Key Points
-
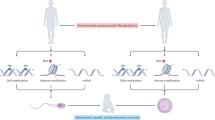
Recent evidence increasingly supports the idea that certain ancestral life experiences acquired in the environment can be inherited by offspring; paternally acquired characteristics can be encoded in the sperm in the form of epigenetic information in addition to DNA sequences.
-
Sperm RNAs, in particular sperm microRNAs (miRNAs) and tRNA-derived small RNAs (tsRNAs), can mediate intergenerational transmission of paternally acquired phenotypes such as diet-induced metabolic disorders and mental stress phenotypes.
-
The mechanisms by which sperm RNAs respond to environmental changes and encode the acquired traits remain unclear but may involve environmental–somatic–germline interactions that may be mediated by extracellular vesicles (EVs) and mobile RNAs, and involve a breach of the somatic–germline barrier.
-
Sperm RNAs may initiate a transcriptional cascade of effects throughout embryonic development to induce a paternally acquired phenotype in offspring; how the initial effects caused by sperm RNAs are converted to a stable form of information to allow transgenerational inheritance remains a major puzzle but possibly involves interplay among transposable elements, DNA methylation and chromatin structure.
-
Emerging evidence suggests that RNA modifications in sperm RNAs have an essential role in modulating epigenetic memory. Novel methods are required to map the locations of multiple RNA modifications in each RNA species, especially for tsRNAs and miRNAs that can induce offspring phenotypes.
-
It remains unknown how many types of acquired traits can be transmitted to offspring through the germ line and under what circumstances this is likely to occur.
Abstract
Once deemed heretical, emerging evidence now supports the notion that the inheritance of acquired characteristics can occur through ancestral exposures or experiences and that certain paternally acquired traits can be 'memorized' in the sperm as epigenetic information. The search for epigenetic factors in mammalian sperm that transmit acquired phenotypes has recently focused on RNAs and, more recently, RNA modifications. Here, we review insights that have been gained from studying sperm RNAs and RNA modifications, and their roles in influencing offspring phenotypes. We discuss the possible mechanisms by which sperm become acquisitive following environmental–somatic–germline interactions, and how they transmit paternally acquired phenotypes by shaping early embryonic development.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Landman, O. E. The inheritance of acquired characteristics. Annu. Rev. Genet. 25, 1–20 (1991).
Rechavi, O. et al. Starvation-induced transgenerational inheritance of small RNAs in C. elegans. Cell 158, 277–287 (2014).
Ost, A. et al. Paternal diet defines offspring chromatin state and intergenerational obesity. Cell 159, 1352–1364 (2014).
Ng, S. F. et al. Chronic high-fat diet in fathers programs β-cell dysfunction in female rat offspring. Nature 467, 963–966 (2010).
Carone, B. R. et al. Paternally induced transgenerational environmental reprogramming of metabolic gene expression in mammals. Cell 143, 1084–1096 (2010). References 4 and 5 show that acquired metabolic disorders due to paternal diets could be inherited by the offspring.
Dias, B. G. & Ressler, K. J. Parental olfactory experience influences behavior and neural structure in subsequent generations. Nat. Neurosci. 17, 89–96 (2014).
Anway, M. D., Cupp, A. S., Uzumcu, M. & Skinner, M. K. Epigenetic transgenerational actions of endocrine disruptors and male fertility. Science 308, 1466–1469 (2005); correction, 328, 690 (2010).
Rodgers, A. B., Morgan, C. P., Bronson, S. L., Revello, S. & Bale, T. L. Paternal stress exposure alters sperm microRNA content and reprograms offspring HPA stress axis regulation. J. Neurosci. 33, 9003–9012 (2013).
Chen, Q. et al. Sperm tsRNAs contribute to intergenerational inheritance of an acquired metabolic disorder. Science 351, 397–400 (2016). This study demonstrates that zygotic injection of sperm total RNAs or tsRNAs from males fed a HFD can induce metabolic disorders in offspring, and that RNA modifications in sperm tsRNAs are sensitive to a HFD.
