Key Points
-
Seasonal influenza virus vaccines are an effective countermeasure against influenza if the vaccine strains and the circulating viruses are well matched; vaccine efficacy drops sharply if mismatched viruses are circulating.
-
Viruses from the animal reservoir, including H3N2v, H5N1, H5N6, H6N1, H7N3, H7N9 and H10N8, have recently caused morbidity and mortality in humans. Although these viruses are unable to transmit efficiently among humans, the development of pre-pandemic vaccine candidates that could enhance pandemic preparedness is warranted.
-
Pandemic influenza virus vaccines must be produced in a timely manner to effectively reduce the impact of a novel pandemic virus on the global human population. Technological advances such as gene synthesis, reverse genetics and recombinant production systems will facilitate the production of vaccines more rapidly in response to future influenza pandemics.
-
Novel human monoclonal antibody technology has helped provide a better understanding of the humoral (crossreactive) immune responses against the influenza virus surface glycoproteins haemagglutinin and neuraminidase.
-
Glycosylation of haemagglutinin and neuraminidase has a role in the immunogenicity of influenza virus vaccines and vaccine candidates.
-
Broadly protective antibodies against the haemagglutinin stalk domain and neuraminidase guide the design of novel, broadly protective vaccines. Novel influenza virus vaccine candidates that induce broad or universal influenza virus coverage are currently in preclinical and clinical development.
-
Broadly protective or universal influenza virus vaccines could abolish the need for annual reformulation and re-administration of seasonal influenza virus vaccines and could improve our pandemic preparedness.
Abstract
Influenza virus infections are a major public health concern and cause significant morbidity and mortality worldwide. Current influenza virus vaccines are an effective countermeasure against infection but need to be reformulated almost every year owing to antigenic drift. Furthermore, these vaccines do not protect against novel pandemic strains, and the timely production of pandemic vaccines remains problematic because of the limitations of current technology. Several improvements have been made recently to enhance immune protection induced by seasonal and pandemic vaccines, and to speed up production in case of a pandemic. Importantly, vaccine constructs that induce broad or even universal influenza virus protection are currently in preclinical and clinical development.
Similar content being viewed by others
Main
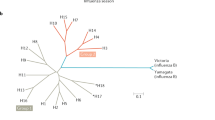
Seasonal influenza virus epidemics are estimated to cause 2–5 million cases of severe illness and up to 250,000–500,000 deaths per year worldwide1. Global annual infection rates are estimated to be 5–10% in adults and 20–30% in children1. Although current influenza virus vaccines are an effective countermeasure against disease, the vaccines induce narrow and strain-specific immunity (see Box 1 for mechanisms of anti-influenza immunity) and have to be updated in a complex, costly and time-consuming process almost every year because of antigenic drift. Four distinct types of influenza viruses are currently co-circulating in the human population: two are influenza A viruses (the 2009 H1N1 pandemic strain and H3N2) and the other two are divergent lineages of the influenza B virus2. Vaccine formulations have to contain at least the two influenza A virus strains and one influenza B virus strain, which further complicates the manufacturing process of such vaccines2.
In addition to seasonal epidemics, influenza viruses cause pandemics at irregular intervals. The influenza virus pandemic of 1918 claimed approximately 40 million lives and was caused by an H1N1 virus3,4. Since then, pandemics have been caused by H2N2 in 1957, by H3N2 in 1968 and again by H1N1 in 2009 (Refs 3,5). Pandemics are caused by influenza viruses that have crossed the species barrier from the animal reservoir (for example, avian species and swine) and acquire the ability to efficiently grow in humans and transmit among the population (Box 2). Importantly, these viruses are often reassortants of haemagglutinin and neuraminidase (HA and NA) genomic segments from animal viruses and several internal genomic segments from human, or at least mammalian, virus origin3. Seasonal influenza virus vaccines are usually ineffective against novel pandemic viruses; therefore, a strain-specific vaccine has to be produced (Fig. 1). Unfortunately, the production of a strain-specific vaccine is time-consuming and the vaccine might be distributed and administered too late, as was the case in 2009 in the United States6.
This figure shows the vaccine production process in response to a new pandemic. Orange stars indicate steps that could be accelerated by using novel technology. For example, seed strain preparation and verification can be facilitated by using gene synthesis, reverse genetics and deep sequencing. Once seed strains are prepared, viruses can be rescued in a backbone that has been optimized for growth on the selected substrate. This strategy can reduce the time required for growth optimization. Bulk manufacturing can be expedited by using novel production technologies that are easy to scale up (for example, cell culture or recombinant protein technologies). Adapted from Ref. 227, World Health Organization.
Here, we describe improvements that have been made in the production process of both seasonal and pandemic influenza virus vaccines to overcome these problems. Furthermore, we discuss novel vaccine constructs, vaccination regimens and adjuvants that induce broader and sustained protection. Finally, we review novel findings regarding the immune response towards haemagglutinin and neuraminidase, and provide an overview of several universal influenza virus vaccine approaches that could lead to vaccines with lifelong protection from any type of influenza virus7.
Improving seasonal influenza virus vaccines
Vaccines against influenza A and B viruses were invented in the 1940s. These early vaccines, termed whole-virus inactivated vaccines, were generated in embryonated chicken eggs (a technology that is still predominant today) and consisted of crudely purified whole virus inactivated with formalin and phenylmercuric nitrate8,9. The effect of antigenic drift made it necessary to reformulate vaccines after only 2 years of use, and the World Health Organization soon established an influenza surveillance network for the early detection of drifted strains10,11. The 1968 pandemic led to the development of trivalent inactivated vaccines (TIVs) against influenza viruses12. Furthermore, studies on reactogenicity to different vaccine formulations in children ultimately led to the development of split and subunit vaccines13. These vaccines are split using ether and/or detergent, and haemagglutinin and neuraminidase are, in the case of subunit vaccines, purified and enriched2. In addition to inactivated influenza vaccines (IIVs), live attenuated influenza vaccines (LAIVs) are also used. LAIVs are usually temperature-sensitive and cold-adapted and will efficiently replicate in the upper respiratory tract but not in the lower respiratory tract. LAIVs, which are administered by nasal spray, have been developed in parallel in Russia (licensed in 1980) and in the United States (licensed in 2003)14,15,16.
A recent study that evaluated 34 randomized clinical trials concluded that the vaccine efficacy of LAIVs in children (the age group for which this type of vaccine is indicated and thought to be most effective) is approximately 83% and the efficacy of TIVs in adults is approximately 75% (Ref. 17). Furthermore, a study on the use of IIVs in pregnant women in Bangladesh showed that vaccination reduced the incidence of influenza virus infection in mothers and newborns, and also significantly decreased the number of stillbirths and increased birth weight18,19. Collectively, these studies demonstrate that current seasonal influenza virus vaccines confer good protection against infection and are an important public health tool. However, a vaccine efficacy of 75% is far from optimal and drops sharply in the elderly who are more susceptible to influenza virus infection20,21. Furthermore, the duration of protection is short22,23. Mismatches between vaccine strains and circulating strains also occasionally occur and are usually associated with lower vaccine efficacy24.
Recently, improvements in vaccine formulations have been made with the goal of eliciting better protection against seasonal influenza virus strains. To induce a stronger, broader and more sustained immune response — specifically in the elderly — several novel formulations have been tested (Table 1). These formulations range from high-dose vaccines for the elderly, which have been licensed in the United States25,26, to the development of several adjuvanted vaccines. Two of the most advanced adjuvant formulations — MF59 and AS03 — have been tested with seasonal influenza virus vaccines and were able to enhance the efficacy of the vaccines27. MF59 adjuvanted seasonal vaccines for the elderly population have been licensed and marketed in more than 25 countries under the brand name Fluad (Novartis)27,28. AS03 adjuvanted influenza vaccines are also under consideration for use in the elderly population29. An additional improvement in seasonal influenza virus vaccines is the inclusion of a second influenza B virus strain. The novel quadrivalent influenza virus vaccine is now licensed in the United States as an IIV and a LAIV, but debate regarding the added value of these vaccines compared with TIVs is ongoing30,31,32.
Another strategy that can be used to induce a broader and more sustained immune response against seasonal influenza virus strains is based on heterologous prime–boost regimens. This type of regimen has been tested in mice, in ferrets and in nonhuman primates. A DNA vaccine expressing a haemagglutinin from a seasonal influenza virus is administered first (prime), and a typical TIV is subsequently administered (boost). Mice that received the prime–boost regimen showed broader immunity and had a more than 50-fold higher neutralizing titre than that induced by TIVs only33. Pre-existing immunity to influenza virus, which occurs in humans, did not have a negative effect on this vaccination regimen34. Several clinical trials that translated these findings into humans have recently been completed (ClinicalTrials.gov identifiers: NCT01609998, NCT01676402, NCT00995982 and NCT01498718). Another study showed that vaccination with ferritin particles displaying influenza virus haemagglutinin trimers induced stronger and broader immune responses than TIVs35.
Improvements on the vaccine production side include the US licensure of the first recombinant influenza virus vaccine (FluBlok; Protein Sciences Corporation) and the US licensure of the first cell-culture-derived seasonal influenza virus vaccine (Flucelvax; Novartis)36,37. These developments in vaccine production have also had a high impact on improving the speed at which pandemic influenza virus vaccines can be produced (Fig. 1).
Improving pandemic preparedness
During the past decades, several avian influenza viruses have caused zoonotic outbreaks in the human population. These outbreaks were sporadic and were usually associated with close contact to infected poultry or other avian species. Highly pathogenic H5N1 viruses in humans were first detected in Hong Kong in 1997 and reappeared in 2003 (Refs 38,39). These viruses express a haemagglutinin with a multibasic cleavage site and are therefore able to replicate to high titres in many tissues in infected birds40. The H5N1 virus is now distributed over Eurasia and Africa and has evolved into a number of antigenically distinct clades39. Humans have been occasionally infected and the high fatality rate of the infection, together with the wide geographical spread of the H5N1 virus, has raised concerns about its pandemic potential41 (see The WHO Influenza Monthly Risk Assessment Summaries; Influenza at the Human–Animal Interface (in Further information)). However, serological data suggest that a high number of infections with the virus — for example, in Southeast Asia — remain subclinical in humans42. Furthermore, the H5N1 virus expresses an N1 subtype of neuraminidase that is closely related to the neuraminidase of the currently circulating pandemic H1N1 virus43. As such, the human population would not be completely naive to a pandemic strain of H5N1.
Many other zoonotic viruses, including H5N6, H6N1, H7N9 and H10N8, have recently caused morbidity and mortality in humans in Asia44,45,46,47,48. In addition, H3N2 variant viruses that transmit from pigs to humans, seal H3N8 and H10N7 viruses, and highly pathogenic avian H5N8 and H7N3 viruses have raised concerns about their potential to spread in the human population in Europe and in North America49,50,51,52,53. It is difficult to predict the strain or subtype that will cause the next influenza virus pandemic. As the human population expands, the interface between the animal reservoir of influenza viruses and the human population grows. This expanded interface makes it more likely for a virus to cross the species barrier. Therefore, the development of vaccines for influenza virus strains with pandemic potential is warranted to improve our pandemic preparedness.
