Key Points
-
A large number of tools are available for the simulation of genomic data for all current next-generation sequencing (NGS) platforms, with partially overlapped functionality. Here we review 23 of these tools, highlighting their distinct functionalities, requirements and potential applications.
-
The parameterization of these simulators is often complex. The user may decide between using existing sets of parametric values called profiles or re-estimating them from their own data.
-
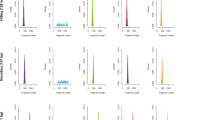
Parameters that can be modulated in these simulations include the effects of the PCR amplification of the libraries, read features and quality scores, base-calling errors, variation of sequencing depth across the genomes and the introduction of genomic variants.
-
Several types of genomic variants can be introduced in the simulated reads, such as single-nucleotide polymorphisms, insertions and deletions, inversions, translocations, copy-number variants and short-tandem repeats.
-
Reads can be generated from single or multiple genomes, and with distinct ploidy levels. NGS data from metagenomic communities can be simulated when given an 'abundance profile' that reflects the proportion of taxa in a given sample.
-
Many of the simulators have not been formally described and/or tested in dedicated publications. We encourage the formal publication of these tools and the realization of comprehensive, comparative benchmarking processes.
-
Choosing among the different genomic NGS simulators is not easy. Here, we provide a decision tree to help users choose a suitable tool for their specific interests.
Abstract
Computer simulation of genomic data has become increasingly popular for assessing and validating biological models or for gaining an understanding of specific data sets. Several computational tools for the simulation of next-generation sequencing (NGS) data have been developed in recent years, which could be used to compare existing and new NGS analytical pipelines. Here we review 23 of these tools, highlighting their distinct functionality, requirements and potential applications. We also provide a decision tree for the informed selection of an appropriate NGS simulation tool for the specific question at hand.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Metzker, M. L. Sequencing technologies — the next generation. Nat. Rev. Genet. 11, 31–46 (2010).
Nielsen, R., Paul, J. S., Albrechtsen, A. & Song, Y. S. Genotype and SNP calling from next-generation sequencing data. Nat. Rev. Genet. 12, 443–451 (2011).
Koboldt, D. C., Steinberg, K. M., Larson, D. E., Wilson, R. K. & Mardis, E. R. The next-generation sequencing revolution and its impact on genomics. Cell 155, 27–38 (2013).
Wang, X. V., Blades, N., Ding, J., Sultana, R. & Parmigiani, G. Estimation of sequencing error rates in short reads. BMC Bioinformatics 13, 185 (2012).
Liu, L. et al. Comparison of next-generation sequencing systems. J. Biomed. Biotechnol. 2012, 1–11 (2012).
Holtgrewe, M. Mason — a read simulator for second generation sequencing data. http://publications.mi.fu-berlin.de/962 (FU Berlin, 2010).
Angly, F. E., Willner, D., Rohwer, F., Hugenholtz, P. & Tyson, G. W. Grinder: a versatile amplicon and shotgun sequence simulator. Nucleic Acids Res. 40, e94 (2012).
Huang, W., Li, L., Myers, J. R. & Marth, G. T. ART: a next-generation sequencing read simulator. Bioinformatics 28, 593–594 (2012). This paper describes probably the most popular NGS simulator nowadays, with well-supported and detailed documentation.
Hu, X. et al. pIRS: profile-based Illumina pair-end reads simulator. Bioinformatics 28, 1533–1535 (2012).
Caboche, S., Audebert, C., Lemoine, Y. & Hot, D. Comparison of mapping algorithms used in high-throughput sequencing: application to Ion Torrent data. BMC Genomics 15, 264 (2014).
Hoban, S., Bertorelle, G. & Gaggiotti, O. E. Computer simulations: tools for population and evolutionary genetics. Nat. Rev. Genet. 13, 110–122 (2012).
Shendure, J. & Aiden, E. L. The expanding scope of DNA sequencing. Nat. Biotechnol. 30, 1084–1094 (2012).
Shcherbina, A. FASTQSim: platform-independent data characterization and in silico read generation for NGS datasets. BMC Res. Notes 7, 533 (2014).
Knudsen, B., Forsberg, R. & Miyamoto, M. M. A computer simulator for assessing different challenges and strategies of de novo sequence assembly. Genes 1, 263–282 (2010).