Huypens, P. et al. Epigenetic germline inheritance of diet-induced obesity and insulin resistance. Nat. Genet. 48, 497–499 (2016).
Sharma, U. et al. Biogenesis and function of tRNA fragments during sperm maturation and fertilization in mammals. Science 351, 391–396 (2016). This paper provides evidence suggesting that sperm can gain tsRNAs during epididymal transition, which can influence embryonic gene expression.
Lane, M., Robker, R. L. & Robertson, S. A. Parenting from before conception. Science 345, 756–760 (2014).
Feng, S., Jacobsen, S. E. & Reik, W. Epigenetic reprogramming in plant and animal development. Science 330, 622–627 (2010).
Wei, Y. et al. Paternally induced transgenerational inheritance of susceptibility to diabetes in mammals. Proc. Natl Acad. Sci. USA 111, 1873–1878 (2014).
Daxinger, L. & Whitelaw, E. Understanding transgenerational epigenetic inheritance via the gametes in mammals. Nat. Rev. Genet. 13, 153–162 (2012).
Shea, J. M. et al. Genetic and epigenetic variation, but not diet, shape the sperm methylome. Dev. Cell 35, 750–758 (2015). This study suggests that DNA methylation is not the primary epigenetic carrier for diet-induced sperm changes that are responsible for reprogramming of offspring metabolism.
Radford, E. J. et al. In utero effects. In utero undernourishment perturbs the adult sperm methylome and intergenerational metabolism. Science 345, 1255903 (2014). This article shows that in utero nutritional exposure during critical windows of germ-cell development alters the sperm methylome of offspring, but the altered methylation does not persist in the next generation.
Iqbal, K. et al. Deleterious effects of endocrine disruptors are corrected in the mammalian germline by epigenome reprogramming. Genome Biol. 16, 59 (2015).
de Waal, E. et al. Primary epimutations introduced during intracytoplasmic sperm injection (ICSI) are corrected by germline-specific epigenetic reprogramming. Proc. Natl Acad. Sci. USA 109, 4163–4168 (2012).
Hammoud, S. S. et al. Distinctive chromatin in human sperm packages genes for embryo development. Nature 460, 473–478 (2009).
Brykczynska, U. et al. Repressive and active histone methylation mark distinct promoters in human and mouse spermatozoa. Nat. Struct. Mol. Biol. 17, 679–687 (2010).
Ragunathan, K., Jih, G. & Moazed, D. Epigenetics. Epigenetic inheritance uncoupled from sequence-specific recruitment. Science 348, 1258699 (2015).
Jodar, M. et al. The presence, role and clinical use of spermatozoal RNAs. Hum. Reprod. Update 19, 604–624 (2013).
Gapp, K. et al. Implication of sperm RNAs in transgenerational inheritance of the effects of early trauma in mice. Nat. Neurosci. 17, 667–669 (2014). This study provides the first functional evidence for sperm RNAs in transferring the acquired phenotype, showing that the injection of sperm RNAs from a traumatized father induces corresponding phenotypes in offspring.
Rodgers, A. B., Morgan, C. P., Leu, N. A. & Bale, T. L. Transgenerational epigenetic programming via sperm microRNA recapitulates effects of paternal stress. Proc. Natl Acad. Sci. USA 112, 13699–13704 (2015).
Grandjean, V. et al. RNA-mediated paternal heredity of diet-induced obesity and metabolic disorders. Sci. Rep. 5, 18193 (2015).
Kiani, J. et al. RNA-mediated epigenetic heredity requires the cytosine methyltransferase Dnmt2. PLoS Genet. 9, e1003498 (2013). This paper shows that DNMT2 is required for RNA-mediated non-Mendelian epigenetic inheritance in mice.