Currently, there are two major problems relating to pandemic influenza vaccines that need to be addressed. The first is the lag between pandemic virus identification and vaccine development and distribution. When a novel pandemic virus is identified, it takes months to develop, test, distribute and administer the new vaccine. After vaccination of an individual, it takes an additional 2–3 weeks until a protective immune response is mounted (Fig. 1). A stark example of this problem is the situation in 2009, when the majority of the pandemic H1N1 vaccine was distributed only after the second wave of the pandemic hit the US population6. The second issue is low immunogenicity. For example, current pandemic candidate vaccines against H5N1 and H7N9 induce relatively weak immune responses as measured by the traditional correlate of protection, the haemagglutination inhibition (HI) titre54,55,56,57. The cause of this low immunogenicity is currently debated, and vaccine formulations and regimens to overcome this problem are being developed.
Vaccine candidates for potentially pandemic viruses have been developed using a range of different production platforms. The IIV platform — in the split and whole virus format — has advanced the furthest, and vaccines made using this platform have been used for stockpiling58,59. In the case of vaccines against highly pathogenic H5N1 strains, seed strains have been generated using reverse genetics to remove the multibasic cleavage site of the haemagglutinin and to change the backbone to that of a high-growth A/Puerto Rico/8/1934 H1N1 strain59. These modifications render the vaccine strains safer and production possible because highly pathogenic influenza A viruses usually kill embryonated eggs, resulting in low yields of the vaccine59. A number of these H5N1 and H7 vaccines have been tested in humans and a high antigen dose or the use of an adjuvant (or a combination of both) was necessary to induce reliable haemagglutination inhibition titres above 1:40, which is the titre needed for approval by US and European regulatory authorities59,60. Even under these conditions, immune responses were low. A recent clinical trial of a H7N9 vaccine candidate resulted in a vaccine efficacy of approximately 60% despite the use of an adjuvant61. As described below, it has been hypothesized that vaccination with H5 (group 1 haemagglutinin) or H7 (group 2 haemagglutinin) vaccines primarily boosts antibodies against the conserved stalk domain of the haemagglutinin structure to which humans have low levels of pre-existing immunity62,63,64. This might explain why adjuvants and multiple vaccinations are necessary to yield sufficient vaccine efficacy.
As described above, two LAIV backbones (cold adapted A/Ann Arbor/6/1960 and A/Leningrad/134/17/1957) are currently available. Both backbones, as well as experimental LAIV constructs, have been used to generate and test pre-pandemic vaccines, including H2-, H5-, H6- and H7-expressing candidates65,66,67,68,69,70,71,72,73,74. Immune responses measured upon vaccination with these constructs in humans are moderate to weak depending on the ability of the vaccine virus to replicate in the upper respiratory tract65,66,67,68,69,70,71,72,73. The inability of vaccine viruses to replicate in the upper respiratory tract may be due to the absence of a specific glycan structure in this part of the anatomy of humans75.
A novel strategy that can improve the efficacy of pandemic vaccines is the use of a LAIV or DNA vaccine prime followed by an IIV boost. The LAIV or DNA vaccine immunologically primes subjects — often without a measurable seroconversion — and this immune response can subsequently be recalled by administering an IIV boost. Several clinical trials have demonstrated the value of this approach76,77,78. Strategies to prime particular groups of the human population (for example, health-care workers) with H5 or H7 LAIVs to induce a rapid and strong recall of the immune response in case of a pandemic are currently being discussed.
Novel platforms for rapid vaccine production. Rapid vaccine production in response to a novel pandemic influenza virus strain is vital for reducing global morbidity and mortality. Several novel technologies that improve the vaccine production process have been described in recent years (Fig. 1). The use of cellular substrates could make influenza virus vaccine production independent of the global embryonated egg supply and enable easy scaling up of the process. Several cell lines, including Madin–Darbey canine kidney cells, Vero cells (African green monkey) and Per.C6 cells (human), have been tested and established for influenza virus vaccine production55,79,80. In addition, novel gene synthesis technologies combined with influenza virus reverse genetics now enable the generation of custom-made seed strains within very short time frames80,81. These novel technologies can be used for both IIV and LAIV candidates, abolish the need for time-consuming classical reassortment and could significantly shorten their production time.
Recombinant protein expression has several advantages for the production of pandemic influenza virus vaccines. These include rapid vaccine production, the absence of infectious virus during production, the independence from egg supplies, the ease of scale up, the ability to use sequences derived directly from clinical specimens without egg- or cell-culture passage history and — for many recombinant expression systems — the low cost of production. A disadvantage of this approach is the reliance on one influenza virus antigen, usually haemagglutinin. These vaccines therefore lack the multifaceted immune response against other influenza virus proteins that might confer protection.
Numerous recombinant protein vaccines, mostly haemagglutinin-based, are currently in preclinical and clinical development. Additionally, the trivalent seasonal recombinant haemagglutinin vaccine FluBlok, which is produced in insect cells, has already been licensed by the US Food and Drug Administration and paved the way for pandemic vaccines to be produced in the same manner37. Popular expression systems for influenza virus vaccines and vaccine candidates include the following: baculovirus and insect cell expression systems82,83; Agrobacterium species-driven expression in plants such as the Nicotiana species84; and bacterial expression in Escherichia coli85,86. Furthermore, vaccine candidates have been expressed in Lactobacillus species87, algae88, yeast89,90 and cell-free expression systems91. Haemagglutinins expressed in insect and plant cell expression systems are relatively similar to those expressed in mammalian cells, with the exception of the N-linked glycosylation pattern, and are usually correctly folded. By contrast, haemagglutinin expressed in E. coli is not glycosylated, forms inclusion bodies and has to be refolded85,92.
Another platform developed for the production of influenza virus vaccines is the use of virus-like particles (VLPs). VLPs can be produced by co-expression of influenza virus structural proteins in mammalian cells, insect cells or plants83,93,94,95,96,97,98,99,100. Pandemic influenza VLP vaccines have been clinically tested and have shown good safety and efficacy profiles94,101,102.
In addition, several DNA and virus-vectored pandemic influenza virus vaccines are currently in preclinical and clinical development103,104. Many virus-vectored vaccines are based on modified vaccinia virus Ankara (MVA) because of its excellent safety profile. Several H5N1 and H7N9 MVA constructs have been tested in animal models and can induce strong cellular and humoral immune responses105,106,107,108,109,110. The efficacy of these vaccines in humans is currently being tested in clinical trials111.
Immunity to haemagglutinin and neuraminidase
Recent advances in human monoclonal antibody (mAb) technology, including phage library technology and expression cloning of antibodies from plasmablast and memory B-cell populations, have made it possible to gain new insight into the immune responses towards the influenza virus surface glycoproteins haemagglutinin and neuraminidase112,113,114,115,116,117 (Fig. 2). The rediscovery of haemagglutinin stalk-reactive antibodies that was facilitated by these techniques was a major milestone towards the development of a universal influenza virus vaccine. The first stalk-reactive antibody, mAb C179, was isolated in 1992 using traditional murine hybridoma technology118. However, stalk-reactive antibodies are rare in humans, and the first human antibodies with this specificity — CR6261, F10 and a small number of mAbs generated from an antibody library of Turkish H5N1 survivors — were only isolated in 2008–2009 (Refs 115,116,119). In addition to haemagglutinin stalk-reactive antibodies, several broadly reactive antibodies against the haemagglutinin globular head domain and neuraminidase have been discovered120,121,122,123,124,125.
Antibodies directed against the haemagglutinin (HA) globular head domain (red) and the stalk domain (green) or against neuraminidase (NA) (blue) may confer protection via a number of mechanisms. As viruses enter the host and come in contact with mucosal surfaces they are trapped by highly glycosylated innate defence proteins called mucins (step 1). NA helps the virus to pierce through this layer, and this activity may be inhibited by NA-specific antibodies. Antibodies that bind to the globular head domain of HA sterically block interactions between HA and sialic acid on cellular receptors, effectively inhibiting attachment of the virus to the cell (step 2). In rare cases, NA-specific antibodies can also have this function (also working through steric hindrance). Stalk-reactive antibodies bind to the HA on virus particles and prevent the fusion of viral and endosomal membranes by blocking the rearrangement of the HA fusion machinery (step 3). Head-, stalk- and NA-reactive antibodies may inhibit budding and viral egress (step 4). Stalk-reactive antibodies bound to HA sterically inhibit HA maturation (step 5). Stalk-reactive antibodies and possibly NA-reactive antibodies also act through antibody-dependent cell-mediated cytotoxicity (step 6) or complement-dependent cytotoxicity (step 7). cRNA, complementary RNA; FcR, Fc receptor; NK, natural killer; vRNA, viral RNA.
Haemagglutinin stalk-reactive antibodies. Stalk-reactive antibodies are particularly interesting because they bind epitopes on the membrane proximal, conserved portion of haemagglutinin and therefore show broad binding to divergent haemagglutinins. The binding pattern of most stalk-reactive antibodies follows the phylogeny of the influenza virus haemagglutinins and they bind to either group 1 (H1, H2, H5, H6, H8, H9, H11, H12, H13, H16, H17 and H18) or group 2 (H3, H4, H7, H10, H14 and H15) haemagglutinins116,118,126,127,128,129. However, some stalk mAbs have a narrower binding pattern and only recognize haemagglutinin of one subtype (for example, mAb 6F12 shows pan-H1 binding, and mAb 12D1 shows pan-H3 binding), whereas other exceptionally rare antibodies bind to all influenza A haemagglutinins or even crossreact between influenza A and B haemagglutinins130,131,132,133,134. Importantly, most stalk-reactive antibodies seem to bind preferentially to conformational epitopes but do not recognize denatured haemagglutinin116,126,135. Furthermore, they do not show haemagglutination inhibition activity136. In contrast to antibodies with haemagglutination inhibition activity (Fig. 2), which mostly neutralize by inhibiting the interaction between haemagglutinin and sialic acid residues on cellular receptors, stalk-reactive antibodies may protect through several mechanisms (Fig. 2).
Upon binding to haemagglutinin, stalk-reactive antibodies lock the haemagglutinin trimer in a pre-fusion conformation and prevent pH-triggered conformational change when the virus is taken up into the endosome (Fig. 2). Therefore, no fusion of the viral and endosomal membranes can occur and the virus is trapped in the endosome116,126,130,137. In addition, antibody binding sterically blocks access of proteases to the basic cleavage site between the HA1 and HA2 subunits of haemagglutinin, which is located in the stalk domain126,137 (Fig. 2). Uncleaved haemagglutinin (HA0) is unable to undergo the necessary conformational changes for fusion, and this mechanism might also contribute to the protection against infection. Finally, stalk-reactive antibodies also retain newly formed haemagglutinin on the cell surface and may inhibit virus budding129 (Fig. 2). In addition to mechanisms that directly neutralize the virus, other mechanisms such as antibody-dependent cell-mediated cytotoxicity (ADCC) and complement-dependent cytotoxicity might contribute to protection conferred by stalk-reactive antibodies in vivo138,139,140,141,142 (Fig. 2). Specifically, ADCC is an important factor and can potentiate the protective efficacy of stalk-reactive antibodies in vivo139.
Stalk-reactive antibodies are not induced at significant levels by currently used IIVs. The stalk domain seems to be immunosubdominant compared to the immunodominant globular head domain to which most antibodies are directed63,113,114. However, natural infection is able to induce a baseline level of these antibodies in mice and humans143,144,145. Interestingly, stalk-reactive antibody levels were boosted significantly by infection with the 2009 pandemic H1N1 virus, and these antibodies were also isolated from individuals who survived an H5N1 infection119,146,147. This led to the hypothesis that exposure to haemagglutinins that have a divergent head domain to which humans are naive (for example, H5N1 or pH1N1) and to stalk domains with conserved epitopes can boost stalk-reactive antibody titres. Additional support for this hypothesis comes from the analysis of clinical trials with pandemic vaccine candidates — including H5N1, H7N1 and swine-origin H1N1 strains — which induced preferentially stalk-reactive antibodies62,63,64,148,149,150.