Mavromatis, K. et al. Use of simulated data sets to evaluate the fidelity of metagenomic processing methods. Nat. Methods 4, 495–500 (2007). This paper describes the use of NGS simulations for benchmarking NGS analytical methods.
McElroy, K. E., Luciani, F. & Thomas, T. GemSIM: general, error-model based simulator of next-generation sequencing data. BMC Genomics 13, 74 (2012).
Pattnaik, S., Gupta, S., Rao, A. A. & Panda, B. SInC: an accurate and fast error-model based simulator for SNPs, indels and CNVs coupled with a read generator for short-read sequence data. BMC Bioinformatics 15, 40 (2014).
Rothberg, J. M. et al. An integrated semiconductor device enabling non-optical genome sequencing. Nature 475, 348–352 (2011).
Eid, J. et al. Real-time DNA sequencing from single polymerase molecules. Science 323, 133–138 (2009).
Shendure, J. & Ji, H. Next-generation DNA sequencing. Nat. Biotechnol. 26, 1135–1145 (2008).
Shendure, J., Mitra, R. D., Varma, C. & Church, G. M. Advanced sequencing technologies: methods and goals. Nat. Rev. Genet. 5, 335–344 (2004).
Quail, M. et al. A tale of three next generation sequencing platforms: comparison of Ion Torrent, Pacific Biosciences and Illumina MiSeq sequencers. BMC Genomics 13, 341 (2012).
Pratas, D., Pinho, A. J. & O. S. Rodrigues, J. M. XS: a FASTQ read simulator. BMC Res. Notes 7, 40 (2014).
Lee, H. et al. Error correction and assembly complexity of single molecule sequencing reads. bioRxiv http://dx.doi.org/10.1101/006395 (2014).
Earl, D. et al. Assemblathon 1: a competitive assessment of de novo short read assembly methods. Genome Res. 21, 2224–2241 (2011).
Johnson, S., Trost, B., Long, J. R., Pittet, V. & Kusalik, A. A better sequence-read simulator program for metagenomics. BMC Bioinformatics 15, S14 (2014).
Jia, B. et al. NeSSM: a next-generation sequencing simulator for metagenomics. PLoS ONE 8, e75448 (2013).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Altschul, S. F., Gish, W., Miller, W., Myers, E. W. & Lipman, D. J. Basic local alignment search tool. J. Mol. Biol. 215, 403–410 (1990).
Li, R., Li, Y., Kristiansen, K. & Wang, J. SOAP: short oligonucleotide alignment program. Bioinformatics 24, 713–714 (2008).
Li, R. et al. SOAP2: an improved ultrafast tool for short read alignment. Bioinformatics 25, 1966–1967 (2009).
Keegan, K. P. et al. A platform-independent method for detecting errors in metagenomic sequencing data: DRISEE. PLoS Comput. Biol. 8, e1002541 (2012).
Frampton, M. & Houlston, R. Generation of artificial FASTQ files to evaluate the performance of next-generation sequencing pipelines. PLoS ONE 7, e49110 (2012).
Mardis, E. R. The impact of next-generation sequencing technology on genetics. Trends Genet. 24, 133–141 (2008).
Morozova, O. & Marra, M. A. Applications of next-generation sequencing technologies in functional genomics. Genomics 92, 255–264 (2008).
Aird, D. et al. Analyzing and minimizing PCR amplification bias in Illumina sequencing libraries. Genome Biol. 12, R18 (2011).
Haas, B. J. et al. Chimeric 16S rRNA sequence formation and detection in Sanger and 454-pyrosequenced PCR amplicons. Genome Res. 21, 494–504 (2011).
Balzer, S., Malde, K., Lanzén, A., Sharma, A. & Jonassen, I. Characteristics of 454 pyrosequencing data — enabling realistic simulation with flowsim. Bioinformatics 27, i420–i425 (2010). This paper presents one of the most popular simulators for 454 pyrosequencing long reads.
Balzer, S., Malde, K. & Jonassen, I. Systematic exploration of error sources in pyrosequencing flowgram data. Bioinformatics 27, 304–309 (2011).
