, K. et al. The hok killer gene family in gram-negative bacteria. New Biol. 2, 946–956 (1990).
Hurst, L. D. scat+ is a selfish gene analogous to Medea of Tribolium castaneum. Cell 75, 407–408 (1993).
Marshall, J. M. The toxin and antidote puzzle: new ways to control insect pest populations through manipulating inheritance. Bioeng. Bugs 2, 235–240 (2011).
Chen, C.-H. et al. A synthetic maternal-effect selfish genetic element drives population replacement in Drosophila. Science 316, 597–600 (2007).
Phadnis, N. & Orr, H. A. A single gene causes both male sterility and segregation distortion in Drosophila hybrids. Science 323, 376–379 (2009).
Hurst, L. D. & Pomiankowski, A. Causes of sex ratio bias may account for unisexual sterility in hybrids: a new explanation of Haldane's rule and related phenomena. Genetics 128, 841–858 (1991).
Nielsen, R. et al. A scan for positively selected genes in the genomes of humans and chimpanzees. PLoS Biol. 3, e170 (2005).
Kosiol, C. et al. Patterns of positive selection in six mammalian genomes. PLoS Genet. 4, e1000144 (2008).
Goriely, A. et al. Gain-of-function amino acid substitutions drive positive selection of FGFR2 mutations in human spermatogonia. Proc. Natl Acad. Sci. USA 102, 6051–6056 (2005).
Suenaga, Y. et al. NCYM, a cis-antisense gene of MYCN, encodes a de novo evolved protein that inhibits GSK3β resulting in the stabilization of MYCN in human neuroblastomas. PLoS Genet. 10, e1003996 (2014).
Samusik, N., Krukovskaya, L., Meln, I., Shilov, E. & Kozlov, A. P. PBOV1 is a human de novo gene with tumor-specific expression that is associated with a positive clinical outcome of cancer. PLoS ONE 8, e56162 (2013).
Zendman, A. J. W., Ruiter, D. J. & Van Muijen, G. N. P. Cancer/testis-associated genes: identification, expression profile, and putative function. J. Cell. Physiol. 194, 272–288 (2003).
Simpson, A. J. G., Caballero, O. L., Jungbluth, A., Chen, Y.-T. & Old, L. J. Cancer/testis antigens, gametogenesis and cancer. Nat. Rev. Cancer 5, 615–625 (2005).
Hofmann, O. et al. Genome-wide analysis of cancer/testis gene expression. 105, 20422–20427 (2008).
Kohn, D. B., Sadelain, M. & Glorioso, J. C. Occurrence of leukaemia following gene therapy of X-linked SCID. Nat. Rev. Cancer 3, 477–488 (2003).
Bornberg-Bauer, E. & Alba, M. M. Dynamics and adaptive benefits of modular protein evolution. Curr. Opin. Struct. Biol. 23, 459–466 (2013).
Heinen, T. J. A. J., Staubach, F., Häming, D. & Tautz, D. Emergence of a new gene from an intergenic region. Curr. Biol. 19, 1527–1531 (2009).
Broustas, C. G. et al. BRCC2, a novel BH3-like domain-containing protein, induces apoptosis in a caspase-dependent manner. J. Biol. Chem. 279, 26780–26788 (2004).
Broustas, C. G. et al. The proapoptotic molecule BLID interacts with Bcl-XL and its downregulation in breast cancer correlates with poor disease-free and overall survival. Clin. Cancer Res. 16, 2939–2948 (2010).
Andrews, S. J. & Rothnagel, J. A. Emerging evidence for functional peptides encoded by short open reading frames. Nat. Rev. Genet. 15, 193–204 (2014).
Ji, Z., Song, R., Regev, A. & Struhl, K. Many lncRNAs, 5′UTRs, and pseudogenes are translated and some are likely to express functional proteins. eLife 4, e08890 (2015).
Buhl, A. M. et al. Identification of a gene on chromosome 12q22 uniquely overexpressed in chronic lymphocytic leukemia. Blood 107, 2904–2911 (2006).
Lin, B. et al. PART-1: a novel human prostate-specific, androgen-regulated gene that maps to chromosome 5q12. Cancer Res. 60, 858–863 (2000).
Pekarsky, Y., Rynditch, A., Wieser, R., Fonatsch, C. & Gardiner, K. Activation of a novel gene in 3q21 and identification of intergenic fusion transcripts with ecotropic viral insertion site I in leukemia. Cancer Res. 57, 3914–3919 (1997).
Kaushal, A. et al. A novel transcript from the KLKP1 gene is androgen regulated, down-regulated during prostate cancer progression and encodes the first non-serine protease identified from the human kallikrein gene locus. Prostate 68, 381–399 (2008).
Acknowledgements
A.M. and L.D.H. are supported by funding from the European Research Council grant agreements 309834 and 669207, respectively.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Glossary
- Apert syndrome
-
A congenital disorder that is caused by the failure of appropriate apoptosis to occur during fetal development, resulting in malformed skull, face, hands and feet.
