Key Points
-
RNA-based measurements have the potential for application across diverse areas of human health, including disease diagnosis, prognosis and therapeutic selection. Current clinical applications include infectious diseases, cancer, transplant medicine and fetal monitoring.
-
RNA sequencing (RNA-seq) allows for the detection of a wide variety of RNA species, including mRNA, non-coding RNA, pathogen RNA, chimeric gene fusions, transcript isoforms and splice variants, and provides the capability to quantify known, pre-defined RNA species and rare RNA transcript variants within a sample. In addition to differential expression and detection of novel transcripts, RNA-seq also supports the detection of mutations and germline variation for hundreds to thousands of expressed genetic variants, facilitating assessment of allele-specific expression of these variants.
-
Circulating RNAs and small regulatory RNAs, such as microRNAs, are very stable. These RNA species are vigorously being tested for their potential as biomarkers. However, there are currently few agreed upon methods for isolation or quantitative measurements and a current lack of quality controls that can be used to test platform accuracy and sample preparation quality.
-
Analytical, bioinformatic and regulatory challenges exist, and ongoing efforts toward the establishment of benchmark standards, assay optimization for clinical conditions and demonstration of assay reproducibility are required to expand the clinical utility of RNA-seq.
Abstract
With the emergence of RNA sequencing (RNA-seq) technologies, RNA-based biomolecules hold expanded promise for their diagnostic, prognostic and therapeutic applicability in various diseases, including cancers and infectious diseases. Detection of gene fusions and differential expression of known disease-causing transcripts by RNA-seq represent some of the most immediate opportunities. However, it is the diversity of RNA species detected through RNA-seq that holds new promise for the multi-faceted clinical applicability of RNA-based measures, including the potential of extracellular RNAs as non-invasive diagnostic indicators of disease. Ongoing efforts towards the establishment of benchmark standards, assay optimization for clinical conditions and demonstration of assay reproducibility are required to expand the clinical utility of RNA-seq.
Similar content being viewed by others
Main
RNA is a dynamic and diverse biomolecule with an essential role in numerous biological processes. From a molecular diagnostic standpoint, RNA-based measurements have the potential for broad application across diverse areas of human health, including disease diagnosis, prognosis and therapeutic selection. Technological advancements have continually shaped the way that RNA-based measurements are used in the clinic (Box 1). With the evolution of next-generation sequencing (NGS) technologies, the use of RNA sequencing (RNA-seq) to investigate the vast diversity of RNA species is an obvious and exciting application, which opens up entirely new opportunities for improving diagnosis and treatment of human disease. RNA-seq provides an in-depth view of the transcriptome, detecting novel RNA transcript variation1. Beyond operating as an open platform technology, RNA-seq has a number of potential advantages over gene expression microarrays, including an increased dynamic range of expression, measurement of focal changes (such as single nucleotide variants (SNVs), insertions and deletions), detection of different transcript isoforms, splice variants and chimeric gene fusions (including previously unidentified genes and/or transcripts), and, fundamentally, it can be performed on any species. Although RNA-seq assays are now commercially available2,3, these early tests belie the considerable promise for broader applicability of RNA-seq-based clinical tests.
Here we review a selection of current and potential clinical applications of RNA-seq, focusing on differential expression, rare or fusion transcript detection and allele-specific expression, before discussing the emerging areas in pathogen detection and measurement of non-coding RNA (ncRNA) species. An overview of the clinically relevant RNA species discussed within the Review is summarized within Fig. 1, focusing on those RNA species that hold the greatest promise for directly impacting current and future clinical testing, with an understanding that the full diversity of RNA species and their putative roles are covered in other reviews4,5. In addition, we do not review NGS assays and technologies, as this topic is well reviewed and beyond our intended scope. We finish by discussing the challenges faced in translating this technology into clinical practice, including the regulatory environment and ongoing efforts to establish reference standards and the best practices for RNA-seq as a clinical test that is capable of high reproducibility, accuracy and precision.
Various RNA sequencing (RNA-seq) methodologies can be used to measure diverse, clinically relevant RNA species. Small RNA deep sequencing uses size selection to sequence various small non-coding RNAs, including microRNAs (miRNAs) and PIWI-interacting RNAs (piRNAs). Precursor RNAs can be measured using random primer amplification and oligo(dT) primers can be used to select polyadenylated transcripts. RNA-seq also allows for detection and measurement of alternative transcripts, chimeric gene fusion transcripts and viral RNA transcripts, as well as evaluation for allele-specific expression. HPV, human papillomavirus; lncRNA, long non-coding RNA; poly(A), polyadenylation; qRT-PCR, quantitative reverse transcription PCR; rRNA, ribosomal RNA; snoRNA, small nucleolar RNA; VUSs, variants of undetermined significance.
Opportunities enabled by RNA-seq
As in whole-genome and whole-exome sequencing, RNA-seq involves sequencing samples with billions of bases across tens to hundreds of millions of paired or unpaired short-reads. This vast amount of short-read RNA-seq data must be bioinformatically realigned and assembled to detect and measure expression of hundreds of thousands of RNA transcripts. Not only can RNA-seq detect underlying genomic alterations at single nucleotide resolution within expressed regions of the genome, it can also quantify expression levels and capture variation not detected at the genomic level, including the expression of alternative transcripts1,6. Similar to serial analysis of gene expression (SAGE), a predecessor tag-based sequencing method for genome-wide expression analysis7, RNA-seq allows quantification of transcripts without pre-defining the RNA targets of interest and provides improved detection of RNA splice events1,6. Unlike most historical platforms for clinical RNA measurement, such as microarrays and quantitative reverse transcription PCR (qRT-PCR), RNA-seq is fundamentally an open platform technology, allowing both quantification of known or pre-defined RNA species and the capability to detect and quantify rare and novel RNA transcript variants within a sample1. RNA-seq also has a greater dynamic range for quantifying transcript expression compared to microarray technology8, providing the potential for increased detection of variation within a sample. Overall, RNA-seq can identify thousands of differentially expressed genes, tens of thousands of differentially expressed gene isoforms and can detect mutations and germline variations for hundreds to thousands of expressed genetic variants (thus facilitating the assessment of allele-specific expression of these variants), as well as detecting chimeric gene fusions, transcript isoforms and splice variants5,9. In addition, RNA-seq can characterize previously unidentified transcripts and diverse types of ncRNAs, including microRNAs (miRNAs), PIWI-interacting RNAs (piRNAs) and tRNAs5. Indeed, the open platform of RNA-seq for detecting and measuring temporally dynamic RNA species sets the stage for considerable challenges and even more considerable opportunities associated with RNA-seq moving into the clinical test environment.
Detecting aberrant transcription in human disease
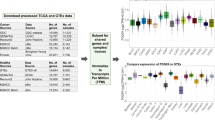
mRNA expression profiling. Multigene mRNA signature-based assays are being increasingly incorporated into clinical management. These assays use various technology platforms to measure mRNA expression of different multigene panels and have broad clinical application (Table 1). For example, in breast cancer, recent clinical guidelines support the use of multigene mRNA-based prognostic assays to assist in treatment decisions in conjunction with clinicopathological factors10,11. Indeed, the OncotypeDx 21-gene expression assay was recently validated in a prospectively conducted study in breast cancer12. Clinically relevant breast cancer gene expression signatures were compared using microarrays and RNA-seq and reported strong correlation for expression of genes from the OncotypeDx and MammoPrint signatures across platforms (Spearman correlations of 0.965 and 0.97, respectively)13. In other work14, systematic evaluation of RNA-seq-based and microarray-based classifiers found that RNA-seq outperformed arrays in characterizing the transcriptome of cancer and performed similarly to arrays in clinical endpoint prediction.
AlloMap is a non-invasive gene expression-based blood test that is used to manage the clinical care of heart transplant recipients, providing a quantified score for the risk of rejection based on the measurement of expression of 20 genes, a subset of which are related to immune system activation and signalling15,16. The potential for using RNA-seq in immune-related diseases is expanding rapidly, and the ability to quickly target and sequence the repertoire of T cell and B cell receptors from patients is beginning to mature, using techniques such as those from Adaptive Biotechnologies and ImmunoSeq. These strategies allow examination of immune-related diseases and immunotherapy response in new ways, as exemplified in a recent report in which RNA-seq and exome sequencing were used together to evaluate mutation load, expressed neoantigens and immune microenvironment expression as predictors of response to immune checkpoint inhibitor therapy in melanoma17.
Gene fusions. Oncogenic gene fusions are well recognized for their pathogenic role in cancer. In some cases, recurrent gene fusions correlate with specific tumour subtypes, allowing gene fusion status to be used for diagnostic purposes. According to the 2008 WHO (World Health Organization) classification, diagnosis of acute myeloid leukaemia (AML) can be made regardless of blast count based on detection of recurrent genetic abnormalities, such as the t(8;21)(q22;q22) translocation, RUNX1–RUNX1T1 fusion (involving isoforms of runt-related transcription factor 1 (RUNX1); this fusion is also known as AML1–ETO)18. Gene fusions have also been associated with prognosis and have been suggested as biomarkers for screening and assessment of cancer risk, as exemplified by the TMPRSS2–ERG fusion (involving transmembrane protease serine 2 gene (TMPRSS2) and v-ets avian erythroblastosis virus E26 oncogene homologue (ERG)) in prostate cancer19.
Several US Food and Drug Administration (FDA)-approved targeted agents have clinical biomarkers amenable to RNA-seq, including agents with activity against known oncogenic fusions. The prototypical example is the marked efficacy of kinase inhibitors (for example, imatinib) in BCR–ABL1-positive (involving breakpoint cluster region (BCR) and tyrosine-protein kinase ABL1 (ABL1)) chronic myeloid leukaemia (CML)20. While karyotyping is typically used for diagnosis of CML, qRT-PCR measurement of BCR–ABL1 transcripts is recommended to monitor the molecular response during kinase inhibitor treatment21. Continued advances in long-read RNA-seq technology promise to further expand the utility of fusion transcript measurements, allowing detection of mutation phasing by measuring across the full fusion transcript. Recently, single-molecule long-read RNA-seq was applied to longitudinal samples from patients with BCR–ABL1-positive CML with poor treatment response22. The results provided a clonal view of the range of resistance mutations, distinguishing between compound mutations in the same RNA molecule and independent alterations present on different molecules of the BCR–ABL1 fusion transcript. The authors reported sensitive detection that resulted in the identification of several mutations that escaped detection by routine clinical analysis and, for one of the proof-of-concept cases, detected a known drug-resistant mutation in a longitudinal patient sample four months earlier than detected by Sanger sequencing22.