Ledergerber, C. & Dessimoz, C. Base-calling for next-generation sequencing platforms. Brief. Bioinform. 12, 489–497 (2011).
Ewing, B. et al. Base-calling of automated sequencer traces using phred. I. Accuracy assessment. Genome Res. 8, 175–185 (1998).
Ewing, B. et al. Base-calling of automated sequencer traces using phred. II. Error probabilities. Genome Res. 8, 186–194 (1998).
Kao, W.-C., Stevens, K. & Song, Y. S. BayesCall: a model-based base-calling algorithm for high-throughput short-read sequencing. Genome Res. 19, 1884–1895 (2009).
Illumina. Technical note: Sequencing. Quality scores for next-generation sequencing: assessing sequencing accuracy using Phred quality scoring. Illumina http://www.illumina.com/documents/products/technotes/technote_Q-Scores.pdf (2011).
Dohm, J. C., Lottaz, C., Borodina, T. & Himmelbauer, H. Substantial biases in ultra-short read data sets from high-throughput DNA sequencing. Nucleic Acids Res. 36, e105 (2008). This paper describes the most relevant biases that affect the generation of NGS data.
Kircher, M. & Kelso, J. High-throughput DNA sequencing - concepts and limitations. BioEssays 32, 524–536 (2010).
Loman, N. J. et al. Performance comparison of benchtop high-throughput sequencing platforms. Nat. Biotechnol. 30, 434–439 (2012).
Robasky, K., Lewis, N. E. & Church, G. M. The role of replicates for error mitigation in next-generation sequencing. Nat. Rev. Genet. 15, 56–62 (2013).
Yang, X., Chockalingam, S. P. & Aluru, S. A survey of error-correction methods for next-generation sequencing. Brief. Bioinform. 14, 56–66 (2013).
Ekblom, R., Smeds, L. & Ellegren, H. Patterns of sequencing coverage bias revealed by ultra-deep sequencing of vertebrate mitochondria. BMC Genomics 15, 467 (2014).
Ono, Y., Asai, K. & Hamada, M. PBSIM: PacBio reads simulator — toward accurate genome assembly. Bioinformatics 29, 119–121 (2013). This paper presents one of the most popular simulators for the PacBio sequencing platform.
Richter, D. C., Ott, F., Auch, A. F., Schmid, R. & Huson, D. H. MetaSim — a sequencing simulator for genomics and metagenomics. PLoS ONE 3, e3373 (2008).
Margulies, M. et al. Genome sequencing in microfabricated high-density picolitre reactors. Nature 437, 376–380 (2005).
Nakamura, K. et al. Sequence-specific error profile of Illumina sequencers. Nucleic Acids Res. 39, e90 (2011).
Kwon, S., Park, S., Lee, B. & Yoon, S. In-depth analysis of interrelation between quality scores and real errors in Illumina reads. Conf. Proc. IEEE Eng. Med. Biol. Soc. 2013, 635–638 (2013).
Lander, E. S. & Waterman, M. S. Genomic mapping by fingerprinting random clones: a mathematical analysis. Genomics 2, 231–239 (1988).
Sims, D., Sudbery, I., Ilott, N. E., Heger, A. & Ponting, C. P. Sequencing depth and coverage: key considerations in genomic analyses. Nat. Rev. Genet. 15, 121–132 (2014).
Li, B. et al. Evaluation of de novo transcriptome assemblies from RNA-Seq data. Genome Biol. 15, 553 (2014).
Ross, M. G. et al. Characterizing and measuring bias in sequence data. Genome Biol. 14, R51 (2013).
Glenn, T. C. Field guide to next-generation DNA sequencers. Mol. Ecol. Resour. 11, 759–769 (2011).
Gilles, A. et al. Accuracy and quality assessment of 454 GS-FLX Titanium pyrosequencing. BMC Genomics 12, 245 (2011).
Quick, J., Quinlan, A. R. & Loman, N. J. A reference bacterial genome dataset generated on the MinION portable single-molecule nanopore sequencer. GigaScience 3, 22 (2014).
Loman, N. J., Quick, J. & Simpson, J. T. A complete bacterial genome assembled de novo using only nanopore sequencing data. bioRxiv http://dx.doi.org/10.1101/015552 (2015).