Nelson, V. R., Heaney, J. D., Tesar, P. J., Davidson, N. O. & Nadeau, J. H. Transgenerational epigenetic effects of the Apobec1 cytidine deaminase deficiency on testicular germ cell tumor susceptibility and embryonic viability. Proc. Natl Acad. Sci. USA 109, E2766–E2773 (2012). This report shows that APOBEC1 is involved in transgenerational epigenetic inheritance of a phenotype.
Bohacek, J. & Mansuy, I. M. Molecular insights into transgenerational non-genetic inheritance of acquired behaviours. Nat. Rev. Genet. 16, 641–652 (2015).
Roemer, I., Reik, W., Dean, W. & Klose, J. Epigenetic inheritance in the mouse. Curr. Biol. 7, 277–280 (1997).
Fullston, T. et al. Paternal obesity initiates metabolic disturbances in two generations of mice with incomplete penetrance to the F2 generation and alters the transcriptional profile of testis and sperm microRNA content. FASEB J. 27, 4226–4243 (2013).
Wu, L. et al. Paternal psychological stress reprograms hepatic gluconeogenesis in offspring. Cell Metab. 23, 735–743 (2016).
Paris, L. et al. Transgenerational inheritance of enhanced susceptibility to radiation-induced medulloblastoma in newborn Ptch1+/− mice after paternal irradiation. Oncotarget 6, 36098–36112 (2015).
Morgan, H. D., Sutherland, H. G., Martin, D. I. & Whitelaw, E. Epigenetic inheritance at the agouti locus in the mouse. Nat. Genet. 23, 314–318 (1999).
Rakyan, V. K. et al. Transgenerational inheritance of epigenetic states at the murine AxinFu allele occurs after maternal and paternal transmission. Proc. Natl Acad. Sci. USA 100, 2538–2543 (2003).
Siklenka, K. et al. Disruption of histone methylation in developing sperm impairs offspring health transgenerationally. Science 350, aab2006 (2015). This study demonstrates that defects induced by genetically disrupting histone methylation cause transgenerational inheritance of phenotypes in mammals.
Ziller, M. J., Hansen, K. D., Meissner, A. & Aryee, M. J. Coverage recommendations for methylation analysis by whole-genome bisulfite sequencing. Nat. Methods 12, 230–232 (2015).
Herman, H. et al. Trans allele methylation and paramutation-like effects in mice. Nat. Genet. 34, 199–202 (2003).
Rassoulzadegan, M., Magliano, M. & Cuzin, F. Transvection effects involving DNA methylation during meiosis in the mouse. EMBO J. 21, 440–450 (2002).
Hatada, I. et al. Aberrant methylation of an imprinted gene U2af1-rs1(SP2) caused by its own transgene. J. Biol. Chem. 272, 9120–9122 (1997).
Grandjean, V. et al. The miR-124-Sox9 paramutation: RNA-mediated epigenetic control of embryonic and adult growth. Development 136, 3647–3655 (2009).
Wagner, K. D. et al. RNA induction and inheritance of epigenetic cardiac hypertrophy in the mouse. Dev. Cell 14, 962–969 (2008).
Rassoulzadegan, M. et al. RNA-mediated non-Mendelian inheritance of an epigenetic change in the mouse. Nature 441, 469–474 (2006). This article provides the first evidence that sperm RNAs can mediate epigenetic inheritance in mammals.
Chong, S. et al. Modifiers of epigenetic reprogramming show paternal effects in the mouse. Nat. Genet. 39, 614–622 (2007).
de Castro Barbosa, T. et al. High-fat diet reprograms the epigenome of rat spermatozoa and transgenerationally affects metabolism of the offspring. Mol. Metab. 5, 184–197 (2016).
Peng, H. et al. A novel class of tRNA-derived small RNAs extremely enriched in mature mouse sperm. Cell Res. 22, 1609–1612 (2012). This is the first report that tsRNAs are highly enriched in mature sperm.
Schuster, A., Skinner, M. K. & Yan, W. Ancestral vinclozolin exposure alters the epigenetic transgenerational inheritance of sperm small noncoding RNAs. Environ. Epigenet. 2, dvw001 (2016).