Aside from monitoring BCR–ABL1 fusion transcript levels during treatment, current clinical guidelines rarely recommend RNA-based detection of gene fusions. The EML4–ALK fusion (involving echinoderm microtubule associated protein like 4 (EML4) and anaplastic lymphoma receptor tyrosine kinase (ALK)) was originally reported in a subset of non-small-cell lung cancers (NSCLC) in 2007, and the ALK inhibitors crizotinib and ceritinib gained FDA approval in ALK-rearrangement-positive NSCLC in 2011 and 2015, respectively. EML4–ALK is typically detected by fluorescence in situ hybridization (FISH) using commercial break-apart probes that flank a highly conserved translocation breakpoint in the ALK genomic locus23, with the emerging use of immunohistochemistry-based strategies to detect overexpression of ALK protein24. Recent clinical guidelines recommend against using qRT-PCR-based ALK fusion detection for treatment selection in lung cancer owing to the challenges of RNA sample quality in routine formalin-fixed paraffin-embedded (FFPE) pathology samples, as well as the risk of false negatives resulting from limitations in detecting fusions involving novel ALK fusion partners25.
In recent years, clinical detection of gene fusions has advanced beyond assays to detect individual fusion events to the introduction of RNA-seq assays, which allow a more comprehensive evaluation of potential gene fusions. For example, the FoundationOne Heme assay uses RNA-seq with genomic sequencing to detect common gene fusions in haematological cancers and sarcomas2,3. Reports are emerging of clinical responses by patients receiving treatment on the basis of gene fusion detection with this assay2,3. RNA-seq promises to expand the repertoire of detectable gene fusions, not only by capturing more subtle intrachromosomal rearrangements but also allowing detection of fusion products with uncharacterized fusion partners. Efforts are underway to catalogue gene fusions detected across various tumour types using RNA-seq data26, although additional studies are needed to define the clinical value for identified fusions.
Alternative transcripts. Alternative transcript variants, arising from splicing alterations or structural variants, have been identified and implicated in a range of human diseases, including developmental disorders27, neurodegenerative disorders28,29 and cancers30. There is also growing evidence that the presence of alternative transcripts can have therapeutic implications. For example, expression of the alternatively spliced androgen receptor variant 7 (AR-V7) has been detected in the circulating tumour cells of ~30% of men with castration-resistant prostate cancer and is associated with reduced response to androgen receptor-directed therapies31,32. Similarly, expression of the tumour-specific epidermal growth factor receptor (EGFR) variant III (EGFRvIII) transcript is well described in glioblastoma, arising from an in-frame deletion encompassing exons 2–7 (Ref. 33); clinical trials targeting EGFRvIII are underway34. In some cases, the mechanisms contributing to the generation of alternative transcripts may be missed by exome sequencing. Alternative breast cancer 1 (BRCA1) transcripts have been identified in a subset of patients with breast cancer who have a family history of breast and/or ovarian cancer35. Notably, these patients had previously tested negative for pathogenic BRCA1 or BRCA2 mutations by conventional genomic analysis35. It is anticipated that RNA-seq data will provide a more complete view of altered splicing and disease-specific transcripts, and that the growing body of transcriptome data will be a rich resource for discovery of disease-specific isoform transcripts as potential diagnostic markers and therapeutic targets.
Allele-specific expression. Allele-specific expression (ASE) can arise through multiple mechanisms, including genetic imprinting, X-chromosome inactivation and allele-specific transcription36; in some cases, ASE has been associated with predisposition to disease37,38. One of the most common mechanisms for ASE is genomic imprinting, whereby one allele is silenced through DNA methylation and histone modifications, leaving biased expression of the transcribed single nucleotide polymorphisms (SNPs) in a parent-of-origin specific manner. Imprinted gene clusters are frequently associated with human disease, as disease syndromes can arise from alterations on the single non-silenced, parental allele. For example, Angelman syndrome, a neurogenetic disorder associated with intellectual disability, speech impairment and a risk of seizures, is a well-studied imprinting disorder caused by deficient maternal allele expression of ubiquitin protein ligase E3A (UBE3A) in the brain39. In addition to epigenetic and transcriptional regulation, post-transcriptional mechanisms such as alternative splicing and protein-truncating mutations can also contribute to ASE40. For example, variants affecting splicing can cause exons to be skipped leading to ASE of variants contained within the exon41; likewise, a premature stop codon can lead to nonsense-mediated decay of one allele, resulting in ASE of the other42,43.
Evaluating ASE within RNA-seq data can inform our understanding of regulatory variation and aid in the functional interpretation of genetic variants44. Initial applications of ASE-RNA-seq focused on genomic regions that contribute to variation in transcript expression levels, termed expression quantitative trait loci, looking at Nigerian individuals from the International HapMap project. RNA-seq of lymphoblastoid cell lines derived from these individuals, coupled with the corresponding genotypes from the HapMap project, resulted in the identification of over 1,000 genes for which genetic variation influenced transcript levels or splicing and showed high concordance between polymorphisms located near genes and ASE45. More recently, as part of the Genotype–Tissue Expression (GTEx) Program, RNA-seq is being carried out on samples from a broad range of tissues from hundreds of post-mortem donors to, in part, examine the influence of genetic variation on gene expression. By analysing ASE in the pilot GTEx data, the effects of truncating mutations on nonsense-mediated transcript decay were characterized, demonstrating the utility of using RNA-seq and ASE analysis to aid in the functional interpretation of genetic variants at the DNA level43.
Given that some events may be difficult to detect or predict, ASE can be an important correlative biomarker towards identifying a pathogenically relevant genetic variant. Overexpression of a mutant allele may indicate the presence of an otherwise unidentified promoter mutation or intronic variant impacting splicing. For example, ASE analysis of transforming growth factor beta receptor 1 (TGFBR1) has been observed and associated with an increased risk of colon cancer, even though the mechanism for ASE has not been identified37. Within our own work, we used RNA-seq and ASE analysis to characterize both the chromosomal parent-of-origin and the extent of X-inactivation in a female child with mild cognitive impairment46. By performing family-trio (child and both parents) whole-exome sequencing and RNA-seq, we defined a de novo heterozygous deletion encompassing 1.6 kb on chromosome X, which contained several genes associated with neurological dysfunction; using SNPs as phasing markers, we demonstrated that the focal deletion was present on the paternal allele. The RNA-seq data further provided the ability to use ASE analysis to estimate skewed X-chromosome inactivation, demonstrating favoured expression of the cytogenetically normal maternal allele, which we suggested contributes to the observed modest phenotype. Notably, RNA-seq provided a unique advantage over the traditional Humara assay, which measures DNA methylation of the androgen receptor gene on chromosome X, providing both a parent-of-origin and chromosome-wide view of X-inactivation46.
Degner et al.47 provided one of the first reports to ascertain RNA-seq read-mapping allele specificity, demonstrating a mapping bias for alleles with SNPs represented in the reference sequence, compared to that of the alternative allele, thus producing reference-biased ASE45,47. Filtering to remove biased sites resulted in an enrichment of SNPs with ASE in genes with previously reported cis-regulatory mechanisms or gene imprinting47. Measurement of ASE can also be confounded by difficulty aligning RNA split reads that harbour neighbouring SNPs and small indels, which can also lead to reference-biased ASE42. Recent reports provide recommendations for bioinformatic analysis and data processing to address these and other challenges and introduce tools for improved detection of ASE from RNA-seq data42,48.
Extracellular RNAs. The investigation of extracellular RNAs (exRNAs) in biofluids to monitor disease is a rapidly growing area of diagnostic research. exRNAs are released from all cells in the body and are protected from degradation by carriage within extracellular vesicles or association with RNA-binding proteins (RBPs) and lipoproteins49,50,51,52,160. Measurement of exRNA is appealing as a non-invasive method for monitoring disease; as biofluids are more readily accessible than tissues, more frequent longitudinal sampling can occur. Transcripts from many tissue types, including neurons from the brain, have been shown to be accessible and detectable in plasma samples53. One obvious drawback is a lack of tissue specificity, as biofluids contain exRNAs and RBPs released from many tissue types. However, the size of the organ or tissue and proximity to the biofluid can influence the RNAs detected. For example, plasma samples have high levels of transcripts released from the liver and heart, while saliva has abundant transcripts from salivary glands and the oesophagus (K.R.V.K-J., unpublished observations). More recently, researchers are addressing this challenge by testing methods to selectively pull down extracellular vesicles derived from specific tissues, such as by immunoprecipitation for specific membrane proteins (such as L1 cell adhesion molecule (L1CAM) for neuronally derived vesicles)54,55. Use of exRNAs for cancer detection has parallels to extracellular DNA in that mutations can be detected and measured in RNA transcripts, provided that the mutations are transcribed. Potential advantages of exRNAs are that there are many more copies of the RNA than the DNA (making assessment potentially more sensitive) and differences in expression level can indicate that an organ or tissue is injured or diseased, in a way that cannot be described by DNA measurements. The catalogue of exRNA contains a large number of mRNAs and a range of regulatory RNAs that can be thoroughly evaluated by RNA-seq. There is growing interest in using this non-invasive analysis of exRNAs to monitor disease, using changes in exRNAs as readouts for key disease pathways and indicators of therapeutic efficacy. Companies such as Exosome Diagnostics are developing exRNA-based Clinical Laboratory Improvement Amendments (CLIA) diagnostic tests to monitor key gene fusions (EML4–ALK) and mutations (EGFR T790M) from plasma samples56. ExoDx Lung (ALK) is the first such test, measuring EML4–ALK transcripts isolated from exosomes in plasma from patients with NSCLC. Notably, several groups have also used circulating RNA information to provide feedback about fetal health57.
The US National Center for Advancing Translational Sciences (NCATS) has recently launched the Extracellular RNA Communication Consortium58 to develop the use of exRNA as a diagnostic tool59. They have funded several groups to help develop a catalogue of exRNAs in healthy individuals and in a number of diseases60. With increasing support for exRNA research, there should be substantial gains in understanding how to best examine these biomolecules and overcome variability in detection.