Interestingly, studies with H5N1 vaccines showed that the first vaccine administration induces high levels of stalk-reactive antibodies, whereas the second vaccination with the same vaccine formulation predominantly induces a response against the globular head domain63,64. This result indicates that the globular head domain regains immunodominance over the stalk domain once the immune system is primed for these novel head domain epitopes. Importantly, polyclonal anti-stalk responses induced by H5N1 vaccines are highly crossreactive towards group 1 haemagglutinins but do not significantly crossreact with group 2 haemagglutinins when measured using quantitative methods63,64.
Broadly reactive antibodies against the haemagglutinin globular head domain and neuraminidase. In addition to broadly neutralizing stalk-specific antibodies, a small number of human antibodies that can neutralize a broad panel of influenza viruses through binding to the haemagglutinin head domain have been isolated121,122,123,124. Some of these antibodies bind to the receptor-binding site of haemagglutinin by mimicking sialic acid, the substrate to which haemagglutinin binds122,123,124. This molecular mimicry explains the binding breadth of these antibodies, which sometime spans several subtypes. However, the antibodies need to insert one of their binding loops deep into the receptor-binding site, and the addition of glycans on the rim around the receptor-binding site can sterically prevent binding without forcing the virus to change the conserved receptor-binding domain. Most of these antibodies are exceptionally rare but some light has been shed recently on the induction of broadly neutralizing antibodies against the H1 head domain of haemagglutinin151,152. Similar to stalk-reactive antibodies, these antibodies seem to be mostly induced when individuals are exposed to highly divergent H1 haemagglutinins over time. In this context, the specific exposure history of an individual, and especially the virus to which the individual was first exposed, seem to have a major role151,152.
Several antibodies against the second surface glycoprotein, neuraminidase, have also shown exceptional breadth153. A rabbit mAb against a conserved linear epitope on neuraminidase showed a broadly inhibitory effect on divergent neuraminidases from influenza A and B viruses and showed limited protection in passive transfer experiments154,155. However, it is unclear whether similar antibodies are induced by natural infection or influenza virus vaccination. In addition, murine antibodies with broad reactivity to the N1 subtype of neuraminidase have been reported recently120. Several of these have neuraminidase inhibition (NI) activity (Fig. 2) and are able to reduce virus cell-to-cell spread in vitro. In general, neuraminidase inhibition activity seems to correlate with in vivo protection for these antibodies. However, protection was also seen in cases in which mAbs did not have neuraminidase inhibition activity against the challenge virus, suggesting that alternative mechanisms such as ADCC and complement-dependent cytotoxicity might also have a role in vivo120.
Glycans: in the context of broadly reactive immune responses, size matters. Both the influenza virus haemagglutinin and neuraminidase are glycoproteins that have several putative N-glycosylation motifs, and glycosylation might have an important role in the folding and biology of these proteins156 (Fig. 3). Haemagglutinin has a variable number of glycosylation sites in the head domain, whereas glycosylation sites in the stalk domain are relatively conserved across haemagglutinin groups156. Haemagglutinin glycosylation has a strong influence on the pathogenicity and antigenicity of haemagglutinin, whereas the role of N-linked glycosylation on neuraminidase is less well understood157. Recent studies suggest that the number and size of glycans on haemagglutinin also influence the breadth of the immune response. Pre-pandemic seasonal H1, pandemic H1 or H5 haemagglutinins that were enzymatically treated to reduce the number of glycan structures to one N-acetylglucosamine showed broader immune responses and protection against challenge with heterologous strains than fully glycosylated haemagglutinins158,159. However, complete deglycosylation led to reduced protection, which is probably due to the loss of important conformational epitopes.
a–c | Haemagglutinin trimers without glycan structures attached (part a), with oligomannose glycan structures attached (part b) and with complex glycan structures attached (part c). Structures are based on the H3 subtype of haemagglutinin of a recent seasonal H3N2 isolate, A/Victoria/361/2011 (RCSB Protein Data Bank (PDB) ID: 4O5N228), which is heavily glycosylated (12 potential N-glycosylation sites). d–f | Structure of the neuraminidase tetramer of the 2009 pandemic H1N1 virus (PDB ID: 3NSS229) (part d). This neuraminidase possesses eight potential N-linked glycosylation sites, which are shown with oligomannosidic structures (part e) and with complex glycans (part f). These glycan structures, although flexible at physiological temperatures, might restrict the access of B cell receptors and antibodies to haemagglutinin and may shield important antigens. Glycans were attached to the haemagglutinin and neuraminidase structures using GlyProt and visualized using PyMOL230.
Reduction of the glycan size seems to lead to stronger immune responses against conserved epitopes that are probably less accessible when shielded by large glycans. This hypothesis is supported by studies showing that binding of broadly neutralizing stalk-reactive antibodies to fully glycosylated haemagglutinin is inhibited at low temperature (4 °C), which is when glycan structures are becoming rigid160. Interestingly, this effect was not seen with haemagglutinin produced in insect cells, which has smaller paucimannose-like non-complex glycan structures.
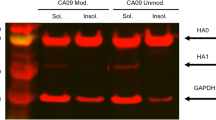
Glycan size on haemagglutinin is strongly influenced by the production method. Mammalian-cell-derived haemagglutinins (on average 12 monosaccharide units, sialylated if expressed without neuraminidase) have the largest glycans followed by egg-derived haemagglutinins (8–9 monosaccharide units, highly branched, no sialic acid). Insect-cell-derived haemagglutinins have glycans that are 5–6 monosaccharide units in length161 (Fig. 3). Therefore, vaccines made in production platforms that produce haemagglutinins with smaller glycans — such as insect cells83 — might be more suitable for inducing broad immune responses. However, some insect cell lines are known to add α-1,3-linked fucose to their glycans, which can be allergenic162.
Large glycan structures can shield epitopes from immune recognition on haemagglutinin157. The introduction of additional glycosylation sites on the immunodominant head domain might therefore be used to skew the immune response towards immunosubdominant epitopes in the stalk domain. A recent study demonstrated that hyperglycosylated H1 haemagglutinin produced in mammalian cells induces broadly protective immune responses against the stalk domain163. Similar results have been reported with prime–boost H5 vaccine strategies with vaccine constructs that had additional glycosylation sites grafted on the head domain164,165.
Development of universal influenza virus vaccines
Stalk-based vaccine constructs. Attempts to construct vaccines based on the stalk domain by removing the immunodominant head domain (producing a headless haemagglutinin) were made as early as 1983 (Ref. 166). The methodology used to remove the head domain, or more specifically the HA1 subunit of the haemagglutinin, involved an acid treatment followed by treatment with a reducing agent. However, this treatment induced significant conformational changes in the HA2 portion of the stalk domain and completely removed the HA1 portion of the stalk domain, therefore destroying important conformational epitopes. Following the discovery of the stalk-reactive mAb C179, a genetic approach to remove the globular head domain was developed167. A modified H2 haemagglutinin was expressed in mammalian cells and used to vaccinate mice, where it achieved limited protection against a heterosubtypic H1N1 challenge167. Another approach with an H1-based headless haemagglutinin displayed on VLPs showed success in the mouse model and was able to induce antibodies that crossreacted with H2 and H5 haemagglutinin168. Several other stalk-only and headless haemagglutinin constructs have been designed and expressed in E. coli and cell-free expression systems and have shown limited efficacy in a mouse model with low challenge doses169,170,171,172,173,174. The main obstacle to overcome for the development of successful headless haemagglutinin constructs is the correct folding of conformational neutralizing epitopes, and better approaches to design stable structures are needed.
A novel approach to induce high levels of stalk-reactive antibodies is based on chimeric haemagglutinins (cHAs)7,175,176 (Fig. 4a). Chimeric haemagglutinins consist of H1 (group 1), H3 (group 2) or influenza B haemagglutinin stalk domains in combination with 'exotic' globular head domains, mostly of avian origin. A disulfide bond between Cys52 and Cys277 (H3 numbering) forms the demarcation line between stalk and head domains. Amino acids between these two cysteine residues belong to the membrane distal globular head domain, whereas amino acids of the haemagglutinin ectodomain that are N-terminal of Cys52 and C-terminal of Cys277 belong to the stalk domain. Importantly, the stalk domain includes parts of the HA1 and the HA2 subunits. The presence of an exotic head domain on these chimeric haemagglutinins stabilizes important conformational epitopes in the stalk domain. Chimeric haemagglutinins are fully functional, and recombinant influenza viruses expressing them grow to high titres in embryonated eggs and in cell cultures175.
Chimeric haemagglutinins (cHAs) consist of 'exotic' globular head domains (for example, from H5 or H6 HA) in combination with H1, H3 or influenza B virus stalk domains. a | An example of a cH5/1 HA (an H5 head (yellow) on top of an H1 stalk (green)). b | Humans have pre-existing immunity to H1 (group 1), H3 (group 2) and influenza B viruses, which is mostly directed against the HA head domains but also includes low levels of anti-stalk immunity. Upon vaccination with a cHA — for example, a cH5/1 HA — antibody levels against the stalk domain are boosted, whereas only a primary response is induced against the novel globular head domain to which humans are naive. A boost with a second cHA that possesses the same stalk domain but a different head domain (for example, cH6/1 HA) could further increase stalk-specific antibody levels. Structures are based on RCSB Protein Data Bank ID 1RU7, and were visualized using Protein Workshop231.
Chimeric haemagglutinins with different head domains have been used in a sequential vaccination regimen to induce stalk-reactive antibodies. After the first exposure to a chimeric haemagglutinin — for example, cH6/1 HA (an H6 head on top of an H1 stalk) — the immune system induces a strong primary response against the exotic head domain but only a weak, almost undetectable, response against the stalk domain. Sequential vaccination with a second chimeric haemagglutinin that expresses a different head domain — for example, cH5/1 HA (an H5 head on top of an H1 stalk) — induces a primary response against the novel head domain but boosts antibodies against the stalk domain because both chimeric haemagglutinins have this domain in common. A third vaccination with yet another different chimeric haemagglutinin — for example, cH8/1 HA (an H8 head on top of an H1 stalk) — again boosts stalk-reactive antibodies whereas only a primary response against the H8 head domain is mounted (Fig. 4b). Using this strategy, it is possible to break the immunodominance of the head domain and to induce high titres of stalk-reactive antibodies.
As discussed above, the breadth of stalk-reactive antibodies is mostly restricted to one haemagglutinin group (group 1, group 2 or B haemagglutinins). Therefore, a successful chimeric haemagglutinin-based universal vaccine candidate needs a group 1 component, a group 2 component and an influenza B haemagglutinin component. Group 1 constructs based on the H1 stalk domain have so far been successfully tested in mice and ferrets and protect from heterologous (H1N1) and heterosubtypic challenge (for example, H5N1 and H6N1), but not from challenge with group 2 viruses (for example, H3N2)177,178. Group 2 constructs based on the H3 stalk domain can protect against various H3N2 viruses and against heterosubtypic challenge viruses such as H7N1 and H7N9 (Refs 179,180). Although most of these studies were performed using experimental DNA and recombinant protein vaccines, it should be mentioned that the chimeric haemagglutinin technology is platform independent and can potentially be used in the form of IIVs, LAIVs, virus vectors, recombinant protein vaccines, VLPs, DNA vaccines, and other forms. As described above, adults already have low levels of B cells with specificities against the stalk domain and would therefore probably only require boosting of these B cell populations with chimeric haemagglutinin constructs to increase the production of virus-specific antibodies (Fig. 4b). Evidence for this hypothesis comes from trials with H5N1 and H7N1 vaccine candidates62,63,64. As described above, these vaccines, which possess exotic head domains but have conserved group 1 or group 2 stalk domains, induced high levels of stalk-reactive antibodies in humans. However, one of these trials showed that the immune response against the stalk domain in the context of inactivated vaccines was as short lived as the immune response against the head domain, with titres returning to baseline 6 months post-vaccination64.