Donkin, I. et al. Obesity and bariatric surgery drive epigenetic variation of spermatozoa in humans. Cell Metab. 23, 369–378 (2015).
Surani, M. A. Breaking the germ line-soma barrier. Nat. Rev. Mol. Cell Biol. 17, 136 (2016).
Liu, Y. A new perspective on Darwin's pangenesis. Biol. Rev. Camb. Philos. Soc. 83, 141–149 (2008).
Koch, S., Acebron, S. P., Herbst, J., Hatiboglu, G. & Niehrs, C. Post-transcriptional Wnt signaling governs epididymal sperm maturation. Cell 163, 1225–1236 (2015).
Sarkies, P. & Miska, E. A. Small RNAs break out: the molecular cell biology of mobile small RNAs. Nat. Rev. Mol. Cell Biol. 15, 525–535 (2014).
Lo Cicero, A., Stahl, P. D. & Raposo, G. Extracellular vesicles shuffling intercellular messages: for good or for bad. Curr. Opin. Cell Biol. 35, 69–77 (2015).
Machtinger, R., Laurent, L. C. & Baccarelli, A. A. Extracellular vesicles: roles in gamete maturation, fertilization and embryo implantation. Hum. Reprod. Update 22, 182–193 (2016).
Mahmoudi, K., Ezrin, A. & Hadjipanayis, C. Small extracellular vesicles as tumor biomarkers for glioblastoma. Mol. Aspects Med. 45, 97–102 (2015).
Vader, P., Mol, E. A., Pasterkamp, G. & Schiffelers, R. M. Extracellular vesicles for drug delivery. Adv. Drug Deliv. Rev. http://dx.doi.org/10.1016/j.addr.2016.02.006 (2016).
Vojtech, L. et al. Exosomes in human semen carry a distinctive repertoire of small non-coding RNAs with potential regulatory functions. Nucleic Acids Res. 42, 7290–7304 (2014).
Zhang, Y. et al. Identification and characterization of an ancient class of small RNAs enriched in serum associating with active infection. J. Mol. Cell. Biol. 6, 172–174 (2014).
Dhahbi, J. M. et al. 5′ tRNA halves are present as abundant complexes in serum, concentrated in blood cells, and modulated by aging and calorie restriction. BMC Genomics 14, 298 (2013).
Li, H., Wu, C., Aramayo, R., Sachs, M. S. & Harlow, M. L. Synaptic vesicles contain small ribonucleic acids (sRNAs) including transfer RNA fragments (trfRNA) and microRNAs (miRNA). Sci. Rep. 5, 14918 (2015).
Anderson, P. & Ivanov, P. tRNA fragments in human health and disease. FEBS Lett. 588, 4297–4304 (2014).
Cossetti, C. et al. Soma-to-germline transmission of RNA in mice xenografted with human tumour cells: possible transport by exosomes. PLoS ONE 9, e101629 (2014).
Zeybel, M. et al. Multigenerational epigenetic adaptation of the hepatic wound-healing response. Nat. Med. 18, 1369–1377 (2012).
Devanapally, S., Ravikumar, S. & Jose, A. M. Double-stranded RNA made in C. elegans neurons can enter the germline and cause transgenerational gene silencing. Proc. Natl Acad. Sci. USA 112, 2133–2138 (2015).
Curran, S. P., Wu, X., Riedel, C. G. & Ruvkun, G. A soma-to-germline transformation in long-lived Caenorhabditis elegans mutants. Nature 459, 1079–1084 (2009).
Shirayama, M. et al. piRNAs initiate an epigenetic memory of nonself RNA in the C. elegans germline. Cell 150, 65–77 (2012).
Buckley, B. A. et al. A nuclear Argonaute promotes multigenerational epigenetic inheritance and germline immortality. Nature 489, 447–451 (2012).
Ashe, A. et al. piRNAs can trigger a multigenerational epigenetic memory in the germline of C. elegans. Cell 150, 88–99 (2012).