- Biased gene conversion
-
Gene conversion (that is, the replacement of a DNA sequence by homologous sequence from the other allele at the same locus, or from elsewhere in the genome) involving a process that repairs mismatches in a non-random fashion. Gene conversion in mammals is thought to be weakly biased towards GC residues over AT residues.
- dN/dS analysis
-
(Also known as Ka/Ks analysis). Analysis to determine the ratio of nonsynonymous substitutions per nonsynonymous site (dN) to synonymous substitutions per synonymous site (dS), which is indicative of the mode of evolution acting on a protein-coding gene. This is interpreted as purifying selection if less than 1; positive selection if greater than 1; and neutral evolution if effectively 1. The numbers of substitutions are estimated by counting the observed differences in orthologous genes identified in at least two different species.
- Domesticated genes
-
Exogenous genetic material that has become incorporated into a genome and subsequently adapted for a host function.
- Fixed
-
An allele is said to be fixed in a population once it rises to 100% frequency.
- Independent lineage sorting
-
This phenomenon is observed when population polymorphism segregating in an ancestral species is maintained past two (or more) speciation events, such that the descendent species each contains alleles that date back to before the speciation events. The descendent species may each independently fix one or other of the ancestral alleles (independently sorting the alleles). When sister species fix different alleles to each other the phylogenetic relationship of the genes is different from the phylogenetic relationship of the species.
- Maternal effect lethals
-
Loci in which the maternal genotype determines the viability of the zygote.
- Meiotic drive
-
Any process that causes a given allele to be overrepresented in the gametes following meiosis. Most commonly, the term is restricted to cases in which the distorted segregation ratios affect whole chromosomes rather than just a particular chromosomal location.
- Neofunctionalization
-
Evolution of a novel function, which may exist alongside an ancestral function or replace an ancestral function.
- Open chromatin
-
Decondensed chromosomal structure associated with gene expression.
- Orthologues
-
Homologous genes that diverged following a speciation event.
- Paralogues
-
Homologous genes that diverged following a gene duplication event; duplicated genes.
- Phylostratigraphic approach
-
An approach for estimating gene age based on its phylogenetic distribution. Commonly, genes are inferred to have been present in the common ancestor of any organisms in which they are detectable by sequence similarity search (such as BLAST), and their origin is assigned to the branch on the tree just prior to the node corresponding to that common ancestor. The term is a portmanteau of phylogenetics and stratigraphy, the latter being the study and dating of rock layers.
- Purifying selection
-
(Also known as negative selection). Removal of deleterious mutations from a population by selection.
- Red Queen co-evolution
-
Named after the Red Queen in Lewis Carroll's Alice Through the Looking Glass who is continually running to stay in the same place. This describes an evolutionary scenario in which two interacting loci (often one from a parasite and one from a host) are both rapidly evolving but the relationship (the interaction) has no qualitative change.
- Selective sweeps
-
Positive selection on a DNA mutation that incidentally carries closely linked variation to high frequency, thus reducing the genetic diversity in the surrounding region of the genome.
- Site frequency spectrum
-
Distribution of allele frequencies at a set of loci. The shape of the distribution can be used to infer demography and natural selection (for example, through hitchhiking).
- Spurious gene expression
-
Gene expression with no selective advantage. A necessary concept but one that in practice is hard to demonstrate, not least because the strength of selective effects relevant to the evolutionary process is typically more subtle than the effects measured in the laboratory. Selective advantage may be defined with respect to the bearer genome or to a selfish element.
- Subfunctionalization
-
Partitioning of functions of an ancestral, multifunctional gene between daughter paralogues.
- Synteny
-
Meaning 'same chromosome', this describes the physical genetic linkage of two or more loci on a chromosome. A region of shared synteny between genomes (where the orthologous genes have an equivalent relative location) is indicative of genome arrangement conservation since their most recent common ancestor. In the context of de novo genes, the syntenic location in the ancestral genome is the expected location of origin of the gene.
Rights and permissions
About this article
Cite this article
McLysaght, A., Hurst, L. Open questions in the study of de novo genes: what, how and why. Nat Rev Genet 17, 567–578 (2016). https://doi.org/10.1038/nrg.2016.78
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrg.2016.78
This article is cited by
-
The origin and structural evolution of de novo genes in Drosophila
Nature Communications (2024)
-
ADAMTS9-AS1 inhibits tumor growth and drug resistance in clear cell renal cell carcinoma via recruiting HuR to enhance ADAMTS9 mRNA stability
Cancer Nanotechnology (2023)
-
Evolution of termination codons of proteins and the TAG-TGA paradox
Scientific Reports (2023)
-
Pan-cancer surveys indicate cell cycle-related roles of primate-specific genes in tumors and embryonic cerebrum
Genome Biology (2022)
-
The Origins and Functions of De Novo Genes: Against All Odds?
Journal of Molecular Evolution (2022)