Clearly, a universal influenza virus vaccine that is protective for only a short duration is of limited use. However, it should be noted that stalk-directed immune responses induced by natural infection (and potentially by whole-virus inactivated vaccines) have long half-lives143,148. Moreover, adjuvants can drastically improve the immune response induced by chimeric haemagglutinin-based vaccines179,181. An adjuvanted chimeric haemagglutinin vaccine, possibly in the context of a heterologous prime–boost regimen (for example, an LAIV followed by an IIV or a DNA vaccine, followed by an IIV) could therefore be used to induce a long-lasting anti-stalk immune response. Clinical trials to test this hypothesis have been initiated. Importantly, novel potency assays and correlates of protection have to be established for these vaccine candidates because current assays and correlates are focused on globular-head-directed immunity.
Broadly protective vaccines based on the globular head domain of haemagglutinin, neuraminidase or M2e. In addition to universal vaccine approaches that are based on the conserved stalk domain, approaches to induce a broader response towards the globular head domain are in development182,183,184. These approaches are restricted to a subtype or even to specific clades within a subtype but could still result in vaccines that last for several years, which is a clear advantage over current vaccines that have to be reformulated almost every year. This concept is based on 'centralized' sequences182, ancestral sequences184 or computationally optimized broadly reactive antigens (COBRAs), which are synthetic haemagglutinins representing an optimized merged sequence of representative strains183,185. COBRA-based vaccines have been shown to successfully induce protection against highly pathogenic H5N1 viruses in mice, ferrets and nonhuman primates186,187,188. Candidates for seasonal influenza viruses are currently in development. Similar to chimeric haemagglutinin constructs, these COBRA-based haemagglutinins are fully functional and vaccine platform independent.
As described above, crossprotective mAbs against the second surface glycoprotein of the influenza virus, neuraminidase, demonstrate that neuraminidase-based immunity has the potential to confer at least intra-subtypic crossprotection. This is also supported by the fact that neuraminidase antigenic drift rates are generally lower than antigenic drift rates of the globular head domain of haemagglutinin189,190,191. However, the immune response to homologous neuraminidase after influenza virus vaccination and infection is not well characterized and understood153. IIVs are not standardized for their neuraminidase content, and the functionality and correct folding of the neuraminidase in these vaccines is not assessed on a regular basis. Recent efforts to gain a better understanding of the neuraminidase content in IIVs and the immune response that they induce showed marked differences in neuraminidase content and anti-neuraminidase immune responses for commercially available vaccines. Immune responses in mice varied from no induction to neuraminidase inhibition titres of 1:1,280 (Ref. 192). However, it has been demonstrated that neuraminidase-based immunity drastically reduces viral replication and clinical signs of infection in humans193. In general, it is assumed that neuraminidase, similar to the stalk domain of haemagglutinin, is immunosubdominant if it is associated with an immunodominant haemagglutinin globular head domain194,195 (Fig. 5a–c). However, it is possible to restore neuraminidase immunogenicity by using neuraminidase-only vaccines195,196,197 (Fig. 5d).
In association with haemagglutinin (HA) (the globular head domain is coloured blue and the stalk domain is coloured green), the viral neuraminidase (NA; black) is immunologically subdominant and the majority of the immune response is directed against the HA globular head domain (part a). Strategies to improve the immunogenicity of NA include supplementing seasonal trivalent inactivated vaccines with recombinant NA (part b), the use of chimeric HA (cHA)-based vaccines (red and green) that break the immunodominance of the HA globular head domain (part c) or vaccination with recombinant or purified NA only (part d). Structures are based on RCSB Protein Data Bank IDs 1RU7 (HA) and 3B7E (NA), and were visualized using Protein Workshop231,232.
Recent studies in ferrets using neuraminidase-only immunogens that induce high titres of anti-neuraminidase immunity clearly showed crossprotection to viruses expressing divergent N1 neuraminidases198. An alternative strategy to increase neuraminidase immunity would be to decrease the immunodominance of the associated haemagglutinin globular head. H7N2 vaccines can boost anti-neuraminidase immunity to high titres in humans, whereas control H3N2 vaccines have failed to do so153,199. As discussed above, the H7 globular head domain appears to be less immunodominant in humans who are naive to this subtype. It could be hypothesized that LAIV-based or IIV-based chimeric haemagglutinin vaccines that have an associated neuraminidase could also induce high titres of anti-neuraminidase immunity. In such a scenario, the immunodominance of the haemagglutinin head domain is also reduced (Fig. 5c). In conclusion, vaccine approaches that induce strong anti-neuraminidase immune responses could improve protection against homologous and heterologous influenza virus strains and would certainly represent a valuable addition to the armamentarium to fight influenza virus infections.
M2 is the third influenza virus surface transmembrane protein and is also of interest for the development of broadly protective influenza virus vaccines. Specifically, the 22–23-amino-acid short ectodomain of M2 (M2e) is promising because of its high conservation and surface exposure200. The development of M2e-based vaccines began in 1999 (Ref. 201) and since then many M2e vaccine constructs, including tetrameric M2e, VLP-displayed M2e, flagellin-fused M2e and multimeric M2e, have been successfully tested for efficacy against a panel of divergent influenza viruses201,202,203,204,205,206. M2 is present at very low copy numbers on virions but is abundant on infected cells. Protection conferred by M2e-based vaccines is probably mediated by ADCC200,207. M2e-specific antibodies are usually non-neutralizing and do not induce sterilizing immunity; however, passive transfer studies in humans demonstrated a reduction in clinical signs and nasal wash virus titres upon challenge with a human H3N2 influenza virus isolate208.
T-cell- or epitope-based universal influenza virus vaccines. Recently, a number of virus-vectored universal vaccine candidates have been developed. Several of these vaccines are based on MVA, which is an excellent platform to induce strong CD4 and CD8 T cell responses and is therefore preferentially used to boost cellular immunity. An MVA vector expressing a fusion protein of the conserved matrix (M1) and nucleoprotein has been tested in clinical trials and was found to be safe and effective in inducing cellular immune responses against influenza viruses209,210. However, the vaccine showed only weak protection in human challenge studies with an H3N2 strain211. Importantly, this study only assessed protection from mild upper respiratory infections, and the vaccine — owing to the nature of T-cell-based immunity — probably has a much stronger effect on lower respiratory infections with long durations (the study was stopped on day 5 post-infection using the antiviral drug oseltamivir)211. The same vaccine candidate is now being tested as an additive to a TIV and shows promising results in this context in preclinical experiments and clinical studies212,213. In addition, a prime–boost regimen with MVA and an adenovirus expressing M1-nucleoprotein showed successful induction of heterosubtypic immunity (Box 3) in mice214. A similar approach used an MVA vector expressing several influenza virus proteins — including haemagglutinin, neuraminidase, nucleoprotein, M1 and M2 — from H5N1 strains and interleukin-15 as a molecular adjuvant215. Challenge studies in mice showed antibody-independent heterosubtypic immunity against H1N1, H3N2 and H7N7 with an efficacy of 80–100% (Ref. 215). However, the mice experienced relatively high weight loss (between 15% and 20% of their initial weight)215.
In addition to viral vectors, numerous vaccine candidates, based on influenza viruses that are either severely attenuated or restricted to single-cycle replication, have been tested in recent years216,217,218. Heterosubtypic immunity has been demonstrated for these constructs — mostly in the absence of neutralizing antibodies — suggesting that T-cell-based protection was induced. Several vaccine candidates composed of single or multiple B- or T-cell epitopes are also in development219,220,221. A vaccine based on an E. coli-expressed fusion peptide containing different epitopes, Multimeric-001, has been tested in clinical trials and was found to be safe222. This vaccine candidate was also assessed in combination with regular TIV and was shown to induce T cell responses and increased haemagglutination inhibition responses to TIV strains in the elderly223.
Although many of these T-cell-based approaches might have the potential to protect from severe morbidity and mortality224,225,226, it is unclear whether they would also protect from the upper respiratory infection that drives transmission of the virus. Furthermore, it is unclear how long protective T cell responses against influenza viruses last. These questions will most likely be addressed in future clinical trials.
Conclusions
Both seasonal and pandemic influenza virus vaccines and vaccine production processes have been significantly improved since the 2009 H1N1 pandemic. Novel production platforms that enable rapid production have been established and several improved influenza virus vaccines have been licensed by the US Food and Drug Administration. Furthermore, the development of novel technologies for a detailed analysis of the human immune response to influenza virus infection and vaccination has led to an improved understanding of protection against influenza. It is now imperative to translate this knowledge into vaccines that provide broad protection from influenza virus infection and, ideally, lifelong universal coverage against all influenza A and B virus strains.
Change history
06 March 2015
The citation number for reference 41, a World Health Organization monthly risk asssesment summary, was missing in the article text and reference list. This has been corrected in the online version. The statement at the end of the legend for Figure 1, related to the permission to adapt a figure in reference 227, has also been modified.
References
World Health Organization. Influenza (seasonal) fact sheet. World Health Organization [online], (2014).
Gerdil, C. The annual production cycle for influenza vaccine. Vaccine 21, 1776–1779 (2003).
Palese, P. Influenza: old and new threats. Nature Med. 10, S82–S87 (2004).
Johnson, N. P. & Mueller, J. Updating the accounts: global mortality of the 1918–1920 “Spanish” influenza pandemic. Bull. Hist. Med. 76, 105–115 (2002).
Palese, P. & Wang, T. T. Why do influenza virus subtypes die out? A hypothesis. MBio 2, e00150-11 (2011).
Racaniello, V. Pandemic influenza vaccine was too late in 2009. Virology Blog [online], (2010).
Krammer, F. & Palese, P. Universal influenza virus vaccines: need for clinical trials. Nature Immunol. 15, 3–5 (2014).
Francis, T., Salk, J. E., Pearson, H. E. & Brown, P. N. Protective effect of vaccination against induced influenza A. J. Clin. Invest. 24, 536–546 (1945).
Salk, J. E., Pearson, H. E., Brown, P. N. & Francis, T. Protective effect of vaccination against induced influenza B. J. Clin. Invest. 24, 547–553 (1945).
Salk, J. E. & Suriano, P. C. Importance of antigenic composition of influenza virus vaccine in protecting against the natural disease; observations during the winter of 1947–1948. Am. J. Public Health Nations Health 39, 345–355 (1949).
Payne, A. M. The influenza programme of WHO. Bull. World Health Organ. 8, 755–774 (1953).
Allison, J. E., Glezen, W. P., Taber, L. H., Paredes, A. & Webster, R. G. Reactogenicity and immunogenicity of bivalent influenza A and monovalent influenza B virus vaccines in high-risk children. J. Infect. Dis. 136, S672–S676 (1977).
Davenport, F. M. et al. Comparisons of serologic and febrile responses in humans to vaccination with influenza A viruses or their hemagglutinins. J. Lab. Clin. Med. 63, 5–13 (1964).
Jin, H. & Subbarao, K. Live attenuated influenza vaccine. Curr. Top. Microbiol. Immunol. 386, 181–204 (2014).