Javurek, A. B. et al. Discovery of a novel seminal fluid microbiome and influence of estrogen receptor alpha genetic status. Sci. Rep. 6, 23027 (2016); corrigendum 6, 25216 (2016).
Muraca, M., Putignani, L., Fierabracci, A., Teti, A. & Perilongo, G. Gut microbiota-derived outer membrane vesicles: under-recognized major players in health and disease? Discov. Med. 19, 343–348 (2015).
Hoshino, A. et al. Tumour exosome integrins determine organotropic metastasis. Nature 527, 329–335 (2015).
Ridder, K. et al. Extracellular vesicle-mediated transfer of genetic information between the hematopoietic system and the brain in response to inflammation. PLoS Biol. 12, e1001874 (2014).
Gregor, M. F. & Hotamisligil, G. S. Inflammatory mechanisms in obesity. Annu. Rev. Immunol. 29, 415–445 (2011).
Cohen, S. et al. Chronic stress, glucocorticoid receptor resistance, inflammation, and disease risk. Proc. Natl Acad. Sci. USA 109, 5995–5999 (2012).
Fan, Y. et al. Diet-induced obesity in male C57BL/6 mice decreases fertility as a consequence of disrupted blood-testis barrier. PLoS ONE 10, e0120775 (2015).
Nargund, V. H. Effects of psychological stress on male fertility. Nat. Rev. Urol. 12, 373–382 (2015).
Crean, A. J., Kopps, A. M. & Bonduriansky, R. Revisiting telegony: offspring inherit an acquired characteristic of their mother's previous mate. Ecol. Lett. 17, 1545–1552 (2014).
Crean, A. J., Adler, M. I. & Bonduriansky, R. Seminal fluid and mate choice: new predictions. Trends Ecol. Evol. 31, 253–255 (2016).
Arroyo, J. D. et al. Argonaute2 complexes carry a population of circulating microRNAs independent of vesicles in human plasma. Proc. Natl Acad. Sci. USA 108, 5003–5008 (2011).
Feinberg, E. H. & Hunter, C. P. Transport of dsRNA into cells by the transmembrane protein SID-1. Science 301, 1545–1547 (2003).
McEwan, D. L., Weisman, A. S. & Hunter, C. P. Uptake of extracellular double-stranded RNA by SID-2. Mol. Cell 47, 746–754 (2012).
Ivanov, P. et al. G-quadruplex structures contribute to the neuroprotective effects of angiogenin-induced tRNA fragments. Proc. Natl Acad. Sci. USA 111, 18201–18206 (2014).
Chin, A. R. et al. Cross-kingdom inhibition of breast cancer growth by plant miR159. Cell Res. 26, 217–228 (2016).
Yang, J., Farmer, L. M., Agyekum, A. A. & Hirschi, K. D. Detection of dietary plant-based small RNAs in animals. Cell Res. 25, 517–520 (2015).
Mlotshwa, S. et al. A novel chemopreventive strategy based on therapeutic microRNAs produced in plants. Cell Res. 25, 521–524 (2015).
Zhang, L. et al. Exogenous plant MIR168a specifically targets mammalian LDLRAP1: evidence of cross-kingdom regulation by microRNA. Cell Res. 22, 107–126 (2012).
Zhou, Z. et al. Honeysuckle-encoded atypical microRNA2911 directly targets influenza A viruses. Cell Res. 25, 39–49 (2015).
Ji, L. & Chen, X. Regulation of small RNA stability: methylation and beyond. Cell Res. 22, 624–636 (2012).
Skinner, M. K., Guerrero-Bosagna, C. & Haque, M. M. Environmentally induced epigenetic transgenerational inheritance of sperm epimutations promote genetic mutations. Epigenetics 10, 762–771 (2015).
Houri-Ze'evi, L. et al. A tunable mechanism determines the duration of the transgenerational small RNA inheritance in C. elegans. Cell 165, 88–99 (2016).