Non-coding RNA species. Beyond mRNA quantification and detection of alternative transcripts, RNA-seq opens up possibilities to measure a considerable diversity of RNA species including long non-coding RNAs (lncRNAs) and various short RNA species including miRNAs and piRNAs (Table 2). Owing to their stability and regulatory role in health and disease, miRNAs have been extensively examined as potential diagnostic markers of disease. Currently, small RNA-seq of miRNAs and other targeted miRNA-array platforms have fallen short for reliable cross-platform accuracy61,62. A considerable obstacle to using small RNA-seq is the low level of validation observed across PCR and sequencing platforms. The US National Institute of Standards and Technology (NIST) has begun development of small RNA controls; external RNA controls to support the validation of assay results and improve platforms are a necessary next step for the utility of small RNA-seq and are discussed further in later sections.
Of the small regulatory RNAs, miRNAs are the best studied and have an updated, well-curated repository for sequence information: miRBase63. With increasing accessibility and popularity of small RNA-seq, there is growing interest in using this technology in other categories of regulatory RNA. However, correct alignment and categorization is hampered by the state of the small RNA databases. Some small RNA databases are well-maintained and curated (such as the Genomic tRNA Database64), whereas other databases maintain sequences that are predicted but not experimentally validated. There is also substantial sequence overlap between categories of RNA, such as piRNA, tRNA and rRNA, making downstream data analysis challenging.
There are new types of regulatory RNAs for which the diagnostic potential is unknown. Circular RNAs (circRNAs) were recently rediscovered in RNA-seq experiments searching for chromosomal rearrangements in cancer65. Although many groups are identifying new circRNAs and their potential functional roles in the cell, there are few reports of function in disease pathogenesis. However, circRNAs have been found in high abundance in biofluids and tissues and have been found to be more stable than mRNAs, increasing their potential for diagnostic purposes66. Other regulatory RNAs have had new roles identified. For example, a role for tRNAs has been reported in cancer, whereby cleavage of tRNAs produced fragments that could displace the RNA-binding protein Y-box binding protein 1 (YBX1) from oncogenic transcripts, altering stability and suppressing breast cancer growth, invasion and metastasis67. A current challenge is associating new RNA discoveries, newly identified sequence information and the emerging roles for regulatory RNAs that challenge dogma with disease and diagnosis.
RNA-seq for infectious disease diagnosis
RNA-based pathogen diagnostics. Given the large number of clinically important RNA viruses (HIV, the Ebola, West Nile, dengue, hepatitis A, hepatitis D, hepatitis E, coxsackie and influenza viruses, and the severe acute respiratory syndrome (SARS) and Middle East respiratory syndrome (MERS) coronaviruses), qRT-PCR assays have been developed and are commonly used in the clinic for viral detection and typing68,69,70,71. It is likely that many of these targets will be translated into RNA-based sequencing assays in the near future. For example, unbiased non-targeted metagenomic RNA-seq has recently been used to directly detect influenza virus RNA in respiratory fluids, with additional viral pathogens detected in a subset of cases72. In a public health context, RNA-seq was used to track the origin and transmission patterns of the Ebola virus during the 2014 outbreak in West Africa73. RNA-based amplicon sequencing is also being explored for viral quasi-species (that is, mixed allele population) assessment for hepatitis C virus (HCV) and HIV; such analyses are necessary in the clinic to determine the presence and relative quantity of drug-resistance mutations for patient therapy, which can occur as minor components in a larger viral population74,75. However, clinical application of RNA-based diagnostics for infectious disease is still rare beyond the qRT-PCR assays for viral pathogens.
Microbial exogenous small RNA. A tremendous diversity of exogenous RNAs from non-human sources has been seen in human plasma, which indicates there is a relationship between the host and the microbiome, food sources and/or the environment76,77. The sources and importance of these microbial exogenous RNAs — which may or may not be encapsulated in outer-membrane vesicles78 — are still being explored, particularly in the context of infection159. However, they hold a great deal of promise for new diagnostic targets. Extensive analysis (in vitro, ex vivo and in vivo) and cataloguing of small RNAs produced by pathogens has been under way for several years and will provide a comprehensive reference database of exogenous RNA signals that may be useful for future clinical infection studies79,80,81. For example, ex vivo studies on Neisseria meningitidis infections in human blood have yielded dozens of small RNAs that seem to be associated with bacteraemic infections82. Similarly, multiple studies of the Mycobacterium tuberculosis microRNAome have yielded numerous biomarkers that are currently being explored for diagnostic purposes83 and even for phenotype–genotype predictive diagnostics (such as for identifying the presence of multi-drug resistance)84.
Pathogen mRNA. Measurement of microbial mRNA may be a useful marker of infection, as expression may improve detection in cases of low-level infections (for example, bacteraemia and cerebrospinal fluid infections) and could act as a better predictor of disease compared to direct genomic detection. For example, the simple detection of human papilloma virus (HPV) DNA is not sufficient to diagnose HPV-related squamous cell carcinoma (HPV DNA is detectable in ~14% of healthy control women85); thus, RNA-based diagnostics to detect HPV have been developed. HPV early oncoprotein E6/E7 mRNA detection, as a surrogate for active infection, may provide a better predictive value for cervical cancer.
Host RNA. Host response, in the form of mRNA signatures, is also likely to become useful for monitoring specific infections. For example, upregulation of specific host immune factors (interferon beta 1 (IFNB1) interferon lambda 2 (IFNL2) and interferon lambda 3 (IFNL3)) was recently demonstrated for genotype 3 HCV infection, which is associated with accelerated liver fibrosis and is an independent risk factor for hepatocellular carcinoma, compared to non-genotype 3 HCV infection86,87. Host microRNA–gene interactions during the infection response are also proving to be a fruitful source of potential diagnostic biomarkers for specific infections, as well as distinguishing between active and latent infections; for example, the use of such markers to discriminate between latent infection and active disease with M. tuberculosis88. As with the other diagnostic uses of small RNA biomarkers described in this Review, a number of hurdles exist for the development and validation of infection assays, including sensitivity, specificity and, perhaps more difficult to overcome, the normalization of small RNA in clinical samples, which will vary between conditions, tissues and individual hosts and is an active area of study89. However, the application of RNA-seq provides a useful orthogonal approach to genomics-based diagnostics for clinical microbiology.
Challenges moving RNA-seq to the clinic
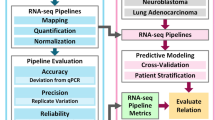
Translating an assay to the clinic. Translation and broader adoption of a laboratory test into the clinic involves evaluation and demonstration of analytical validity, clinical validity and, eventually, clinical utility90 (Fig. 2). Analytical validity generally refers to the ability of the test to measure the intended biomolecules within clinically relevant conditions. Establishment of an analytically valid test can have different meanings depending on the regulatory framework that the test falls under, as discussed below. However, analytical validity generally implies that the test has undergone thorough technical performance characterization. Clinical validity refers to the ability of a test to predict a clinical outcome given a set of events, irrespective of whether the test results can enable an effective therapy. Clinical utility indicates whether a test provides useful information, positive or negative, for the patient being tested. Tests that can either indicate a more effective therapy, such as a companion diagnostic, or provide information on avoiding some therapies may both have clinical utility.
Before initial clinical introduction, a clinical test must demonstrate analytical validity, showing sufficient assay performance to produce accurate and reproducible technical results. Demonstration of analytical validity involves several measures, including sensitivity (true technical positives), specificity (true technical negatives), robustness and limits of detection. Clinical validity follows analytical validity and, depending on the approval path, demonstration of clinical validity can come before (US Food and Drug Administration (FDA) in vitro diagnostic device) or after (Clinical Laboratory Improvement Amendments (CLIA) laboratory-developed test) test clearance or approval. Clinical validity refers to the concordance between the test result and the clinical diagnosis or outcome and involves measures of sensitivity (true clinical positives) and specificity (true clinical negatives), as well as determination of positive and negative predictive values. Demonstration of both analytical validity and clinical validity occurs before that of clinical utility. Clinical utility requires clinical evidence that use of the test has an impact on patient care and includes evaluation of patient outcomes and the economic benefits associated with the test. ELSI, National Human Genome Research Institute's Ethical, Legal and Social Implications Research Program.
Performance metrics and reference standards. To be analytically valid, a laboratory test must deliver accurate information with reproducible and robust performance. Accuracy is determined by evaluating a measured or calculated value compared to a reference 'gold standard', with evaluation of sensitivity (ability to detect true positives) and specificity (ability to detect true negatives). The test must also provide the same or similar results with repeat testing (reproducibility) and withstand small, deliberate changes in pre-analytic or analytic variables associated with testing (robustness).
Establishing reference standards and the best practices for measuring RNA-seq accuracy, reproducibility and robustness has initially been ad hoc with individual groups providing the initial steps. In 2008, Marioni and colleagues8 provided some of the earliest technical assessments of reproducibility for measuring gene expression levels by RNA-seq, reporting high reproducibility across technical replicates for a single RNA sample. Of the genes denoted significantly differentially expressed by microarray analysis, 81% were also differentially expressed with RNA-seq, with fold-change correlations between the two technologies similar or better than those reported in comparisons of different microarray platforms8. RNA-seq detected ~30% more differentially expressed genes than microarray analysis, with qRT-PCR confirmation for a subset, suggesting a large proportion may represent true positives and this difference may be a result of the broader dynamic range of RNA-seq and/or the ability to resolve splicing changes8. In 2013, the Genetic European Variation in Disease (GEUVADIS) consortium demonstrated the feasibility and reproducibility of performing RNA-seq across multiple laboratories, sequencing lymphoblastoid cell lines for 465 individuals, in seven sequencing centres using a single platform91. On the basis of this study, the consortium proposed a set of quality checks to assess technical biases in RNA-seq data, including differences in GC content, fragment size, transcript length and the percentage of reads mapped to annotated exons91.
Although providing necessary guidance on RNA-seq assay metrics, early assessments often used collection methods that may not reflect the conditions observed within the clinic. For example, within oncology, most studies of transcriptome analysis by RNA-seq come from fresh-frozen tumour samples that are stringently collected in terms of cellularity, tumour necrosis and RNA quality, whereas most pathological samples are collected through formalin fixation to preserve the protein and cellular structure. As RNA-seq libraries are typically prepared from total RNA using polyadenylation (poly(A)) enrichment of mRNAs, this method does not adequately capture partially degraded mRNAs, as are found in FFPE samples. Given the clear challenge of low-quality RNA from clinical FFPE samples, concerted effort has focused on optimization and evaluation of protocol modifications, including rRNA depletion (ribo-depletion) protocols to remove rRNA without poly(A) enrichment92 and the use of capture sequencing, such as with oligonucleotide probe hybridization93. The utility of cDNA-Capture sequencing (exome capture and RNA-seq) was demonstrated for differential gene expression analysis from FFPE samples94. In addition to differential expression, this capture protocol was recently applied in the Michigan Oncology Sequencing Center (MI-ONCOSEQ) clinical sequencing programme, which demonstrated that the use of capture libraries improved performance for fusion and splice junction detection, compared to typical poly(A)-enriched RNA-seq using low-quality RNA (FFPE) samples95. Analytical tools are also being developed and evaluated for their effect on test performance. In a comparison study of several common software tools (Cufflinks-Cuffdiff2, DESeq and edgeR) used to analyse differential expression by RNA-seq, using qRT-PCR and microarray results as benchmark standards, Zhang and colleagues96 found that each tool had different strengths and recommended an ensemble-based approach combining two or more tools to reduce false-positives.