Maassab, H. F. Adaptation and growth characteristics of influenza virus at 25 °C. Nature 213, 612–614 (1967).
Alexandrova, G. I. et al. Study of live recombinant cold-adapted influenza bivalent vaccine of type A for use in children: an epidemiological control trial. Vaccine 4, 114–118 (1986).
Tricco, A. C. et al. Comparing influenza vaccine efficacy against mismatched and matched strains: a systematic review and meta-analysis. BMC Med. 11, 153 (2013).
Steinhoff, M. C. et al. Neonatal outcomes after influenza immunization during pregnancy: a randomized controlled trial. CMAJ 184, 645–653 (2012).
Sheffield, J. S. et al. Effect of influenza vaccination in the first trimester of pregnancy. Obstet. Gynecol. 120, 532–537 (2012).
Beyer, W. E. et al. Cochrane re-arranged: support for policies to vaccinate elderly people against influenza. Vaccine 31, 6030–6033 (2013).
Ohmit, S. E. et al. Influenza vaccine effectiveness in the community and the household. Clin. Infect. Dis. 56, 1363–1369 (2013).
Kissling, E. et al. Low and decreasing vaccine effectiveness against influenza A(H3) in 2011/12 among vaccination target groups in Europe: results from the I-MOVE multicentre case–control study. Euro Surveill. 18, 20390 (2013).
Clark, A. et al. A comparison of live and inactivated influenza A (H1N1) virus vaccines. 2. Long-term immunity. J. Hyg. (Lond.) 90, 361–370 (1983).
de Jong, J. C., Beyer, W. E., Palache, A. M., Rimmelzwaan, G. F. & Osterhaus, A. D. Mismatch between the 1997/1998 influenza vaccine and the major epidemic A(H3N2) virus strain as the cause of an inadequate vaccine-induced antibody response to this strain in the elderly. J. Med. Virol. 61, 94–99 (2000).
DiazGranados, C. A. et al. High-dose trivalent influenza vaccine compared to standard dose vaccine in elderly adults: safety, immunogenicity and relative efficacy during the 2009–2010 season. Vaccine 31, 861–866 (2013).
DiazGranados, C. A. et al. Efficacy of high-dose versus standard-dose influenza vaccine in older adults. N. Engl. J. Med. 371, 635–645 (2014).
O'Hagan, D. T., Ott, G. S., Nest, G. V., Rappuoli, R. & Giudice, G. D. The history of MF59® adjuvant: a phoenix that arose from the ashes. Expert Rev. Vaccines 12, 13–30 (2013).
Del Giudice, G. & Rappuoli, R. Inactivated and adjuvanted influenza vaccines. Curr. Top. Microbiol. Immunol. 386, 151–180 (2014).
Ledgerwood, J. E. AS03-adjuvanted influenza vaccine in elderly people. Lancet Infect. Dis. 13, 466–467 (2013).
Tinoco, J. C. et al. Immunogenicity, reactogenicity, and safety of inactivated quadrivalent influenza vaccine candidate versus inactivated trivalent influenza vaccine in healthy adults aged ≥18 years: a phase III, randomized trial. Vaccine 32, 1480–1487 (2014).
Jain, V. K. et al. Vaccine for prevention of mild and moderate-to-severe influenza in children. N. Engl. J. Med. 369, 2481–2491 (2013).
Esposito, S. & Principi, N. Vaccine for prevention of influenza in children. N. Engl. J. Med. 370, 1167 (2014).
Wei, C. J. et al. Induction of broadly neutralizing H1N1 influenza antibodies by vaccination. Science 329, 1060–1064 (2010).
Wei, C. J. et al. Elicitation of broadly neutralizing influenza antibodies in animals with previous influenza exposure. Sci. Transl. Med. 4, 147ra114 (2012).
Kanekiyo, M. et al. Self-assembling influenza nanoparticle vaccines elicit broadly neutralizing H1N1 antibodies. Nature 499, 102–106 (2013).
Centers for Disease Control and Prevention. Table. Influenza vaccines — United States, 2014–15 influenza season. Centers for Disease Control and Prevention [online], (2014).
US Food and Drug Administration. FDA approves new seasonal influenza vaccine made using novel technology. US Food and Drug Administration [online], (2013).
Claas, E. C. et al. Human influenza A H5N1 virus related to a highly pathogenic avian influenza virus. Lancet 351, 472–477 (1998).
Vijaykrishna, D. et al. Evolutionary dynamics and emergence of panzootic H5N1 influenza viruses. PLoS Pathog. 4, e1000161 (2008).
Hatta, M., Gao, P., Halfmann, P. & Kawaoka, Y. Molecular basis for high virulence of Hong Kong H5N1 influenza A viruses. Science 293, 1840–1842 (2001).
World Health Organization. The WHO Influenza Monthly Risk Assessment Summaries. World Health Organization [online], (2015).
Wang, T. T., Parides, M. K. & Palese, P. Seroevidence for H5N1 influenza infections in humans: meta-analysis. Science 335, 1463 (2012).
Garten, R. et al. Antigenic and genetic characteristics of swine-origin 2009 A(H1N1) influenza viruses circulating in humans. Science 325, 197–201 (2009).
Gao, R. et al. Human infection with a novel avian-origin influenza A (H7N9) virus. N. Engl. J. Med. 368, 1888–1897 (2013).
Wei, S. H. et al. Human infection with avian influenza A H6N1 virus: an epidemiological analysis. Lancet Respir. Med. 1, 771–778 (2013).
García-Sastre, A. & Schmolke, M. Avian influenza A H10N8 — a virus on the verge? Lancet 383, 676–677 (2014).
Chen, H. et al. Clinical and epidemiological characteristics of a fatal case of avian influenza A H10N8 virus infection: a descriptive study. Lancet 383, 714–721 (2014).
Park, M. World's first H5N6 bird flu death reported in China. CNN [online], (2014).
Centers for Disease Control and Prevention (CDC). Notes from the field: outbreak of influenza A (H3N2) virus among persons and swine at a county fair — Indiana, July 2012. MMWR Morb. Mortal. Wkly Rep. 61, 561 (2012).
Lopez-Martinez, I. et al. Highly pathogenic avian influenza A(H7N3) virus in poultry workers, Mexico, 2012. Emerg. Infect. Dis. 19, 1531–1534 (2013).
Anthony, S. J. et al. Emergence of fatal avian influenza in New England harbor seals. MBio 3, e00166-12 (2012).
Zohari, S., Neimanis, A., Harkonen, T., Moraeus, C. & Valarcher, J. Avian influenza A(H10N7) virus involvement in mass mortality of harbour seals (Phoca vitulina) in Sweden, March through October 2014. Euro Surveill. 19, 20967 (2014).
[No authors listed.] Avian influenza outbreak in Yorkshire: strain identified as H5N8. Vet. Rec. 175, 495–496 (2014).
Krammer, F. & Cox, R. J. The emergence of H7N9 viruses: a chance to redefine correlates of protection for influenza virus vaccines. Expert Rev. Vaccines 12, 1369–1372 (2013).
Cox, R. J. et al. A phase I clinical trial of a PER.C6® cell grown influenza H7 virus vaccine. Vaccine 27, 1889–1897 (2009).
Couch, R. B. et al. Evaluations for in vitro correlates of immunogenicity of inactivated influenza a H5, H7 and H9 vaccines in humans. PLoS ONE 7, e50830 (2012).
Couch, R. B., Patel, S. M., Wade-Bowers, C. L. & Niño, D. A randomized clinical trial of an inactivated avian influenza A (H7N7) vaccine. PLoS ONE 7, e49704 (2012).
Kistner, O. et al. Cell culture (Vero) derived whole virus (H5N1) vaccine based on wild-type virus strain induces cross-protective immune responses. Vaccine 25, 6028–6036 (2007).
Baz, M., Luke, C. J., Cheng, X., Jin, H. & Subbarao, K. H5N1 vaccines in humans. Virus Res. 178, 78–98 (2013).
Belshe, R. B. et al. Immunogenicity of avian influenza A/Anhui/01/2005(H5N1) vaccine with MF59 adjuvant: a randomized clinical trial. JAMA 312, 1420–1428 (2014).
Mulligan, M. J. et al. Serological responses to an avian influenza A/H7N9 vaccine mixed at the point-of-use with MF59 adjuvant: a randomized clinical trial. JAMA 312, 1409–1419 (2014).
Krammer, F. et al. An H7N1 influenza virus vaccine induces broadly reactive antibody responses against H7N9 in humans. Clin. Vaccine Immunol. 21, 1153–1163 (2014).
Ellebedy, A. H. et al. Induction of broadly cross-reactive antibody responses to the influenza HA stem region following H5N1 vaccination in humans. Proc. Natl Acad. Sci. USA 111, 13133–13138 (2014).
Nachbagauer, R. et al. Induction of broadly-reactive anti-hemagglutinin stalk antibodies by an H5N1 vaccine in humans. J. Virol. 88, 13260–13268 (2014).
Karron, R. A. et al. Evaluation of two live attenuated cold-adapted H5N1 influenza virus vaccines in healthy adults. Vaccine 27, 4953–4960 (2009).
Talaat, K. R. et al. A live attenuated H7N3 influenza virus vaccine is well tolerated and immunogenic in a phase I trial in healthy adults. Vaccine 27, 3744–3753 (2009).
Talaat, K. R. et al. An open label phase I trial of a live attenuated H6N1 influenza virus vaccine in healthy adults. Vaccine 29, 3144–3148 (2011).
Talaat, K. R. et al. An open-label phase I trial of a live attenuated H2N2 influenza virus vaccine in healthy adults. Influenza Other Respir. Viruses 7, 66–73 (2013).
Rudenko, L. et al. Assessment of human immune responses to H7 avian influenza virus of pandemic potential: results from a placebo-controlled, randomized double-blind phase I study of live attenuated H7N3 influenza vaccine. PLoS ONE 9, e87962 (2014).
Rudenko, L., Isakova-Sivak, I. & Donina, S. H7N3 live attenuated influenza vaccine has a potential to protect against new H7N9 avian influenza virus. Vaccine 31, 4702–4705 (2013).
Rudenko, L. et al. Safety and immunogenicity of live attenuated influenza reassortant H5 vaccine (phase I–II clinical trials). Influenza Other Respir. Viruses 2, 203–209 (2008).
Matsuoka, Y. et al. African green monkeys recapitulate the clinical experience with replication of live attenuated pandemic influenza virus vaccine candidates. J. Virol. 88, 8139–8152 (2014).
Min, J. Y. et al. A live attenuated H7N7 candidate vaccine virus induces neutralizing antibody that confers protection from challenge in mice, ferrets, and monkeys. J. Virol. 84, 11950–11960 (2010).
Steel, J. et al. Live attenuated influenza viruses containing NS1 truncations as vaccine candidates against H5N1 highly pathogenic avian influenza. J. Virol. 83, 1742–1753 (2009).
de Graaf, M. & Fouchier, R. A. Role of receptor binding specificity in influenza A virus transmission and pathogenesis. EMBO J. 33, 823–841 (2014).
Ledgerwood, J. E. et al. Prime-boost interval matters: a randomized phase 1 study to identify the minimum interval necessary to observe the H5 DNA influenza vaccine priming effect. J. Infect. Dis. 208, 418–422 (2013).
Talaat, K. R. et al. A live attenuated influenza A(H5N1) vaccine induces long-term immunity in the absence of a primary antibody response. J. Infect. Dis. 209, 1860–1869 (2014).