Piotrowska, K. & Zernicka-Goetz, M. Role for sperm in spatial patterning of the early mouse embryo. Nature 409, 517–521 (2001).
Shi, J. et al. Dynamic transcriptional symmetry-breaking in pre-implantation mammalian embryo development revealed by single-cell RNA-seq. Development 142, 3468–3477 (2015).
Goolam, M. et al. Heterogeneity in Oct4 and Sox2 targets biases cell fate in 4-cell mouse embryos. Cell 165, 61–74 (2016).
White, M. D. et al. Long-lived binding of Sox2 to DNA predicts cell fate in the four-cell mouse embryo. Cell 165, 75–87 (2016). References 92–94 provide the conceptual framework and experimental evidence that the slightest blastomere–blastomere biases during early embryo development can change the trajectory of developmental potential.
Torres-Padilla, M. E., Parfitt, D. E., Kouzarides, T. & Zernicka-Goetz, M. Histone arginine methylation regulates pluripotency in the early mouse embryo. Nature 445, 214–218 (2007).
Plachta, N., Bollenbach, T., Pease, S., Fraser, S. E. & Pantazis, P. Oct4 kinetics predict cell lineage patterning in the early mammalian embryo. Nat. Cell Biol. 13, 117–123 (2011).
Davidson, E. H. Emerging properties of animal gene regulatory networks. Nature 468, 911–920 (2010).
Holoch, D. & Moazed, D. RNA-mediated epigenetic regulation of gene expression. Nat. Rev. Genet. 16, 71–84 (2015).
Watanabe, T. et al. Role for piRNAs and noncoding RNA in de novo DNA methylation of the imprinted mouse Rasgrf1 locus. Science 332, 848–852 (2011).
Zuo, Z. et al. Genome-wide analysis reveals origin of transfer RNA genes from tRNA halves. Mol. Biol. Evol. 30, 2087–2098 (2013).
Wu, T. P. et al. DNA methylation on N6-adenine in mammalian embryonic stem cells. Nature 532, 329–333 (2016).
Greer, E. L. et al. DNA methylation on N6-adenine in C. elegans. Cell 161, 868–878 (2015).
Morris, K. V. & Mattick, J. S. The rise of regulatory RNA. Nat. Rev. Genet. 15, 423–437 (2014).
Engreitz, J. M. et al. The Xist lncRNA exploits three-dimensional genome architecture to spread across the X chromosome. Science 341, 1237973 (2013).
Hacisuleyman, E. et al. Topological organization of multichromosomal regions by the long intergenic noncoding RNA Firre. Nat. Struct. Mol. Biol. 21, 198–206 (2014).
Simon, M. D. et al. High-resolution Xist binding maps reveal two-step spreading during X-chromosome inactivation. Nature 504, 465–469 (2013).
Xiang, J. F. et al. Human colorectal cancer-specific CCAT1-L lncRNA regulates long-range chromatin interactions at the MYC locus. Cell Res. 24, 513–531 (2014).
Dekker, J. & Mirny, L. The 3D genome as moderator of chromosomal communication. Cell 164, 1110–1121 (2016).
Le Thomas, A. et al. Transgenerationally inherited piRNAs trigger piRNA biogenesis by changing the chromatin of piRNA clusters and inducing precursor processing. Genes Dev. 28, 1667–1680 (2014).
Fu, Y., Dominissini, D., Rechavi, G. & He, C. Gene expression regulation mediated through reversible m6A RNA methylation. Nat. Rev. Genet. 15, 293–306 (2014).
Chen, K., Zhao, B. S. & He, C. Nucleic acid modifications in regulation of gene expression. Cell Chem. Biol. 23, 74–85 (2016).
Yan, M. et al. A high-throughput quantitative approach reveals more small RNA modifications in mouse liver and their correlation with diabetes. Anal. Chem. 85, 12173–12181 (2013).