Although these early efforts advanced the field and indeed proposed reference standards and best practices, the most substantial large-scale efforts have mainly emerged in the past few years and include consortium members spanning public regulatory bodies (such as the FDA, NIST and the Centers for Disease Control and Prevention), academic groups and industry. One of the first sets of reference standards for RNA-seq was the development of synthetic RNA spike-in controls by the External RNA Controls Consortium (ERCC)97. The ERCC RNA spike-in controls contain 92 polyadenylated transcripts pre-formulated into two sets (Mix1 and Mix2), each with the full complement of transcripts spanning approximately 106-fold concentration range, present at defined Mix1/Mix2 molar ratios in four subgroups. These stock solutions can be diluted and added to each RNA sample, thus providing a post hoc measurement of assay performance from a set of synthetic transcripts98. Synthetic gene-fusion spike-ins, composed of polyadenylated RNA transcripts corresponding to known oncogenic gene fusions, are also available with corresponding RNA-seq data99. In late 2014, the Sequencing Quality Control (SEQC) project (Phase III of the Microarray Quality Control (MAQC) Consortium)100 and the Association of Biomolecular Resource Facilities (ABRF)101 independently reported on large-scale efforts to define the performance characteristics of RNA-seq. Both studies represented remarkable coordinated analysis of inter-sample, cross-platform and inter-site variability for RNA-seq, compared to gold-standard methods such as microarrays and qRT-PCR, with extensive use of ERCC RNA spike-in controls. Both studies extensively examined how assay differences such as poly(A)-enrichment selection or random priming plus ribo-deletion affect reproducibility. Specifically, the FDA-led SEQC study examined the reproducibility and accuracy of RNA-seq in a multi-site study using different sequencing platforms. The SEQC study found that the correlation of relative gene expression measurements between different RNA-seq platforms (SOLiD, Life Technologies; HiSeq 2000, Illumina) and Affymetrix HuGene U133 Plus 2.0 microarrays with TaqMan qRT-PCR was high (Spearman and Pearson correlation coefficients of >0.9), although none of the platforms provided accurate absolute quantification of transcript levels, based on evaluation of ERCC spike-in control titration values98. Sensitivity, specificity and reproducibility of differential expression calls across sites were dependent on the analysis pipeline and the use of filters. Applying filters for P value, fold-change and expression level improved the false discovery rate for differential gene expression analysis, with most pipelines showing high reproducibility for differential expression calls across sites and greatest concordance for the most highly expressed genes. Consistent with previous reports, the SEQC study found that RNA-seq was seen as fundamentally superior at detecting novel transcripts, such as those resulting from alternative splicing, validating over 80% by qRT-PCR100. The ABRF study examined intra- and inter-laboratory reproducibility at 15 different laboratories, comparing different library preparation methods, sample variables (RNA integrity, size-specific fractionation), analysis algorithms and sequencing platforms. The ABRF study reported an overall high concordance for normalized gene expression measures within platforms (Spearman correlation >0.86) and between platforms (Spearman correlation >0.83). Deficiencies in cross-platform detection were identified and associated with read-length, analysis approaches and technology differences101. Similar to previous reports, library preparation methods influenced transcript enrichment, with poly(A) libraries containing more exonic reads and ribo-depletion libraries containing more intronic reads. Notably, although differences were observed, differential gene expression results were comparable between poly(A) enrichment and ribo-depletion library preparation methods, as well as between degraded and non-degraded RNA samples101.
The importance of these studies goes well beyond the initial reporting, as companion papers helped to frame the specific areas where analytical validity can be substantially improved. Particularly relevant to clinical specimens, a concordance between RNA-seq and microarray was reported for 27 different chemical treatments, showing that differentially expressed genes correlated with the effect size of the treatment given and, whereas both platforms showed high concordance with qRT-PCR data for highly expressed genes, RNA-seq showed higher concordance than microarrays for genes with low expression102. Furthermore, multiple groups have examined the role of normalization methods towards the controlling biases resulting from GC content, sequencing coverage and insert size103,104.
Analysis paralysis and other bioinformatic challenges. The SEQC–MAQC100 and ABRF101 consortium papers identified numerous substantial bioinformatic challenges that must be addressed for RNA-seq to become broadly adopted into clinical laboratories. Recognizing that excellent bioinformatics reviews are available elsewhere105, here we attempt only to highlight the major overarching bioinformatics challenges. We find three large themes that frequently contribute to 'analysis paralysis' during the development of bioinformatics solutions to RNA-seq: first is the lack of consensus by governing bodies advocating best practices and reference standards for validating RNA-seq pipelines; second is the overabundance of software tools, options and combinations thereof for RNA-seq analysis; and third are highly complex pipelines consisting of chaining together multiple tools that are independently developed, maintained and licensed.
First and probably most relevant, RNA-seq analysis has largely grown organically without extensive standards or dominating governing bodies. By comparison, standards were established early on for DNA-based NGS by the 1,000 Genomes Project, including variant call format (VCF), binary alignment/map format (BAM format) and genotype likelihoods, essentially providing the 'best practice approaches' (Refs 106,107,108,109,110,111). Before 2014, reference standards, ERCC spike-in controls and the general MAQC were major contributors towards building a reproducible RNA-seq pipeline, but they pale in comparison to having a consensus germline-genomic reference standard, such as the NA12878 human reference genome112. As a consequence of this lack of standards, RNA-seq provided fertile grounds for the emergence of a multitude of software tools and other options, all competing for relevance as part of a broad range of RNA-seq solutions. RNA-seq pipelines represent the laboratory-specific wrapper scripts, chaining together collections of software tools with the goal of reporting hundreds to thousands of test results from gigabases of data. Unfortunately, having RNA-seq pipelines composed of several independently developed components, each with continual versioning and variable licenses, can be challenging in a clinical testing laboratory. Although excellent for agile software development in a non-regulated and rapidly changing field, this type of fertile environment creates major challenges in a clinical laboratory. For example, whereas genome builds change every few years, transcript definitions often change quarterly.
Although the fundamental challenges of bioinformatics are unlikely to be easily solved, the framework for how they are managed has improved considerably. With the ERCC97, SEQC–MAQC100 and ABRF101 consortia providing initial models and reference standards, the emergence of 'bake-offs' and 'best practices' will be an essential next step in reducing some of the bioinformatics challenges of clinical RNA-seq. Moreover, the emergence of tools that allow containerization, such as Docker113, provide a platform for distributing fully contained pipelines such that a pipeline run in one facility could be reproducibly deployed in another laboratory. Some full-packaged pipelines have recently emerged with the expressed intent of being relevant for clinical applications114,115.
Regulatory considerations. Deployment of a clinical assay in the United States involves two paths, each with accompanying regulations administered under the US Department of Health and Human Services (HHS). The first are those approved through the CLIA of 1988 that allow 'laboratory-developed tests' (LDTs). LDTs are in vitro diagnostic tests that are developed and used within a single approved laboratory and are not marketed towards any other laboratory. CLIA regulations monitor the laboratory process to ensure the accuracy, reliability and appropriateness of laboratory testing, from sample acquisition, handling and storage to the interpretation and reporting of test results. The guidelines for approving CLIA laboratories are established by accredited professional organizations, such as the College of American Pathologists or by other agencies approved by the Centers for Medicare and Medicaid Services (CMS), such as in the state of New York. CLIA regulations of LDTs do not address the clinical validity or clinical utility of an assay, but instead provide a framework whereby clinical laboratories validate analytical performance measures of the LDTs within their own laboratory facility. The second set of regulations for clinical assay deployment are the Medical Device Amendments of 1976, which expanded FDA oversight for the marketing of in vitro diagnostic devices (IVDs). FDA premarket review of IVDs assures the assay has established analytical and clinical validity; with the exception of companion diagnostics, the FDA does not typically require demonstration of clinical utility for clearance or approval of IVDs. Demonstration of clinical validity (for LDTs) and clinical utility (for LDTs and IVDs) can follow the initial clearance or approval of a diagnostic test; clinical utility, in particular, requires broader clinical evaluation across multiple sites and/or within clinical trials. For example, the Afirma (Veracyte) microarray-based gene expression classifier for thyroid nodule assessment was launched in 2011 as a CLIA-regulated LDT. Subsequent studies have reported on clinical validity116 and clinical utility117, the latter involving a multi-site study that demonstrated the effects of the Afirma test on clinical care recommendations, which resulted in a reduction of unnecessary surgeries117.
In the US, the distinction over when NGS assays are under regulatory oversight by the FDA or the CMS is emerging as an area of regulatory and legislative debate. In late 2014, the FDA proposed a regulatory framework for LDTs118,119 that will, in all likelihood, alter the regulatory landscape discussion for RNA-seq assays moving to the clinic. The FDA also provided a perspective on the mammoth shifts created by technological advances associated with NGS, and the requirement for the agency to change from the current 'general enforcement discretion' — in which the FDA has generally not enforced regulations with respect to LDTs — to having a more active role, with proposed premarket review and quality system regulation requirements120. Under this proposed LDT framework, the CMS (under CLIA) would oversee the laboratory operations and testing processes and the FDA would monitor compliance with quality system regulations.