Luke, C. J. & Subbarao, K. Improving pandemic H5N1 influenza vaccines by combining different vaccine platforms. Expert Rev. Vaccines 13, 873–883 (2014).
Kistner, O. et al. Development of a mammalian cell (Vero) derived candidate influenza virus vaccine. Vaccine 16, 960–968 (1998).
Dormitzer, P. R. Rapid production of synthetic influenza vaccines. Curr. Top. Microbiol. Immunol. 386, 237–273 (2015).
Fodor, E. et al. Rescue of influenza A virus from recombinant DNA. J. Virol. 73, 9679–9682 (1999).
Cox, M. M. Recombinant protein vaccines produced in insect cells. Vaccine 30, 1759–1766 (2012).
Krammer, F. & Grabherr, R. Alternative influenza vaccines made by insect cells. Trends Mol. Med. 16, 313–320 (2010).
Jul-Larsen, Å. et al. The human potential of a recombinant pandemic influenza vaccine produced in tobacco plants. Hum. Vaccin. Immunother. 8, 653–661 (2012).
Aguilar-Yáñez, J. M. et al. An influenza A/H1N1/2009 hemagglutinin vaccine produced in Escherichia coli. PLoS ONE 5, e11694 (2010).
Taylor, D. N. et al. Development of VAX128, a recombinant hemagglutinin (HA) influenza–flagellin fusion vaccine with improved safety and immune response. Vaccine 30, 5761–5769 (2012).
Shi, S. H. et al. Immunoprotection against influenza virus H9N2 by the oral administration of recombinant Lactobacillus plantarum NC8 expressing hemagglutinin in BALB/c mice. Virology 464–465, 166–176 (2014).
Bayne, A. C. et al. Vaccination against influenza with recombinant hemagglutinin expressed by Schizochytrium sp. confers protective immunity. PLoS ONE 8, e61790 (2013).
Saelens, X. et al. Protection of mice against a lethal influenza virus challenge after immunization with yeast-derived secreted influenza virus hemagglutinin. Eur. J. Biochem. 260, 166–175 (1999).
Murugan, S. et al. Recombinant haemagglutinin protein of highly pathogenic avian influenza A (H5N1) virus expressed in Pichia pastoris elicits a neutralizing antibody response in mice. J. Virol. Methods 187, 20–25 (2013).
Welsh, J. P., Lu, Y., He, X. S., Greenberg, H. B. & Swartz, J. R. Cell-free production of trimeric influenza hemagglutinin head domain proteins as vaccine antigens. Biotechnol. Bioeng. 109, 2962–2969 (2012).
DuBois, R. M. et al. The receptor-binding domain of influenza virus hemagglutinin produced in Escherichia coli folds into its native, immunogenic structure. J. Virol. 85, 865–872 (2011).
Khurana, S. et al. H5N1 virus-like particle vaccine elicits cross-reactive neutralizing antibodies in humans that preferentially bind to oligomeric form of influenza hemagglutinin. J. Virol. 85, 10945–10954 (2011).
López-Macías, C. et al. Safety and immunogenicity of a virus-like particle pandemic influenza A (H1N1) 2009 vaccine in a blinded, randomized, placebo-controlled trial of adults in Mexico. Vaccine 29, 7826–7834 (2011).
Krammer, F. et al. Swine-origin pandemic H1N1 influenza virus-like particles produced in insect cells induce hemagglutination inhibiting antibodies in BALB/c mice. Biotechnol. J. 5, 17–23 (2010).
Smith, G. E. et al. Development of influenza H7N9 virus like particle (VLP) vaccine: homologous A/Anhui/1/2013 (H7N9) protection and heterologous A/chicken/Jalisco/CPA1/2012 (H7N3) cross-protection in vaccinated mice challenged with H7N9 virus. Vaccine 31, 4305–4313 (2013).
Klausberger, M. et al. One-shot vaccination with an insect cell-derived low-dose influenza A H7 virus-like particle preparation protects mice against H7N9 challenge. Vaccine 32, 355–362 (2014).
Margine, I., Martinez-Gil, L., Chou, Y. Y. & Krammer, F. Residual baculovirus in insect cell-derived influenza virus-like particle preparations enhances immunogenicity. PLoS ONE 7, e51559 (2012).
D'Aoust, M. et al. Influenza virus-like particles produced by transient expression in Nicotiana benthamiana induce a protective immune response against a lethal viral challenge in mice. Plant Biotechnol. J. 6, 930–940 (2008).
D'Aoust, M. A. et al. The production of hemagglutinin-based virus-like particles in plants: a rapid, efficient and safe response to pandemic influenza. Plant Biotechnol. J. 8, 607–619 (2010).
Fries, L. F., Smith, G. E. & Glenn, G. M. A recombinant viruslike particle influenza A (H7N9) vaccine. N. Engl. J. Med. 369, 2564–2566 (2013).
Landry, N. et al. Preclinical and clinical development of plant-made virus-like particle vaccine against avian H5N1 influenza. PLoS ONE 5, e15559 (2010).
Ledgerwood, J. E. et al. Influenza virus H5 DNA vaccination is immunogenic by intramuscular and intradermal routes in humans. Clin. Vaccine Immunol. 19, 1792–1797 (2012).
Tripp, R. A. & Tompkins, S. M. Virus-vectored influenza virus vaccines. Viruses 6, 3055–3079 (2014).
Rimmelzwaan, G. F. & Sutter, G. Candidate influenza vaccines based on recombinant modified vaccinia virus Ankara. Expert Rev. Vaccines 8, 447–454 (2009).
Kreijtz, J. H. et al. Evaluation of a modified vaccinia virus Ankara (MVA)-based candidate pandemic influenza A/H1N1 vaccine in the ferret model. J. Gen. Virol. 91, 2745–2752 (2010).
Altenburg, A. F. et al. Modified vaccinia virus Ankara (MVA) as production platform for vaccines against influenza and other viral respiratory diseases. Viruses 6, 2735–2761 (2014).
Kreijtz, J. H. et al. Recombinant modified vaccinia virus Ankara expressing the hemagglutinin gene confers protection against homologous and heterologous H5N1 influenza virus infections in macaques. J. Infect. Dis. 199, 405–413 (2009).
Kreijtz, J. H. et al. A single immunization with an MVA-based influenza virus H7 vaccine affords protection in the H7N9 pneumonia ferret model. J. Infect. Dis. http://dx.doi.org/10.1093/infdis/jiu528 (2014).
Prabakaran, M. et al. Progress toward a universal H5N1 vaccine: a recombinant modified vaccinia virus Ankara-expressing trivalent hemagglutinin vaccine. PLoS ONE 9, e107316 (2014).
Kreijtz, J. H. et al. Safety and immunogenicity of a modified-vaccinia-virus-Ankara-based influenza A H5N1 vaccine: a randomised, double-blind phase 1/2a clinical trial. Lancet Infect. Dis. 14, 1196–1207 (2014).
Wrammert, J. et al. Broadly cross-reactive antibodies dominate the human B cell response against 2009 pandemic H1N1 influenza virus infection. J. Exp. Med. 208, 181–193 (2011).
Wrammert, J. et al. Rapid cloning of high-affinity human monoclonal antibodies against influenza virus. Nature 453, 667–671 (2008).
Moody, M. A. et al. H3N2 influenza infection elicits more cross-reactive and less clonally expanded anti-hemagglutinin antibodies than influenza vaccination. PLoS ONE 6, e25797 (2011).
Throsby, M. et al. Heterosubtypic neutralizing monoclonal antibodies cross-protective against H5N1 and H1N1 recovered from human IgM+ memory B cells. PLoS ONE 3, e3942 (2008).
Sui, J. et al. Structural and functional bases for broad-spectrum neutralization of avian and human influenza A viruses. Nature Struct. Mol. Biol. 16, 265–273 (2009).
Corti, D. et al. Heterosubtypic neutralizing antibodies are produced by individuals immunized with a seasonal influenza vaccine. J. Clin. Invest. 120, 1663–1673 (2010).
Okuno, Y., Isegawa, Y., Sasao, F. & Ueda, S. A common neutralizing epitope conserved between the hemagglutinins of influenza A virus H1 and H2 strains. J. Virol. 67, 2552–2558 (1993).
Kashyap, A. K. et al. Combinatorial antibody libraries from survivors of the Turkish H5N1 avian influenza outbreak reveal virus neutralization strategies. Proc. Natl Acad. Sci. USA 105, 5986–5991 (2008).
Wan, H. et al. Molecular basis for broad neuraminidase immunity: conserved epitopes in seasonal and pandemic H1N1 as well as H5N1 influenza viruses. J. Virol. 87, 9290–9300 (2013).
Krause, J. C. et al. A broadly neutralizing human monoclonal antibody that recognizes a conserved, novel epitope on the globular head of the influenza H1N1 virus hemagglutinin. J. Virol. 85, 10905–10908 (2011).
Krause, J. C. et al. Human monoclonal antibodies to pandemic 1957 H2N2 and pandemic 1968 H3N2 influenza viruses. J. Virol. 86, 6334–6340 (2012).
Ekiert, D. C. et al. Cross-neutralization of influenza A viruses mediated by a single antibody loop. Nature 489, 526–532 (2012).
Whittle, J. R. et al. Broadly neutralizing human antibody that recognizes the receptor-binding pocket of influenza virus hemagglutinin. Proc. Natl Acad. Sci. USA 108, 14216–14221 (2011).
Ohshima, N. et al. Naturally occurring antibodies in a human can neutralize a broad spectrum of influenza strains including H3, H1, H2 and H5. J. Virol. 85, 11048–11057 (2011).
Ekiert, D. C. et al. Antibody recognition of a highly conserved influenza virus epitope. Science 324, 246–251 (2009).
Friesen, R. H. et al. A common solution to group 2 influenza virus neutralization. Proc. Natl Acad. Sci. USA 111, 445–450 (2014).
Ekiert, D. C. et al. A highly conserved neutralizing epitope on group 2 influenza A viruses. Science 333, 843–850 (2011).
Tan, G. S. et al. Characterization of a broadly neutralizing monoclonal antibody that targets the fusion domain of group 2 influenza A virus hemagglutinin. J. Virol. 88, 13580–13592 (2014).
Tan, G. S. et al. A pan-h1 anti-hemagglutinin monoclonal antibody with potent broad-spectrum efficacy in vivo. J. Virol. 86, 6179–6188 (2012).
Wang, T. T. et al. Broadly protective monoclonal antibodies against H3 influenza viruses following sequential immunization with different hemagglutinins. PLoS Pathog. 6, e1000796 (2010).
Dreyfus, C. et al. Highly conserved protective epitopes on influenza B viruses. Science 337, 1343–1348 (2012).
Corti, D. et al. A neutralizing antibody selected from plasma cells that binds to group 1 and group 2 influenza A hemagglutinins. Science 333, 850–856 (2011).
Nakamura, G. et al. An in vivo human-plasmablast enrichment technique allows rapid identification of therapeutic influenza A antibodies. Cell Host Microbe 14, 93–103 (2013).
Krammer, F. et al. A carboxy-terminal trimerization domain stabilizes conformational epitopes on the stalk domain of soluble recombinant hemagglutinin substrates. PLoS ONE 7, e43603 (2012).
Harris, A. K. et al. Structure and accessibility of HA trimers on intact 2009 H1N1 pandemic influenza virus to stem region-specific neutralizing antibodies. Proc. Natl Acad. Sci. USA 110, 4592–4597 (2013).
Brandenburg, B. et al. Mechanisms of hemagglutinin targeted influenza virus neutralization. PLoS ONE 8, e80034 (2013).