Machnicka, M. A. et al. MODOMICS: a database of RNA modification pathways — 2013 update. Nucleic Acids Res. 41, D262–D267 (2013).
Cantara, W. A. et al. The RNA Modification Database, RNAMDB: 2011 update. Nucleic Acids Res. 39, D195–D201 (2011).
Schaefer, M. et al. RNA methylation by Dnmt2 protects transfer RNAs against stress-induced cleavage. Genes Dev. 24, 1590–1595 (2010).
Goll, M. G. et al. Methylation of tRNAAsp by the DNA methyltransferase homolog Dnmt2. Science 311, 395–398 (2006).
Tuorto, F. et al. RNA cytosine methylation by Dnmt2 and NSun2 promotes tRNA stability and protein synthesis. Nat. Struct. Mol. Biol. 19, 900–905 (2012).
Popow, J., Jurkin, J., Schleiffer, A. & Martinez, J. Analysis of orthologous groups reveals archease and DDX1 as tRNA splicing factors. Nature 511, 104–107 (2014).
Han, C. et al. The RNA-binding protein DDX1 promotes primary microRNA maturation and inhibits ovarian tumor progression. Cell Rep. 8, 1447–1460 (2014).
Hildebrandt, M. R., Germain, D. R., Monckton, E. A., Brun, M. & Godbout, R. Ddx1 knockout results in transgenerational wild-type lethality in mice. Sci. Rep. 5, 9829 (2015).
Zheng, G. et al. Efficient and quantitative high-throughput tRNA sequencing. Nat. Methods 12, 835–837 (2015).
Cozen, A. E. et al. ARM-seq: AlkB-facilitated RNA methylation sequencing reveals a complex landscape of modified tRNA fragments. Nat. Methods 12, 879–884 (2015). References 121 and 122 report novel methods to discover previously undetected tRNAs and tsRNAs by RNA-seq.
Kirchner, S. & Ignatova, Z. Emerging roles of tRNA in adaptive translation, signalling dynamics and disease. Nat. Rev. Genet. 16, 98–112 (2015).
Schwartzman, O. & Tanay, A. Single-cell epigenomics: techniques and emerging applications. Nat. Rev. Genet. 16, 716–726 (2015).
Angermueller, C. et al. Parallel single-cell sequencing links transcriptional and epigenetic heterogeneity. Nat. Methods 13, 229–232 (2016).
Hou, Y. et al. Single-cell triple omics sequencing reveals genetic, epigenetic, and transcriptomic heterogeneity in hepatocellular carcinomas. Cell Res. 26, 304–319 (2016).
Ostermeier, G. C., Dix, D. J., Miller, D., Khatri, P. & Krawetz, S. A. Spermatozoal RNA profiles of normal fertile men. Lancet 360, 772–777 (2002).
Ostermeier, G. C., Miller, D., Huntriss, J. D., Diamond, M. P. & Krawetz, S. A. Reproductive biology: delivering spermatozoan RNA to the oocyte. Nature 429, 154 (2004).
Yuan, S. et al. mir-34b/c and mir-449a/b/c are required for spermatogenesis, but not for the first cleavage division in mice. Biol. Open 4, 212–223 (2015).
Liu, W. M. et al. Sperm-borne microRNA-34c is required for the first cleavage division in mouse. Proc. Natl Acad. Sci. USA 109, 490–494 (2012).
Yuan, S. et al. Sperm-borne miRNAs and endo-siRNAs are important for fertilization and preimplantation embryonic development. Development 143, 635–647 (2016).
Zhong, C. et al. Parthenogenetic haploid embryonic stem cells efficiently support mouse generation by oocyte injection. Cell Res. 26, 131–134 (2016).
Li, Z. et al. Birth of fertile bimaternal offspring following intracytoplasmic injection of parthenogenetic haploid embryonic stem cells. Cell Res. 26, 135–138 (2016).
Miller, D. Confrontation, consolidation, and recognition: the oocyte's perspective on the incoming sperm. Cold Spring Harb. Perspect. Med. 5, a023408 (2015).