The effects of expanding regulatory oversight by the FDA on RNA-seq are predicated around the FDA approval process for the first FDA-cleared NGS instrument and NGS in vitro diagnostic tests, the Illumina MiSeqDx and the associated in vitro diagnostic assays for cystic fibrosis, the Illumina MiSeqDx Cystic Fibrosis 139-Variant and Cystic Fibrosis Clinical Sequencing Assays. Accuracy was evaluated using a representative subset of variants, rather than evaluating all possible variants, and relied on publicly available data to support clinical relevance of the variants. Although evaluation of analytical performance may continue to involve this subset-based approach, the proposed new standards, as outlined by the FDA120, could include defined technical metrics for data quality, additional standards for computational approaches and standard best practices for quality assurance. The debate over FDA oversight is largely focused on the presence or absence detection of DNA variants, such as germline cystic fibrosis transmembrane conductance regulator (CFTR) or BRCA2 testing. While the FDA guidance and debate is limited in use of examples, the broad scope of additional regulation on all NGS-developed tests, including RNA-seq, may provide regulatory uncertainty for RNA-seq and impede its adoption in the clinic. The proposed FDA regulations around NGS have not gone without debate, emphasizing that the limited enforcement capabilities and regulatory guidance could unnecessarily stifle adoption and innovation121. International regulatory frameworks vary across jurisdictions122,123, with evolving practice guidelines and regulations for the clinical use of NGS118,119,123,124. For example, in the European Union (EU), IVD tests require a Conformité Européenne (CE) mark to indicate compliance with the EU IVD Directive (98/79/EC). Similar to the US, the EU is reviewing policy changes related to IVDs, with proposed changes to harmonize the IVD market and increase oversight, including the use of a risk-based classification scheme to define clinical evidence requirements, such as analytical and clinical performance, for IVD approval125. The pending regulatory changes, both in the US and internationally, may substantially impact the clinical utility of RNA, particularly until greater consensus is reached towards reference standards.
Integrating DNA and RNA sequencing
In clinical studies, integrative DNA and RNA analysis has provided additional evidence for dysregulation of mutated genes, as well as detection of gene fusions and splicing variants and, in some cases, helped prioritize variants for therapeutic intervention (Box 2). In addition to associating specific genomic alterations with potential therapeutic response, exciting and emerging work suggests that integrated sequencing strategies may also aid in the identification of patient-specific immunogenic neoantigens expressed in the tumour. Recently, RNA-seq and exome sequencing were used to identify and filter for predicted immunogenic neoantigens that were expressed in melanoma tumours, demonstrating an association between these expressed neoantigens and response to the immune-checkpoint inhibitor ipilimumab17.
Integration of DNA sequencing and RNA-seq holds promise beyond oncology. For example, in the transplant field, reports are emerging for the utility of circulating DNA in monitoring for transplant rejection129,130; as discussed earlier, RNA-based measures are already used for the early detection of rejection in heart transplant recipients. The integration of RNA and DNA sequencing to improve transplant rejection diagnosis is an important area currently being examined. The combination of genotyping circulating DNA from donor and recipient, and assessing changes in expression level may provide insights for the degree of rejection, the molecular mechanisms underlying rejection and could suggest possible therapeutic strategies to keep the transplant viable. In the same way, integration of DNA and RNA sequencing could benefit fetal medicine. DNA sequencing of the fetus from maternal blood, in combination with changes in transcript expression levels, could provide additional accuracy and insight for assessing developmental complications.
Although challenges exist, the demonstrated utility of integrative sequencing strategies in research studies is growing across broad health applications and points to the promise for incorporation of RNA-seq into clinical medicine.
Conclusion and perspectives
With its unprecedented ability to simultaneously detect global gene transcript levels and diverse RNA species, RNA-seq has the potential to revolutionize clinical testing for a wide range of diseases. Although recent efforts have set the stage for the establishment of benchmark standards for technical and analytical best practices in order to better standardize RNA-seq accuracy, reproducibility and precision, additional steps toward test proficiency and validation will be required to expand the utility of RNA-seq.
Conceptually, RNA-seq is shot-gun sequencing of the transcriptome, lending to both potential utility and considerable hurdles towards translating RNA to the clinic. The dynamic range and expanding approaches for sample preparation and analysis allow for incredible fine-tuning of sensitivity, specificity and reproducibility. To some extent, the same flexibility and seemingly infinite set of options for RNA-seq that has spurred incredible discoveries into the dynamic nature of human disease has also hindered its path to the clinic. In particular, the establishment of standards has lagged until recently. Several paths forward exist and it is likely that many will be taken towards the direct use of RNA-seq in clinical applications. Once the discovery phase is complete, many diagnostic tests will become targeted assays, sensitive enough to detect small numbers of rare transcripts. The fixed nature of probe sets with microarrays or qRT-PCR offer an accelerated path for clinical test development, as 'the data are the data' without the lure of the latest and newest analysis methods. Therefore, although RNA-seq as a platform has great promise, continuing studies are needed to demonstrate analytical validity and facilitate its adoption within the clinical laboratory setting.
References
Mortazavi, A., Williams, B. A., McCue, K., Schaeffer, L. & Wold, B. Mapping and quantifying mammalian transcriptomes by RNA-Seq. Nat. Methods 5, 621–628 (2008).
Doebele, R. C. et al. An oncogenic NTRK fusion in a soft tissue sarcoma patient with response to the tropomyosin-related kinase (TRK) inhibitor LOXO-101. Cancer Discov. 5, 1049–1057 (2015).
Sonu, R. J., Jonas, B. A., Dwyre, D. M., Gregg, J. P. & Rashidi, H. H. Optimal molecular methods in detecting p190 (BCR-ABL) fusion variants in hematologic malignancies: a case report and review of the literature. Case Rep. Hematol. 2015, 458052 (2015).
Cech, T. R. & Steitz, J. A. The noncoding RNA revolution-trashing old rules to forge new ones. Cell 157, 77–94 (2014).
Wang, Z., Gerstein, M. & Snyder, M. RNA-Seq: a revolutionary tool for transcriptomics. Nat. Rev. Genet. 10, 57–63 (2009).
Wilhelm, B. T. et al. Dynamic repertoire of a eukaryotic transcriptome surveyed at single-nucleotide resolution. Nature 453, 1239–1243 (2008). This is one of the earliest applications of RNA-seq indicating diagnostic potential for detecting and quantifying various RNA species including mRNA, alternative transcripts and non-coding RNA.
Velculescu, V. E., Zhang, L., Vogelstein, B. & Kinzler, K. W. Serial analysis of gene expression. Science 270, 484–487 (1995). This paper provides an early description of multiplexed RNA detection that helped set the stage for use of microarray and spotted arrays within diagnostics.
Marioni, J. C., Mason, C. E., Mane, S. M., Stephens, M. & Gilad, Y. RNA-seq: an assessment of technical reproducibility and comparison with gene expression arrays. Genome Res. 18, 1509–1517 (2008).
Sultan, M. et al. A global view of gene activity and alternative splicing by deep sequencing of the human transcriptome. Science 321, 956–960 (2008).
Senkus, E. et al. Primary breast cancer: ESMO Clinical Practice Guidelines for diagnosis, treatment and follow-up. Ann. Oncol. 26, v8–v30 (2015).
Coates, A. S. et al. Tailoring therapies-improving the management of early breast cancer: St Gallen International Expert Consensus on the Primary Therapy of Early Breast Cancer 2015. Ann. Oncol. 26, 1533–1546 (2015).
Sparano, J. A. et al. Prospective validation of a 21-gene expression assay in breast cancer. N. Engl. J. Med. 373, 2005–2014 (2015). This paper helps establish clinical validity for a prognostic signature for endocrine therapy alone for patients with hormone-receptor-positive, ERBB2-negative, axillary node-negative breast cancer.
Fumagalli, D. et al. Transfer of clinically relevant gene expression signatures in breast cancer: from Affymetrix microarray to Illumina RNA-Sequencing technology. BMC Genomics 15, 1008 (2014).
Zhang, W. et al. Comparison of RNA-seq and microarray-based models for clinical endpoint prediction. Genome Biol. 16, 133 (2015).
Deng, M. C. et al. Noninvasive discrimination of rejection in cardiac allograft recipients using gene expression profiling. Am. J. Transplant. 6, 150–160 (2006).
Starling, R. C. et al. Molecular testing in the management of cardiac transplant recipients: initial clinical experience. J. Heart Lung Transplant. 25, 1389–1395 (2006).
Van Allen, E. M. et al. Genomic correlates of response to CTLA-4 blockade in metastatic melanoma. Science 350, 207–211 (2015).
Vardiman, J. W. et al. The 2008 revision of the World Health Organization (WHO) classification of myeloid neoplasms and acute leukemia: rationale and important changes. Blood 114, 937–951 (2009).
Font-Tello, A. et al. Association of ERG and TMPRSS2-ERG with grade, stage, and prognosis of prostate cancer is dependent on their expression levels. Prostate 75, 1216–1226 (2015).
Druker, B. J. et al. Efficacy and safety of a specific inhibitor of the BCR-ABL tyrosine kinase in chronic myeloid leukemia. N. Engl. J. Med. 344, 1031–1037 (2001).
O'Brien, S. et al. Chronic Myelogenous Leukemia, version 1.2014. J. Natl Compr. Canc. Netw. 11, 1327–1340 (2013).
Cavelier, L. et al. Clonal distribution of BCR-ABL1 mutations and splice isoforms by single-molecule long-read RNA sequencing. BMC Cancer 15, 45 (2015).
Leighl, N. B. et al. Molecular testing for selection of patients with lung cancer for epidermal growth factor receptor and anaplastic lymphoma kinase tyrosine kinase inhibitors: American Society of Clinical Oncology endorsement of the College of American Pathologists/International Association for the study of lung cancer/association for molecular pathology guideline. J. Clin. Oncol. 32, 3673–3679 (2014).
Wynes, M. W. et al. An international interpretation study using the ALK IHC antibody D5F3 and a sensitive detection kit demonstrates high concordance between ALK IHC and ALK FISH and between evaluators. J. Thorac. Oncol. 9, 631–638 (2014).
Lindeman, N. I. et al. Molecular testing guideline for selection of lung cancer patients for EGFR and ALK tyrosine kinase inhibitors: guideline from the College of American Pathologists, International Association for the Study of Lung Cancer, and Association for Molecular Pathology. Arch. Pathol. Lab. Med. 137, 828–860 (2013).
Wang, Y., Wu, N., Liu, J., Wu, Z. & Dong, D. FusionCancer: a database of cancer fusion genes derived from RNA-seq data. Diagn. Pathol. 10, 131 (2015).
Magri, F. et al. Clinical and molecular characterization of a cohort of patients with novel nucleotide alterations of the Dystrophin gene detected by direct sequencing. BMC Med. Genet. 12, 37 (2011).
Liu, F. & Gong, C. X. Tau exon 10 alternative splicing and tauopathies. Mol. Neurodegener. 3, 8 (2008).
La Cognata, V., D'Agata, V., Cavalcanti, F. & Cavallaro, S. Splicing: is there an alternative contribution to Parkinson's disease? Neurogenetics 16, 245–263 (2015).