Terajima, M. et al. Complement-dependent lysis of influenza A virus-infected cells by broadly cross-reactive human monoclonal antibodies. J. Virol. 85, 13463–13467 (2011).
Dilillo, D. J., Tan, G. S., Palese, P. & Ravetch, J. V. Broadly neutralizing hemagglutinin stalk-specific antibodies require FcγR interactions for protection against influenza virus in vivo. Nature Med. 20, 143–151 (2014).
Jegaskanda, S., Weinfurter, J. T., Friedrich, T. C. & Kent, S. J. Antibody-dependent cellular cytotoxicity is associated with control of pandemic H1N1 influenza virus infection of macaques. J. Virol. 87, 5512–5522 (2013).
Jegaskanda, S. et al. Cross-reactive influenza-specific antibody-dependent cellular cytotoxicity antibodies in the absence of neutralizing antibodies. J. Immunol. 190, 1837–1848 (2013).
Jegaskanda, S., Reading, P. C. & Kent, S. J. Influenza-specific antibody-dependent cellular cytotoxicity: toward a universal influenza vaccine. J. Immunol. 193, 469–475 (2014).
Margine, I. et al. H3N2 influenza virus infection induces broadly reactive hemagglutinin stalk antibodies in humans and mice. J. Virol. 87, 4728–4737 (2013).
Krammer, F., Pica, N., Hai, R., Tan, G. S. & Palese, P. Hemagglutinin stalk-reactive antibodies are boosted following sequential infection with seasonal and pandemic H1N1 influenza virus in mice. J. Virol. 86, 10302–10307 (2012).
Miller, M. S. et al. Neutralizing antibodies against previously encountered influenza virus strains increase over time: a longitudinal analysis. Sci. Transl. Med. 5, 198ra107 (2013).
Pica, N. et al. Hemagglutinin stalk antibodies elicited by the 2009 pandemic influenza virus as a mechanism for the extinction of seasonal H1N1 viruses. Proc. Natl Acad. Sci. USA 109, 2573–2578 (2012).
Thomson, C. A. et al. Pandemic H1N1 influenza infection and vaccination in humans induces cross-protective antibodies that target the hemagglutinin stem. Front. Immunol. 3, 87 (2012).
Miller, M. S. et al. 1976 and 2009 H1N1 influenza virus vaccines boost anti-hemagglutinin stalk antibodies in humans. J. Infect. Dis. 207, 98–105 (2013).
Sangster, M. Y. et al. B cell response and hemagglutinin stalk-reactive antibody production in different age cohorts following 2009 H1N1 influenza virus vaccination. Clin. Vaccine Immunol. 20, 867–876 (2013).
Whittle, J. R. et al. Flow cytometry reveals that H5N1 vaccination elicits cross-reactive stem-directed antibodies from multiple Ig heavy-chain lineages. J. Virol. 88, 4047–4057 (2014).
Hensley, S. E. Challenges of selecting seasonal influenza vaccine strains for humans with diverse pre-exposure histories. Curr. Opin. Virol. 8, 85–89 (2014).
Li, Y. et al. Immune history shapes specificity of pandemic H1N1 influenza antibody responses. J. Exp. Med. 210, 1493–1500 (2013).
Wohlbold, T. J. & Krammer, F. In the shadow of hemagglutinin: a growing interest in influenza viral neuraminidase and its role as a vaccine antigen. Viruses 6, 2465–2494 (2014).
Doyle, T. M. et al. A monoclonal antibody targeting a highly conserved epitope in influenza B neuraminidase provides protection against drug resistant strains. Biochem. Biophys. Res. Commun. 441, 226–229 (2013).
Doyle, T. M. et al. Universal anti-neuraminidase antibody inhibiting all influenza A subtypes. Antiviral Res. 100, 567–574 (2013).
Tate, M. D. et al. Playing hide and seek: how glycosylation of the influenza virus hemagglutinin can modulate the immune response to infection. Viruses 6, 1294–1316 (2014).
Medina, R. A. et al. Glycosylations in the globular head of the hemagglutinin protein modulate the virulence and antigenic properties of the H1N1 influenza viruses. Sci. Transl. Med. 5, 187ra70 (2013).
Wang, C. C. et al. Glycans on influenza hemagglutinin affect receptor binding and immune response. Proc. Natl Acad. Sci. USA 106, 18137–18142 (2009).
Chen, J. R. et al. Vaccination of monoglycosylated hemagglutinin induces cross-strain protection against influenza virus infections. Proc. Natl Acad. Sci. USA 111, 2476–2481 (2014).
Magadán, J. G. et al. Biogenesis of influenza A virus hemagglutinin cross-protective stem epitopes. PLoS Pathog. 10, e1004204 (2014).
An, Y. et al. Comparative glycomics analysis of influenza hemagglutinin (H5N1) produced in vaccine relevant cell platforms. J. Proteome Res. 12, 3707–3720 (2013).
Palmberger, D., Ashjaei, K., Strell, S., Hoffmann-Sommergruber, K. & Grabherr, R. Minimizing fucosylation in insect cell-derived glycoproteins reduces binding to IgE antibodies from the sera of patients with allergy. Biotechnol. J. 9, 1206–1214 (2014).
Eggink, D., Goff, P. H. & Palese, P. Guiding the immune response against influenza virus hemagglutinin toward the conserved stalk domain by hyperglycosylation of the globular head domain. J. Virol. 88, 699–704 (2014).
Lin, S. C., Lin, Y. F., Chong, P. & Wu, S. C. Broader neutralizing antibodies against H5N1 viruses using prime-boost immunization of hyperglycosylated hemagglutinin DNA and virus-like particles. PLoS ONE 7, e39075 (2012).
Lin, S. C., Liu, W. C., Jan, J. T. & Wu, S. C. Glycan masking of hemagglutinin for adenovirus vector and recombinant protein immunizations elicits broadly neutralizing antibodies against H5N1 avian influenza viruses. PLoS ONE 9, e92822 (2014).
Graves, P. N., Schulman, J. L., Young, J. F. & Palese, P. Preparation of influenza virus subviral particles lacking the HA1 subunit of hemagglutinin: unmasking of cross-reactive HA2 determinants. Virology 126, 106–116 (1983).
Sagawa, H., Ohshima, A., Kato, I., Okuno, Y. & Isegawa, Y. The immunological activity of a deletion mutant of influenza virus haemagglutinin lacking the globular region. J. Gen. Virol. 77, 1483–1487 (1996).
Steel, J. et al. Influenza virus vaccine based on the conserved hemagglutinin stalk domain. MBio 1, e00018-10 (2010).
Bommakanti, G. et al. Design of an HA2-based Escherichia coli expressed influenza immunogen that protects mice from pathogenic challenge. Proc. Natl Acad. Sci. USA 107, 13701–13706 (2010).
Bommakanti, G. et al. Design of Escherichia coli-expressed stalk domain immunogens of H1N1 hemagglutinin that protect mice from lethal challenge. J. Virol. 86, 13434–13444 (2012).
Mallajosyula, V. V. et al. Influenza hemagglutinin stem-fragment immunogen elicits broadly neutralizing antibodies and confers heterologous protection. Proc. Natl Acad. Sci. USA 111, E2514–E2523 (2014).
Lu, Y., Welsh, J. P. & Swartz, J. R. Production and stabilization of the trimeric influenza hemagglutinin stem domain for potentially broadly protective influenza vaccines. Proc. Natl Acad. Sci. USA 111, 125–130 (2014).
Staneková, Z. et al. Heterosubtypic protection against influenza A induced by adenylate cyclase toxoids delivering conserved HA2 subunit of hemagglutinin. Antiviral Res. 97, 24–35 (2013).
Stropkovská, A. et al. Broadly cross-reactive monoclonal antibodies against HA2 glycopeptide of influenza A virus hemagglutinin of H3 subtype reduce replication of influenza A viruses of human and avian origin. Acta Virol. 53, 15–20 (2009).
Hai, R. et al. Influenza viruses expressing chimeric hemagglutinins: globular head and stalk domains derived from different subtypes. J. Virol. 86, 5774–5781 (2012).
Krammer, F. & Palese, P. Influenza virus hemagglutinin stalk-based antibodies and vaccines. Curr. Opin. Virol. 3, 521–530 (2013).
Krammer, F. et al. Assessment of influenza virus hemagglutinin stalk-based immunity in ferrets. J. Virol. 88, 3432–3442 (2014).
Krammer, F., Pica, N., Hai, R., Margine, I. & Palese, P. Chimeric hemagglutinin influenza virus vaccine constructs elicit broadly protective stalk-specific antibodies. J. Virol. 87, 6542–6550 (2013).
Krammer, F. et al. H3 stalk-based chimeric hemagglutinin influenza virus constructs protect mice from H7N9 challenge. J. Virol. 88, 2340–2343 (2014).
Margine, I. et al. Hemagglutinin stalk-based universal vaccine constructs protect against group 2 influenza A viruses. J. Virol. 87, 10435–10446 (2013).
Goff, P. H. et al. Adjuvants and immunization strategies to induce influenza virus hemagglutinin stalk antibodies. PLoS ONE 8, e79194 (2013).
Weaver, E. A., Rubrum, A. M., Webby, R. J. & Barry, M. A. Protection against divergent influenza H1N1 virus by a centralized influenza hemagglutinin. PLoS ONE 6, e18314 (2011).
Kirchenbaum, G. A. & Ross, T. M. Eliciting broadly protective antibody responses against influenza. Curr. Opin. Immunol. 28, 71–76 (2014).
Ducatez, M. F. et al. Feasibility of reconstructed ancestral H5N1 influenza viruses for cross-clade protective vaccine development. Proc. Natl Acad. Sci. USA 108, 349–354 (2011).
Giles, B. M. & Ross, T. M. Computationally optimized antigens to overcome influenza viral diversity. Expert Rev. Vaccines 11, 267–269 (2012).
Giles, B. M., Bissel, S. J., Dealmeida, D. R., Wiley, C. A. & Ross, T. M. Antibody breadth and protective efficacy are increased by vaccination with computationally optimized hemagglutinin but not with polyvalent hemagglutinin-based H5N1 virus-like particle vaccines. Clin. Vaccine Immunol. 19, 128–139 (2012).
Giles, B. M. & Ross, T. M. A computationally optimized broadly reactive antigen (COBRA) based H5N1 VLP vaccine elicits broadly reactive antibodies in mice and ferrets. Vaccine 29, 3043–3054 (2011).
Giles, B. M. et al. A computationally optimized hemagglutinin virus-like particle vaccine elicits broadly reactive antibodies that protect nonhuman primates from H5N1 infection. J. Infect. Dis. 205, 1562–1570 (2012).
Kilbourne, E. D., Johansson, B. E. & Grajower, B. Independent and disparate evolution in nature of influenza A virus hemagglutinin and neuraminidase glycoproteins. Proc. Natl Acad. Sci. USA 87, 786–790 (1990).
Abed, Y., Hardy, I., Li, Y. & Boivin, G. Divergent evolution of hemagglutinin and neuraminidase genes in recent influenza A:H3N2 viruses isolated in Canada. J. Med. Virol. 67, 589–595 (2002).
Westgeest, K. B. et al. Genetic evolution of the neuraminidase of influenza A (H3N2) viruses from 1968 to 2009 and its correspondence to haemagglutinin evolution. J. Gen. Virol. 93, 1996–2007 (2012).
Sultana, I. et al. Stability of neuraminidase in inactivated influenza vaccines. Vaccine 32, 2225–2230 (2014).