Jain, M. et al. Improved data analysis for the MinION nanopore sequencer. Nat. Methods 12, 351–356 (2015).
Laver, T. et al. Assessing the performance of the Oxford Nanopore Technologies MinION. Biomol. Detect. Quantif. 3, 1–8 (2015).
Madoui, M.-A. et al. Genome assembly using Nanopore-guided long and error-free DNA reads. BMC Genomics 16, 327 (2015).
Carneiro, M. O. et al. Pacific biosciences sequencing technology for genotyping and variation discovery in human data. BMC Genomics 13, 375 (2012).
Koren, S. et al. Hybrid error correction and de novo assembly of single-molecule sequencing reads. Nat. Biotechnol. 30, 693–700 (2012).
Salmela, L. & Rivals, E. LoRDEC: accurate and efficient long read error correction. Bioinformatics 30, 3506–3514 (2014).
Acknowledgements
This work was supported by the European Research Council (ERC-617457- PHYLOCANCER to D.P.) and the Spanish Government (research grants BFU2012-33038 and BFU2015-63774-P to D.P.; Research Personnel Training (FPI) graduate fellowship BES-2013-067181 to M.E.; and a Juan de la Cierva postdoctoral fellowship (FPDI-2013-17503 to S.R.). The authors thank two anonymous reviewers and members of the phylogenomics laboratory for their comments.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Glossary
- Coverage bias
-
A bias in the amount of reads for a particular region. For example, sequencing depth increases in regions of elevated GC content.
- Single end
-
Reads generated by single-read sequencing, which involves sequencing DNA fragments from only one end.
- Paired end
-
In paired-end sequencing, a single fragment is sequenced from both the 5′ and 3′ ends, giving rise to reads in both forward and reverse orientations, in which read one is the forward read and read two is the reverse. The sequenced fragments may be separated by a certain number of bases (depending on insert size and read length) or overlapping.
- Mate pair
-
Mate-pair sequencing means generating long-insert paired-end DNA libraries. The inserts are circularized and fragmented, and the labelled fragments (corresponding to the ends of the original DNA ligated together) are purified, ligated to another set of adapters and finally sequenced at the paired end. The resulting inserts include two DNA segments that were originally separated by 2–5 kb, facilitating mapping and assembly.
- Reference sequence
-
A particular genomic region, multiple genomic regions concatenated, a chromosome or a complete genome from which next-generation sequencing reads will be generated.
- Profile
-
A set of biological (GC content, insertions and deletions, and substitution rates) and/or technological (insert sizes, read lengths, error rates and quality scores) parameter distributions or values that will be used in a specific simulation.
- Abundance profile
-
A set of probabilities that represent the proportion of taxa within a community (and data set).
- Quality scores
-
(Also known as Phred Q scores). Predictions of the probability of an error in a base call.
- Amplicon
-
A piece of DNA or RNA resulting from a natural or artificial amplification event (for example, PCR).
- K-mers
-
The possible sub-sequences of length k that can be obtained from a given sequence.
- Coverage
-
The number of times a certain nucleotide has been sequenced.
- Base calling
-
The analysis of the information obtained from the machine sensors during next-generation sequencing and posterior prediction of the individual bases. This converts the signal into actual sequence data with quality scores.
- Homopolymers
-
Sequences of multiple identical nucleotides.
Rights and permissions
About this article
Cite this article
Escalona, M., Rocha, S. & Posada, D. A comparison of tools for the simulation of genomic next-generation sequencing data. Nat Rev Genet 17, 459–469 (2016). https://doi.org/10.1038/nrg.2016.57
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrg.2016.57
This article is cited by
-
Challenges and best practices in omics benchmarking
Nature Reviews Genetics (2024)
-
Genotyping of European Toxoplasma gondii strains by a new high-resolution next-generation sequencing-based method
European Journal of Clinical Microbiology & Infectious Diseases (2024)
-
SQANTI-SIM: a simulator of controlled transcript novelty for lrRNA-seq benchmark
Genome Biology (2023)
-
Potential persistence mechanisms of the major Anopheles gambiae species complex malaria vectors in sub-Saharan Africa: a narrative review
Malaria Journal (2023)
-
Performance evaluation of six popular short-read simulators
Heredity (2023)