Chen, J. & Weiss, W. A. Alternative splicing in cancer: implications for biology and therapy. Oncogene 34, 1–14 (2015).
Dehm, S. M., Schmidt, L. J., Heemers, H. V., Vessella, R. L. & Tindall, D. J. Splicing of a novel androgen receptor exon generates a constitutively active androgen receptor that mediates prostate cancer therapy resistance. Cancer Res. 68, 5469–5477 (2008).
Antonarakis, E. S. et al. AR-V7 and resistance to enzalutamide and abiraterone in prostate cancer. N. Engl. J. Med. 371, 1028–1038 (2014).
Sugawa, N., Ekstrand, A. J., James, C. D. & Collins, V. P. Identical splicing of aberrant epidermal growth factor receptor transcripts from amplified rearranged genes in human glioblastomas. Proc. Natl Acad. Sci. USA 87, 8602–8606 (1990).
Reardon, D. A. et al. 107 ReACT: overall survival from a randomized Phase II study of rindopepimut (CDX-110) plus Bevacizumab in relapsed glioblastoma. Neurosurgery 62 (Suppl. 1), 198–199 (2015).
Gambino, G., Tancredi, M., Falaschi, E., Aretini, P. & Caligo, M. A. Characterization of three alternative transcripts of the BRCA1 gene in patients with breast cancer and a family history of breast and/or ovarian cancer who tested negative for pathogenic mutations. Int. J. Mol. Med. 35, 950–956 (2015).
Massah, S., Beischlag, T. V. & Prefontaine, G. G. Epigenetic events regulating monoallelic gene expression. Crit. Rev. Biochem. Mol. Biol. 50, 337–358 (2015).
Valle, L. et al. Germline allele-specific expression of TGFBR1 confers an increased risk of colorectal cancer. Science 321, 1361–1365 (2008).
de la Chapelle, A. Genetic predisposition to human disease: allele-specific expression and low-penetrance regulatory loci. Oncogene 28, 3345–3348 (2009).
Lossie, A. C. et al. Distinct phenotypes distinguish the molecular classes of Angelman syndrome. J. Med. Genet. 38, 834–845 (2001).
Li, G. et al. Identification of allele-specific alternative mRNA processing via transcriptome sequencing. Nucleic Acids Res. 40, e104 (2012).
Klimpe, S. et al. Evaluating the effect of spastin splice mutations by quantitative allele-specific expression assay. Eur. J. Neurol. 18, 99–105 (2011).
Wood, D. L. A. et al. Recommendations for accurate resolution of gene and isoform allele-specific expression in RNA-seq data. PLoS ONE 10, e0126911–e0126927 (2015).
Rivas, M. A. et al. Human genomics. Effect of predicted protein-truncating genetic variants on the human transcriptome. Science 348, 666–669 (2015).
Larson, N. B. et al. Comprehensively evaluating cis-regulatory variation in the human prostate transcriptome by using gene-level allele-specific expression. Am. J. Hum. Genet. 96, 869–882 (2015).
Pickrell, J. K. et al. Understanding mechanisms underlying human gene expression variation with RNA sequencing. Nature 464, 768–772 (2010).
Szelinger, S. et al. Characterization of X chromosome inactivation using integrated analysis of whole-exome and mRNA sequencing. PLoS ONE 9, e113036 (2014).
Degner, J. F. et al. Effect of read-mapping biases on detecting allele-specific expression from RNA-sequencing data. Bioinformatics 25, 3207–3212 (2009).
Castel, S. E., Levy-Moonshine, A., Mohammadi, P., Banks, E. & Lappalainen, T. Tools and best practices for data processing in allelic expression analysis. Genome Biol. 16, 195 (2015).
Valadi, H. et al. Exosome-mediated transfer of mRNAs and microRNAs is a novel mechanism of genetic exchange between cells. Nat. Cell Biol. 9, 654–659 (2007).
Arroyo, J. D. et al. Argonaute2 complexes carry a population of circulating microRNAs independent of vesicles in human plasma. Proc. Natl Acad. Sci. USA 108, 5003–5008 (2011).
Michael, A. et al. Exosomes from human saliva as a source of microRNA biomarkers. Oral Dis. 16, 34–38 (2010).
Skog, J. et al. Glioblastoma microvesicles transport RNA and proteins that promote tumour growth and provide diagnostic biomarkers. Nat. Cell Biol. 10, 1470–1476 (2008).
Koh, W. et al. Noninvasive in vivo monitoring of tissue-specific global gene expression in humans. Proc. Natl Acad. Sci. USA 111, 7361–7366 (2014).
Goetzl, E. J. et al. Altered lysosomal proteins in neural-derived plasma exosomes in preclinical Alzheimer disease. Neurology 85, 40–47 (2015).
Shi, M. et al. Plasma exosomal α-synuclein is likely CNS-derived and increased in Parkinson's disease. Acta Neuropathol. 128, 639–650 (2014).
Brock, G., Castellanos-Rizaldos, E., Hu, L., Coticchia, C. & Skog, J. Liquid biopsy for cancer screening, patient stratification and monitoring. Translat. Cancer Res. 4, 280–290 (2015).
Tsui, N. B. et al. Maternal plasma RNA sequencing for genome-wide transcriptomic profiling and identification of pregnancy-associated transcripts. Clin. Chem. 60, 954–962 (2014).
Ainsztein, A. M. et al. The NIH Extracellular RNA Communication Consortium. J. Extracell. Vesicles 4, 27493 (2015).
Quinn, J. F. et al. Extracellular RNAs: development as biomarkers of human disease. J. Extracell. Vesicles 4, 27495 (2015).
Laurent, L. C. et al. Meeting report: discussions and preliminary findings on extracellular RNA measurement methods from laboratories in the NIH Extracellular RNA Communication Consortium. J. Extracell. Vesicles 4, 26533 (2015).
Mestdagh, P. et al. Evaluation of quantitative miRNA expression platforms in the microRNA quality control (miRQC) study. Nat. Methods 11, 809–815 (2014). This paper provides a critical assessment of current platform performance in measuring miRNA expression.
Kelly, H. et al. Cross Platform standardisation of an experimental pipeline for use in the identification of dysregulated human circulating miRNAs. PLoS ONE 10, e0137389 (2015).
Kozomara, A. & Griffiths-Jones, S. miRBase: annotating high confidence microRNAs using deep sequencing data. Nucleic Acids Res. 42, D68–73 (2014).
Chan, P. P. & Lowe, T. M. GtRNAdb: a database of transfer RNA genes detected in genomic sequence. Nucleic Acids Res. 37, D93–D97 (2009).
Salzman, J., Gawad, C., Wang, P. L., Lacayo, N. & Brown, P. O. Circular RNAs are the predominant transcript isoform from hundreds of human genes in diverse cell types. PLoS ONE 7, e30733 (2012).
Memczak, S., Papavasileiou, P., Peters, O. & Rajewsky, N. Identification and characterization of circular RNAs as a new class of putative biomarkers in human blood. PLoS ONE 10, e0141214 (2015).
Goodarzi, H. et al. Endogenous tRNA-derived fragments suppress breast cancer progression via YBX1 displacement. Cell 161, 790–802 (2015). Researchers identified a novel regulatory role for fragments of tRNA; further research may find that other RNA fragments can have novel functions.
Sutter, D. E. et al. Performance of five FDA-approved rapid antigen tests in the detection of 2009 H1N1 influenza A virus. J. Med. Virol. 84, 1699–1702 (2012).
Santiago, G. A. et al. Analytical and clinical performance of the CDC real time RT-PCR assay for detection and typing of dengue virus. PLoS Negl. Trop. Dis. 7, e2311 (2013).
Styer, L. M., Miller, T. T. & Parker, M. M. Validation and clinical use of a sensitive HIV-2 viral load assay that uses a whole virus internal control. J. Clin. Virol. 58 (Suppl. 1), e127–e133 (2013).
Pinsky, B. A. et al. Analytical performance characteristics of the Cepheid GeneXpert Ebola Assay for the detection of Ebola virus. PLoS ONE 10, e0142216 (2015).
Fischer, N. et al. Evaluation of unbiased next-generation sequencing of RNA (RNA-seq) as a diagnostic method in influenza virus-positive respiratory samples. J. Clin. Microbiol. 53, 2238–2250 (2015). This study describes the use of an unbiased RNA-seq application for positive detection and characterization of influenza in respiratory samples, paving the way for this tool as a diagnostic methodology.
Gire, S. K. et al. Genomic surveillance elucidates Ebola virus origin and transmission during the 2014 outbreak. Science 345, 1369–1372 (2014). This paper describes the use of RNA-seq for viral detection and measurement for epidemic emergence and tracking.
Gregori, J. et al. Ultra-deep pyrosequencing (UDPS) data treatment to study amplicon HCV minor variants. PLoS ONE 8, e83361 (2013).
Ekici, H. et al. Cost-efficient HIV-1 drug resistance surveillance using multiplexed high-throughput amplicon sequencing: implications for use in low- and middle-income countries. J. Antimicrob. Chemother. 69, 3349–3355 (2014).
Wang, K. et al. The complex exogenous RNA spectra in human plasma: an interface with human gut biota? PLoS ONE 7, e51009 (2012).
Beatty, M. et al. Small RNAs from plants, bacteria and fungi within the order Hypocreales are ubiquitous in human plasma. BMC Genomics 15, 933 (2014). The authors aptly analyse the small RNA component of human plasma that relates to the microbiome. Although small in sample size, this study helps to pave the way to more robust circulating small RNA studies combining both host and microbial RNA-omes.
Ghosal, A. et al. The extracellular RNA complement of Escherichia coli. Microbiologyopen http://dx.doi.org/10.1002/mbo3.235 (2015).
Chen, S. J. et al. Characterization of Epstein–Barr virus miRNAome in nasopharyngeal carcinoma by deep sequencing. PLoS ONE 5, e12745 (2010).
Meshesha, M. K. et al. The microRNA transcriptome of human cytomegalovirus (HCMV). Open Virol. J. 6, 38–48 (2012).
Cucher, M. et al. High-throughput characterization of Echinococcus spp. metacestode miRNomes. Int. J. Parasitol. 45, 253–267 (2015).
Fagnocchi, L. et al. Global transcriptome analysis reveals small RNAs affecting Neisseria meningitidis bacteremia. PLoS ONE 10, e0126325 (2015).
Latorre, I. et al. A novel whole-blood miRNA signature for a rapid diagnosis of pulmonary tuberculosis. Eur. Respir. J. 45, 1173–1176 (2015).