Couch, R. B., Kasel, J. A., Gerin, J. L., Schulman, J. L. & Kilbourne, E. D. Induction of partial immunity to influenza by a neuraminidase-specific vaccine. J. Infect. Dis. 129, 411–420 (1974).
Johansson, B. E., Moran, T. M. & Kilbourne, E. D. Antigen-presenting B cells and helper T cells cooperatively mediate intravirionic antigenic competition between influenza A virus surface glycoproteins. Proc. Natl Acad. Sci. USA 84, 6869–6873 (1987).
Johansson, B. E. & Kilbourne, E. D. Dissociation of influenza virus hemagglutinin and neuraminidase eliminates their intravirionic antigenic competition. J. Virol. 67, 5721–5723 (1993).
Johansson, B. E. & Kilbourne, E. D. Immunization with purified N1 and N2 influenza virus neuraminidases demonstrates cross-reactivity without antigenic competition. Proc. Natl Acad. Sci. USA 91, 2358–2361 (1994).
Kilbourne, E. D. et al. Purified influenza A virus N2 neuraminidase vaccine is immunogenic and non-toxic in humans. Vaccine 13, 1799–1803 (1995).
Easterbrook, J. D. et al. Protection against a lethal H5N1 influenza challenge by intranasal immunization with virus-like particles containing 2009 pandemic H1N1 neuraminidase in mice. Virology 432, 39–44 (2012).
Kilbourne, E. D., Cerini, C. P., Khan, M. W., Mitchell, J. W. & Ogra, P. L. Immunologic response to the influenza virus neuraminidase is influenced by prior experience with the associated viral hemagglutinin. I. Studies in human vaccinees. J. Immunol. 138, 3010–3013 (1987).
Schotsaert, M., De Filette, M., Fiers, W. & Saelens, X. Universal M2 ectodomain-based influenza A vaccines: preclinical and clinical developments. Expert Rev. Vaccines 8, 499–508 (2009).
Neirynck, S. et al. A universal influenza A vaccine based on the extracellular domain of the M2 protein. Nature Med. 5, 1157–1163 (1999).
De Filette, M. et al. An influenza A vaccine based on tetrameric ectodomain of matrix protein 2. J. Biol. Chem. 283, 11382–11387 (2008).
Wang, L. et al. Nanoclusters self-assembled from conformation-stabilized influenza M2e as broadly cross-protective influenza vaccines. Nanomedicine 10, 473–482 (2014).
Huleatt, J. et al. Potent immunogenicity and efficacy of a universal influenza vaccine candidate comprising a recombinant fusion protein linking influenza M2e to the TLR5 ligand flagellin. Vaccine 26, 201–214 (2008).
De Filette, M. et al. Universal influenza A vaccine: optimization of M2-based constructs. Virology 337, 149–161 (2005).
Ebrahimi, S. M., Dabaghian, M., Tebianian, M. & Jazi, M. H. In contrast to conventional inactivated influenza vaccines, 4xM2e. HSP70c fusion protein fully protected mice against lethal dose of H1, H3 and H9 influenza A isolates circulating in Iran. Virology 430, 63–72 (2012).
El Bakkouri, K. et al. Universal vaccine based on ectodomain of matrix protein 2 of influenza A: Fc receptors and alveolar macrophages mediate protection. J. Immunol. 186, 1022–1031 (2011).
Ramos, E. L. et al. Efficacy and safety of treatment with an anti-M2e monoclonal antibody in experimental human influenza. J. Infect. Dis. http://dx.doi.org/10.1093/infdis/jiu539 (2014).
Berthoud, T. K. et al. Potent CD8+ T-cell immunogenicity in humans of a novel heterosubtypic influenza A vaccine, MVA–NP+M1. Clin. Infect. Dis. 52, 1–7 (2011).
Antrobus, R. D. et al. A T cell-inducing influenza vaccine for the elderly: safety and immunogenicity of MVA–NP+M1 in adults aged over 50 years. PLoS ONE 7, e48322 (2012).
Lillie, P. J. et al. Preliminary assessment of the efficacy of a T-cell-based influenza vaccine, MVA–NP+M1, in humans. Clin. Infect. Dis. 55, 19–25 (2012).
Mullarkey, C. E. et al. Improved adjuvanting of seasonal influenza vaccines: preclinical studies of MVA–NP+M1 coadministration with inactivated influenza vaccine. Eur. J. Immunol. 43, 1940–1952 (2013).
Antrobus, R. D. et al. Coadministration of seasonal influenza vaccine and MVA–NP+M1 simultaneously achieves potent humoral and cell-mediated responses. Mol. Ther. 22, 233–238 (2014).
Lambe, T. et al. Immunity against heterosubtypic influenza virus induced by adenovirus and MVA expressing nucleoprotein and matrix protein-1. Sci. Rep. 3, 1443 (2013).
Valkenburg, S. A. et al. IL-15 adjuvanted multivalent vaccinia-based universal influenza vaccine requires CD4+ T cells for heterosubtypic protection. Proc. Natl Acad. Sci. USA 111, 5676–5681 (2014).
Baker, S. F. et al. Protection against lethal influenza with a viral mimic. J. Virol. 87, 8591–8605 (2013).
Yang, C., Skiena, S., Futcher, B., Mueller, S. & Wimmer, E. Deliberate reduction of hemagglutinin and neuraminidase expression of influenza virus leads to an ultraprotective live vaccine in mice. Proc. Natl Acad. Sci. USA 110, 9481–9486 (2013).
Powell, T. J., Silk, J. D., Sharps, J., Fodor, E. & Townsend, A. R. Pseudotyped influenza A virus as a vaccine for the induction of heterotypic immunity. J. Virol. 86, 13397–13406 (2012).
Wang, T. T. et al. Vaccination with a synthetic peptide from the influenza virus hemagglutinin provides protection against distinct viral subtypes. Proc. Natl Acad. Sci. USA 107, 18979–18984 (2010).
Staneková, Z. et al. Heterosubtypic protective immunity against influenza A virus induced by fusion peptide of the hemagglutinin in comparison to ectodomain of M2 protein. Acta Virol. 55, 61–67 (2011).
Janulíková, J., Staneková, Z., Mucha, V., Kostolanský, F. & Varecková, E. Two distinct regions of HA2 glycopolypeptide of influenza virus hemagglutinin elicit cross-protective immunity against influenza. Acta Virol. 56, 169–176 (2012).
Atsmon, J. et al. Safety and immunogenicity of multimeric-001—a novel universal influenza vaccine. J. Clin. Immunol. 32, 595–603 (2012).
Atsmon, J. et al. Priming by a novel universal influenza vaccine (multimeric-001)—a gateway for improving immune response in the elderly population. Vaccine 32, 5816–5823 (2014).
Sridhar, S. et al. Cellular immune correlates of protection against symptomatic pandemic influenza. Nature Med. 19, 1305–1312 (2013).
Hillaire, M. L. et al. Cross-protective immunity against influenza pH1N1 2009 viruses induced by seasonal influenza A (H3N2) virus is mediated by virus-specific T-cells. J. Gen. Virol. 92, 2339–2349 (2011).
van de Sandt, C. E. et al. Human cytotoxic T lymphocytes directed to seasonal influenza A viruses cross-react with the newly emerging H7N9 virus. J. Virol. 88, 1684–1693 (2013).
World Health Organization. Pandemic influenza vaccine manufacturing process and timeline. World Health Organization [online], (2009).
Lee, P. S. et al. Receptor mimicry by antibody F045-092 facilitates universal binding to the H3 subtype of influenza virus. Nature Commun. 5, 3614 (2014).
Li, Q. et al. The 2009 pandemic H1N1 neuraminidase N1 lacks the 150-cavity in its active site. Nature Struct. Mol. Biol. 17, 1266–1268 (2010).
Bohne-Lang, A. & von der Lieth, C. W. GlyProt: in silico glycosylation of proteins. Nucleic Acids Res. 33, W214–W219 (2005).
Gamblin, S. et al. The structure and receptor binding properties of the 1918 influenza hemagglutinin. Science 303, 1838–1842 (2004).
Xu, X., Zhu, X., Dwek, R. A., Stevens, J. & Wilson, I. A. Structural characterization of the 1918 influenza virus H1N1 neuraminidase. J. Virol. 82, 10493–10501 (2008).
Acknowledgements
The authors thank T. J. Wohlbold for help with GlyProt and PyMOL. Research in the Krammer laboratory is supported by a US National Institutes of Health (NIH) Centres for Excellence in Influenza Research and Surveillance (CEIRS) contract (HHSN272201400008C). P.P. is supported by an NIH CEIRS contract (HHSN272201400008C) and by NIH grants (U19 AI109946 and P01 AI097092).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The Icahn School of Medicine at Mount Sinai has filed several patents regarding influenza virus vaccine constructs.
Related links
Glossary
- Antigenic drift
-
A mechanism by which influenza viruses escape from human 'herd immunity'. The RNA-dependent RNA polymerase of influenza viruses is relatively error prone and has no proofreading mechanism, resulting in a high frequency of point mutations. Immunologic pressure in the human population then selects for mutants that can escape from this herd immunity. The globular head domain of haemagglutinin is — owing to its immuno-dominance and high plasticity — most affected by antigenic drift.
- Haemagglutinin
-
(HA). A homotrimeric viral surface glycoprotein that mediates the attachment of influenza viruses to cells by binding to sialic acids on glycan structures of cellular receptors. Haemagglutinin also mediates the fusion of viral and endosomal membranes, which causes the release of the viral genome into the cytosol. Haemagglutinin is the major antigen of the virus.
- Neuraminidase
-
(NA). A viral homotetrameric viral surface glycoprotein with sialidase activity. Neuraminidase helps transport the virus trough mucosal surfaces and mediates the release of budding viruses from the cell surface.
- Whole-virus inactivated vaccines
-
Whole-virus inactivated vaccines are based on intact virions that have been chemically (for example, with formalin or β-propiolactone) or physically (for example, with ultraviolet light) inactivated. Treatment of these virions with detergent leads to split vaccines. Further (partial) purification of the haemagglutinin and neuraminidase of viruses results in subunit vaccines.
- Haemagglutination inhibition
-
(HI). Haemagglutination activity is the standard correlate of protection used for influenza virus vaccines, and haemagglutination inhibition describes the ability of antibodies to block the binding of the haemagglutinin globular head domain to cellular receptors. Stalk-reactive antibodies are generally haemagglutination inhibition negative.
- Neuraminidase inhibition
-
(NI). NI describes the ability of antibodies to block the sialidase function of neuraminidase.
Rights and permissions
About this article
Cite this article
Krammer, F., Palese, P. Advances in the development of influenza virus vaccines. Nat Rev Drug Discov 14, 167–182 (2015). https://doi.org/10.1038/nrd4529
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrd4529
This article is cited by
-
A structure and knowledge-based combinatorial approach to engineering universal scFv antibodies against influenza M2 protein
Journal of Biomedical Science (2023)
-
Potential cross-species transmission of highly pathogenic avian influenza H5 subtype (HPAI H5) viruses to humans calls for the development of H5-specific and universal influenza vaccines
Cell Discovery (2023)
-
Trends in nano-platforms for the treatment of viral infectious diseases
Korean Journal of Chemical Engineering (2023)
-
Enhanced passive safety surveillance of a quadrivalent inactivated split virion influenza vaccine in Finland during the influenza season 2020/21
BMC Public Health (2022)
-
Generation of a live attenuated influenza A vaccine by proteolysis targeting
Nature Biotechnology (2022)