Chandler, V. L. Paramutation: from maize to mice. Cell 128, 641–645 (2007).
Yuan, S., Oliver, D., Schuster, A., Zheng, H. & Yan, W. Breeding scheme and maternal small RNAs affect the efficiency of transgenerational inheritance of a paramutation in mice. Sci. Rep. 5, 9266 (2015).
Beale, L. S. Pangenesis. Nature 4, 25–26 (1871).
Eberhard, W. G. Postcopulatory sexual selection: Darwin's omission and its consequences. Proc. Natl Acad. Sci. USA 106 (Suppl. 1), 10025–10032 (2009).
Pizzari, T. & Foster, K. R. Sperm sociality: cooperation, altruism, and spite. PLoS Biol. 6, e130 (2008).
Edward, D. A., Stockley, P. & Hosken, D. J. Sexual conflict and sperm competition. Cold Spring Harb. Perspect. Biol. 7, a017707 (2015).
Acknowledgements
The Chen laboratory is currently supported by start-up funds from the University of Nevada, Reno School of Medicine, USA. This work is in part funded by the Ministry of Science and Technology of the People's Republic of China (2015CB943000 to Q.C. and 2016YFA0500903, 2015DFG32640 to E.D.), the National Natural Science Foundation of China (81490742, 31671568 to E.D.), the US National Institutes of Health (HD060858, HD071736 and HD085506 to W.Y.) and the John Templeton Foundation, USA (PID: 50183 to W.Y.). The authors thank all Chen laboratory members for critical discussions on the contents of the manuscript and for help in preparing the graphics and illustrations.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Related links
Glossary
- Parthenogenetic
-
A type of asexual reproduction that occurs when a female gamete develops a new individual without being fertilized by a male gamete
- RNA-editing events
-
Molecular processes by which specific nucleotide sequences in an RNA molecule are changed, such as C-to-U and A-to-I editing.
- Kinky-tail phenotype
-
A mouse phenotype characterized by a kinked tail containing a sharp bend at an angle to the main tail axis.
- Transvection
-
An epigenetic phenomenon that involves the interaction between two homologous chromosomes, resulting in either gene activation or repression at an allele.
- Paramutation
-
Interaction between two alleles at a single locus during meiosis, resulting in epigenetic transfer of information from one allele to another that is heritable for generations.
- Epididymal maturation
-
Spermatozoa from testis undergo a maturation process during transit from the proximal to the distal end of the epididymis, acquiring motility and fertility.
- Polyandrous
-
The mating behaviour of animals in which the females mate with more than one male in a single breeding cycle.
- G-Quadruplex conformation
-
Guanine-rich oligonucleotides that assembled into intra- or inter-molecular guanine tetrad structures, which can be formed by both DNA and RNA.
- Lamarckism
-
The idea that an organism can pass on characteristics that are acquired during its lifetime to its offspring.
Rights and permissions
About this article
Cite this article
Chen, Q., Yan, W. & Duan, E. Epigenetic inheritance of acquired traits through sperm RNAs and sperm RNA modifications. Nat Rev Genet 17, 733–743 (2016). https://doi.org/10.1038/nrg.2016.106
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrg.2016.106
This article is cited by
-
Multigenerational paternal obesity enhances the susceptibility to male subfertility in offspring via Wt1 N6-methyladenosine modification
Nature Communications (2024)
-
Sperm-borne microRNA-34c regulates maternal mRNA degradation and preimplantation embryonic development in mice
Reproductive Biology and Endocrinology (2023)
-
HA-tag CD63 is a novel conditional transgenic approach to track extracellular vesicle interactions with sperm and their transfer at conception
Scientific Reports (2023)
-
Inheritance of paternal lifestyles and exposures through sperm DNA methylation
Nature Reviews Urology (2023)
-
Paternal methotrexate exposure affects sperm small RNA content and causes craniofacial defects in the offspring
Nature Communications (2023)