Ren, N. et al. MicroRNA signatures from multidrugresistant Mycobacterium tuberculosis. Mol. Med. Rep. 12, 6561–6567 (2015).
Lie, A. K. & Kristensen, G. Human papillomavirus E6/E7 mRNA testing as a predictive marker for cervical carcinoma. Expert Rev. Mol. Diagn. 8, 405–415 (2008).
Kanwal, F., Kramer, J. R., Ilyas, J., Duan, Z. & El-Serag, H. B. HCV genotype 3 is associated with an increased risk of cirrhosis and hepatocellular cancer in a national sample of U. S. Veterans with HCV. Hepatol. 60, 98–105 (2014).
Mitchell, A. M. et al. Transmitted/founder hepatitis C viruses induce cell-type- and genotype-specific differences in innate signaling within the liver. MBio 6, e02510 (2015).
Wu, L. S. et al. Systematic expression profiling analysis identifies specific microRNA-gene interactions that may differentiate between active and latent tuberculosis infection. Biomed. Res. Int. 2014, 895179 (2014).
Barry, S. E. et al. Identification of miR-93 as a suitable miR for normalizing miRNA in plasma of tuberculosis patients. J. Cell. Mol. Med. 19, 1606–1613 (2015).
Haddow, J. E. & Palomaki, G. E. in Human genome epidemiology: a scientific foundation for using genetic information to improve health and prevent disease. (eds Khoury, M. J., Little, J. & Burke, W.) 217–233 (Oxford Univ. Press, 2004).
' t Hoen, P. A. et al. Reproducibility of high-throughput mRNA and small RNA sequencing across laboratories. Nat. Biotechnol. 31, 1015–1022 (2013).
Zhao, W. et al. Comparison of RNA-Seq by poly (A) capture, ribosomal RNA depletion, and DNA microarray for expression profiling. BMC Genomics 15, 419 (2014).
Mercer, T. R. et al. Targeted sequencing for gene discovery and quantification using RNA CaptureSeq. Nat. Protoc. 9, 989–1009 (2014).
Cabanski, C. R. et al. cDNA hybrid capture improves transcriptome analysis on low-input and archived samples. J. Mol. Diagn. 16, 440–451 (2014).
Cieslik, M. et al. The use of exome capture RNA-seq for highly degraded RNA with application to clinical cancer sequencing. Genome Res. 25, 1372–1381 (2015). This paper describes the application of exome- capture transcriptome sequencing to FFPE samples in the clinical setting.
Zhang, Z. H. et al. A Comparative study of techniques for differential expression analysis on RNA-seq data. PLoS ONE 9, e103207–e103211 (2014).
Munro, S. A. et al. Assessing technical performance in differential gene expression experiments with external spike-in RNA control ratio mixtures. Nat. Commun. 5, 5125 (2014).
Jiang, L. et al. Synthetic spike-in standards for RNA-seq experiments. Genome Res. 21, 1543–1551 (2011). This article discusses the development of widely adopted synthetic RNA spike-ins providing sensitivity, accuracy and biases in RNA-seq experiments; this work also provides an early characterization of the limits for discovery of rare transcripts in RNA-seq.
Tembe, W. D. et al. Open-access synthetic spike-in mRNA-seq data for cancer gene fusions. BMC Genomics 15, 1–9 (2014). Publicly available RNA-seq data using synthetic spike-ins of known cancer gene fusions.
Su, Z. et al. A comprehensive assessment of RNA-seq accuracy, reproducibility and information content by the Sequencing Quality Control Consortium. Nat. Biotechnol. 32, 903–914 (2014). This study reports results from the SEQC project, evaluating RNA-seq performance assessments such as reproducibility and accuracy across sequencing platforms and collaborative sites.
Li, S. et al. Multi-platform assessment of transcriptome profiling using RNA-seq in the ABRF next-generation sequencing study. Nat. Biotechnol. 32, 915–925 (2014). This study reports results from the ABRF-NGS study assessing the influence of experimental protocols & RNA conditions across sequencing platforms and collaborative sites.
Wang, C. et al. The concordance between RNA-seq and microarray data depends on chemical treatment and transcript abundance. Nat. Biotechnol. 32, 926–932 (2014). This study reports results from the SEQC project, evaluating the influence of chemical treatment on RNA-seq performance.
Li, S. et al. Detecting and correcting systematic variation in large-scale RNA sequencing data. Nat. Biotechnol. 32, 888–895 (2014). References 100–103 describe large-scale efforts from the SEQC and ABRF consortia towards establishing standards for RNA-seq. This study evaluated systematic biases across platforms and laboratory sites, and assessed data analysis approaches for data normalization and bias correction.
Risso, D., Ngai, J., Speed, T. P. & Dudoit, S. Normalization of RNA-seq data using factor analysis of control genes or samples. Nat. Biotechnol. 32, 896–902 (2014).
Garber, M., Grabherr, M. G., Guttman, M. & Trapnell, C. Computational methods for transcriptome annotation and quantification using RNA-seq. Nat. Methods 8, 469–477 (2011).
Genomes Project, C. et al. A map of human genome variation from population-scale sequencing. Nature 467, 1061–1073 (2010).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Handsaker, R. E., Korn, J. M., Nemesh, J. & McCarroll, S. A. Discovery and genotyping of genome structural polymorphism by sequencing on a population scale. Nat. Genet. 43, 269–276 (2011).
Rehm, H. L. et al. ACMG clinical laboratory standards for next-generation sequencing. Genet. Med. 15, 733–747 (2013).
Wang, Y., Lu, J., Yu, J., Gibbs, R. A. & Yu, F. An integrative variant analysis pipeline for accurate genotype/haplotype inference in population NGS data. Genome Res. 23, 833–842 (2013).
DePristo, M. A. et al. A framework for variation discovery and genotyping using next-generation DNA sequencing data. Nat. Genet. 43, 491–498 (2011).
Zook, J. M. et al. Integrating human sequence data sets provides a resource of benchmark SNP and indel genotype calls. Nat. Biotechnol. 32, 246–251 (2014).
Di Tommaso, P. et al. The impact of Docker containers on the performance of genomic pipelines. PeerJ 3, e1273 (2015).
Kalari, K. R. et al. MAP-RSeq: Mayo Analysis Pipeline for RNA sequencing. BMC Bioinformatics 15, 224 (2014).
Nasser, S. et al. An integrated framework for reporting clinically relevant biomarkers from paired tumor/normal genomic and transcriptomic sequencing data in support of clinical trials in personalized medicine. Pac. Symp. Biocomput. 56–67 (2015).
Alexander, E. K. et al. Preoperative diagnosis of benign thyroid nodules with indeterminate cytology. N. Engl. J. Med. 367, 705–715 (2012).
Alexander, E. K. et al. Multicenter clinical experience with the Afirma gene expression classifier. J. Clin. Endocrinol. Metab. 99, 119–125 (2014).
US Department of Health & Human Services. Center for Devices and Radiological Health. FDA notification and medical device reporting for laboratory developed tests (LDTs) — draft guidance. [online] (2014).
US Department of Health & Human Services. Center for Devices and Radiological Health. Framework for regulatory oversight of laboratory developed tests (LDTs) — draft guidance. [online] (2014).
US Department of Health & Human Services. Optimizing FDA' s regulatory oversight of next generation sequencing diagnostic tests — preliminary discussion paper. [online] (2014).
Evans, B. J., Burke, W. & Jarvik, G. P. The FDA and genomic tests—getting regulation right. N. Engl. J. Med. 372, 2258–2264 (2015). This commentary discusses FDA oversight of genome-scale tests, including NGS, framing the discussion around the concepts of analytical and clinical validity.
Tazawa, Y. Perspective for the development of companion diagnostics and regulatory landscape to encourage personalized medicine in Japan. Breast Cancer 23, 19–23 (2015).
Pignatti, F. et al. Cancer drug development and the evolving regulatory framework for companion diagnostics in the European union. Clin. Cancer Res. 20, 1458–1468 (2014).
Matthijs, G. et al. Guidelines for diagnostic next-generation sequencing. Eur. J. Hum. Genet. 24, 2–5 (2016).
European Commission. Proposal for a regulation of the European Parliament and of the council on in vitro diagnostic medical devices. [online] (2012).
Rashid, N. U. et al. Differential and limited expression of mutant alleles in multiple myeloma. Blood 124, 3110–3117 (2014).
Andersson, A. K. et al. The landscape of somatic mutations in infant MLL-rearranged acute lymphoblastic leukemias. Nat. Genet. 47, 330–337 (2015).
Chandrasekharappa, S. C. et al. Massively parallel sequencing, aCGH, and RNA-Seq technologies provide a comprehensive molecular diagnosis of Fanconi anemia. Blood 121, e138–e148 (2013).
De Vlaminck, I. et al. Circulating cell-free DNA enables noninvasive diagnosis of heart transplant rejection. Sci. Transl Med. 6, 241ra77 (2014).
De Vlaminck, I. et al. Noninvasive monitoring of infection and rejection after lung transplantation. Proc. Natl Acad. Sci. USA 112, 13336–133341 (2015).
Heid, C. A., Stevens, J., Livak, K. J. & Williams, P. M. Real time quantitative PCR. Genome Res. 6, 986–994 (1996).
Wang, W. et al. Design of multiplexed detection assays for identification of avian influenza a virus subtypes pathogenic to humans by SmartCycler real-time reverse transcription-PCR. J. Clin. Microbiol. 47, 86–92 (2009).
Lockhart, D. J. et al. Expression monitoring by hybridization to high-density oligonucleotide arrays. Nat. Biotechnol. 14, 1675–1680 (1996).
Mook, S., Van't Veer, L. J., Rutgers, E. J., Piccart-Gebhart, M. J. & Cardoso, F. Individualization of therapy using Mammaprint: from development to the MINDACT Trial. Cancer Genom. Proteom. 4, 147–155 (2007). This paper describes the development and clinical validation trial for the Mammaprint gene expression based breast cancer recurrence test.
Paik, S. et al. A multigene assay to predict recurrence of tamoxifen-treated, node-negative breast cancer. N. Engl. J. Med. 351, 2817–2826 (2004). This paper describes the development and clinical validation trial for the OncotypeDX gene expression based breast cancer recurrence test.
Cooperberg, M. R. et al. Validation of a cell-cycle progression gene panel to improve risk stratification in a contemporary prostatectomy cohort. J. Clin. Oncol. 31, 1428–1434 (2013).
Learn, C. A. et al. Resistance to tyrosine kinase inhibition by mutant epidermal growth factor receptor variant III contributes to the neoplastic phenotype of glioblastoma multiforme. Clin. Cancer Res. 10, 3216–3224 (2004).
Robinson, D. et al. Integrative clinical genomics of advanced prostate cancer. Cell 161, 1215–1228 (2015).
Mody, R. J. et al. Integrative clinical sequencing in the management of refractory or relapsed cancer in youth. JAMA 314, 913–925 (2015).
Sekulic, A. et al. Personalized treatment of Sezary syndrome by targeting a novel CTLA4:CD28 fusion. Mol. Genet. Genom. Med. 3, 130–136 (2015).
Craig, D. W. et al. Genome and transcriptome sequencing in prospective metastatic triple-negative breast cancer uncovers therapeutic vulnerabilities. Mol. Cancer Ther. 12, 104–116 (2013). One of the first papers investigating integration of whole-transcriptome sequencing and genome sequencing for targeted therapy selection in advanced metastatic triple-negative breast cancer.
Borad, M. J. et al. Integrated genomic characterization reveals novel, therapeutically relevant drug targets in FGFR and EGFR pathways in sporadic intrahepatic cholangiocarcinoma. PLoS Genet. 10, e1004135 (2014). This paper describes the discovery of therapeutically actionable events including novel oncogenic fusions in FGFR2 identified by RNA-sequencing.
Greco, F. A., Lennington, W. J., Spigel, D. R. & Hainsworth, J. D. Molecular profiling diagnosis in unknown primary cancer: accuracy and ability to complement standard pathology. J. Natl Cancer Inst. 105, 782–790 (2013).
Paik, S. et al. Gene expression and benefit of chemotherapy in women with node-negative, estrogen receptor-positive breast cancer. J. Clin. Oncol. 24, 3726–3734 (2006).
Albain, K. S. et al. Prognostic and predictive value of the 21-gene recurrence score assay in postmenopausal women with node-positive, oestrogen-receptor-positive breast cancer on chemotherapy: a retrospective analysis of a randomised trial. Lancet Oncol. 11, 55–65 (2010).
Knezevic, D. et al. Analytical validation of the Oncotype DX prostate cancer assay - a clinical RT-PCR assay optimized for prostate needle biopsies. BMC Genomics 14, 690 (2013).
Clark-Langone, K. M., Sangli, C., Krishnakumar, J. & Watson, D. Translating tumor biology into personalized treatment planning: analytical performance characteristics of the OncotypeDX Colon Cancer Assay. BMC Cancer 10, 691 (2010).
Zhang, Y. et al. Breast cancer index identifies early-stage estrogen receptor-positive breast cancer patients at risk for early- and late-distant recurrence. Clin. Cancer Res. 19, 4196–4205 (2013).
Wallden, B. et al. Development and verification of the PAM50-based Prosigna breast cancer gene signature assay. BMC Med. Genom. 8, 54 (2015).
Salazar, R. et al. Gene expression signature to improve prognosis prediction of stage II and III colorectal cancer. J. Clin. Oncol. 29, 17–24 (2011).
Erho, N. et al. Discovery and validation of a prostate cancer genomic classifier that predicts early metastasis following radical prostatectomy. PLoS ONE 8, e66855 (2013).
Meiri, E. et al. A second-generation microRNA-based assay for diagnosing tumor tissue origin. Oncologist 17, 801–812 (2012).
Taubert, H. et al. Expression of the stem cell self-renewal gene Hiwi and risk of tumour-related death in patients with soft-tissue sarcoma. Oncogene 26, 1098–1100 (2007).
Baraniskin, A. et al. Circulating U2 small nuclear RNA fragments as a novel diagnostic biomarker for pancreatic and colorectal adenocarcinoma. Int. J. Cancer 132, E48–E57 (2013).
Su, J. et al. Analysis of small nucleolar RNAs in sputum for lung cancer diagnosis. Oncotarget http://dx.doi.org/10.18632/oncotarget.4219 (2015).
Du, Z. et al. Integrative genomic analyses reveal clinically relevant long noncoding RNAs in human cancer. Nat. Struct. Mol. Biol. 20, 908–913 (2013).
Li, P. et al. Using circular RNA as a novel type of biomarker in the screening of gastric cancer. Clin. Chim. Acta 444, 132–136 (2015).
Guo, Y. et al. Transfer RNA detection by small RNA deep sequencing and disease association with myelodysplastic syndromes. BMC Genomics 16, 727 (2015).
Westermann, A. J. et al. Dual RNA-seq unveils noncoding RNA functions in host-pathogen interactions. Nature 529, 496–501 (2016).
Patton, J. G. et al. Biogenesis, delivery, and function of extracellular RNA. J. Extracell. Vesicles 4, 27494 (2015).
Brinkmann, K. et al. Exosomal RNA-based liquid biopsy detection of EML4-ALK in plasma from NSCLC patients. 16th World Conference on Lung Cancer Abstract 2591 (IASLC, 2015).
Acknowledgements
The authors acknowledge funding from the Ben and Catherine Ivy Foundation and the National Center for Advancing Translational Sciences (exRNA Signatures Predict Outcomes After Brain Injury; UH3-TR000891). Research was also supported by a Stand Up To Cancer – Melanoma Research Alliance Melanoma Dream Team Translational Cancer Research Grant. Stand Up To Cancer is a programme of the Entertainment Industry Foundation administered by the American Association for Cancer Research. The authors apologize to those whose work could not be cited or discussed owing to space constraints.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Glossary
- Next-generation sequencing
-
(NGS). High-throughput, massively parallel sequencing technology that is used in various applications, including whole-genome sequencing, exome sequencing and RNA sequencing.
- Open platform
-
A technology platform that does not depend on genome annotation, or on predesigned species-specific or transcript-specific probes, for transcript measurement. RNA-seq technology functions as an open platform allowing for unbiased detection of both known and novel transcripts.
- Single nucleotide variants
-
(SNVs). Single nucleotide (A, T, G or C) alterations in a DNA sequence.
- Reference standards
-
Highly characterized and standardized control materials that are used to ensure accuracy and comparability of assays.
- Spearman correlation
-
Statistical measure of the strength of association between two rank-ordered variables.
- Phasing
-
Evaluation of closely situated mutations to determine whether they reside on the same or different alleles.
- Break-apart probes
-
A DNA probe system used to detect rearrangements involving specific loci. Probes for a region 5′ of the designated breakpoint are labelled with one colour and probes for a region 3′ of the breakpoint are labelled with another colour. An overlapping signal (such as yellow for red and green probes) indicates a normal pattern, whereas distinct signals (that is, red and green) indicate the presence of a rearrangement.
- Structural variants
-
Genomic variants, other than single-nucleotide variants, involving large regions of DNA, including insertions, deletions, inversions and duplications.
- Single nucleotide polymorphisms
-
(SNPs). Single nucleotide alterations that represent single base-pair variation at a specific DNA position among individuals, the majority of which are inherited.
- Nonsense-mediated decay
-
A translation-coupled RNA decay mechanism whereby aberrant mRNAs with premature stop codons are recognized and degraded.
- Expression quantitative trait loci
-
(eQTLs). Genomic loci that regulate the quantitative phenotypic trait of gene expression. Genetic markers at these loci are associated with measurable changes in gene expression.
- RNA split reads
-
RNA sequencing reads that are split — for example, to accommodate exon junctions.
- Extracellular RNAs
-
(exRNAs). RNAs found outside of the cell, they can be protected within vesicles or in association with RNA-binding proteins, and they can include exogenous sequences.
- Clinical Laboratory Improvement Amendments
-
(CLIA). All laboratory testing on humans in the United States is regulated by The Centers for Medicare and Medicaid Services through CLIA. The purpose is to ensure quality and uniformity of laboratory tests.
- Metagenomic RNA-seq
-
A method of sequencing the entirety of the available RNA in a complex (for example, clinical or environmental) sample, which may or may not include steps to subtract the host RNA to improve or enrich for microbial RNA.
- RNA-based amplicon sequencing
-
A method of direct sequencing of cDNA amplicons of RNA targets from a clinical sample. This can be multiplexed and can involve RNA viral genomes, microbial or host mRNA transcripts, or exogenous RNA targets.
- Microbiome
-
The totality of the genomic content of microbial community members in a complex (for example, clinical or environmental) sample. In the human microbiome, each body site has its own unique microbiome; the entirety of the microbiome on and in an individual person is considered that person's pan-microbiome.
- Companion diagnostic
-
In vitro diagnostic tests that provide information critical for the safe and effective use of a corresponding therapeutic agent. These tests are used to select patients for treatment with specific agents, including identifying patient populations with predicted efficacy as well as those that should not receive the agent due to a low likelihood of effectiveness or possible serious adverse events from the therapy.
- Variant call format
-
Standard text file format for storing genomic sequence variant data, with each line of the file describing a variant present at a specific genomic region or position.
- Binary alignment/map format
-
(BAM format). Standard file format for storing sequencing reads with alignments. BAM files are binary representations of the sequence alignment/map (SAM) format.
- In vitro diagnostic tests
-
Laboratory tests used to detect health conditions, infections or diseases. These diagnostic tests are performed using a sample collected from the patient without direct physical interaction between the test and the patient. In the United States, in vitro diagnostics are regulated by the US Food and Drug Administration (FDA).
- Triple-negative breast cancer
-
A breast cancer subtype characterized as oestrogen receptor-negative, progesterone receptor-negative and ERBB2 (also known as HER2)-negative.
Rights and permissions
About this article
Cite this article
Byron, S., Van Keuren-Jensen, K., Engelthaler, D. et al. Translating RNA sequencing into clinical diagnostics: opportunities and challenges. Nat Rev Genet 17, 257–271 (2016). https://doi.org/10.1038/nrg.2016.10
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrg.2016.10
This article is cited by
-
CryoGrid-PIXUL-RNA: high throughput RNA isolation platform for tissue transcript analysis
BMC Genomics (2023)
-
Integrative phosphoproteomics defines two biologically distinct groups of KMT2A rearranged acute myeloid leukaemia with different drug response phenotypes
Signal Transduction and Targeted Therapy (2023)
-
Prevalence and clinico-genomic characteristics of patients with TRK fusion cancer in China
npj Precision Oncology (2023)
-
Technical assessment of different extraction methods and transcriptome profiling of RNA isolated from small volumes of blood
Scientific Reports (2023)
-
RNA-Seq–based transcriptome analysis of corneal endothelial cells derived from patients with Fuchs endothelial corneal dystrophy
Scientific Reports (2023)