Key Points
-
The ubiquitin–proteasome system (UPS) has links to numerous diseases, and the clinical success of proteasome inhibitors suggests the potential to develop drugs targeting other components of the UPS. In certain cases, the specific inhibition of individual components of the UPS may provide better therapeutic outcomes than does broad inhibition of the proteasome.
-
F-box proteins are the targeting subunits of the SKP1-CUL1-F-box protein (SCF) ubiquitin ligase complexes, the best characterized of the mammalian cullin–RING ligase family of E3 ubiquitin ligases. In mammals, there are approximately 70 F-box proteins, each thought to be able to target multiple substrates, facilitating the regulation of diverse cellular pathways.
-
SCF complexes are dysregulated in several diseases by overexpression of the F-box protein, by mutation (genetic deletion or point mutations) of the F-box protein, or by disrupting the pathways that control F-box protein–substrate targeting.
-
In cases in which an SCF complex is overactive, the inhibition of SCF-mediated degradation can be achieved via competitive inhibition between substrate and F-box protein, allosteric inhibition that disrupts the interaction between substrate and F-box protein, or via the inhibition of SCF complex assembly.
-
In cases in which SCF activity is defective in disease, SCF activity can by reconstituted through bivalent small molecules that re-target the substrate and/or 'molecular glue'-like molecules that restore substrate–F-box protein binding.
-
Although an underdeveloped area, SCF complexes are promising drug targets, and as the substrates and functions of orphan F-box proteins are discovered, further drug targets will become apparent. Several compounds targeting E3 ligases have been developed, hinting that effective drugs targeting other SCF ubiquitin ligase activities may also be developed.
Abstract
The clinical successes of proteasome inhibitors for the treatment of cancer have highlighted the therapeutic potential of targeting this protein degradation system. However, proteasome inhibitors prevent the degradation of numerous proteins, which may cause adverse effects. Increased specificity could be achieved by inhibiting the components of the ubiquitin–proteasome system that target specific subsets of proteins for degradation. F-box proteins are the substrate-targeting subunits of SKP1–CUL1–F-box protein (SCF) ubiquitin ligase complexes. Through the degradation of a plethora of diverse substrates, SCF ubiquitin ligases control a multitude of processes at the cellular and organismal levels, and their dysregulation is implicated in many pathologies. SCF ubiquitin ligases are characterized by their high specificity for substrates, and these ligases therefore represent promising drug targets. However, the potential for therapeutic manipulation of SCF complexes remains an underdeveloped area. This Review explores and discusses potential strategies to target SCF-mediated biological processes to treat human diseases.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Hershko, A. & Ciechanover, A. The ubiquitin system. Annu. Rev. Biochem. 67, 425–479 (1998).
Rieser, E., Cordier, S. M. & Walczak, H. Linear ubiquitination: a newly discovered regulator of cell signalling. Trends Biochem. Sci. 38, 94–102 (2013).
Komander, D. & Rape, M. The ubiquitin code. Annu. Rev. Biochem. 81, 203–229 (2012).
Nijman, S. M. et al. A genomic and functional inventory of deubiquitinating enzymes. Cell 123, 773–786 (2005).
Frescas, D. & Pagano, M. Deregulated proteolysis by the F-box proteins SKP2 and β-TrCP: tipping the scales of cancer. Nature Rev. Cancer 8, 438–449 (2008).
Skaar, J. R., Pagan, J. K. & Pagano, M. Mechanisms and function of substrate recruitment by F-box proteins. Nature Rev. Mol. Cell Biol. 14, 369–381 (2013).
Schmidt, M. & Finley, D. Regulation of proteasome activity in health and disease. Biochim. Biophys. Acta 1843, 13–25 (2014).
Suzuki, E. et al. Molecular mechanisms of bortezomib resistant adenocarcinoma cells. PLoS ONE 6, e27996 (2011).
Ruschak, A. M., Slassi, M., Kay, L. E. & Schimmer, A. D. Novel proteasome inhibitors to overcome bortezomib resistance. J. Natl Cancer Inst. 103, 1007–1017 (2011).
Kubiczkova, L., Pour, L., Sedlarikova, L., Hajek, R. & Sevcikova, S. Proteasome inhibitors — molecular basis and current perspectives in multiple myeloma. J. Cell. Mol. Med. 18, 947–961 (2014).
Mattern, M. R., Wu, J. & Nicholson, B. Ubiquitin-based anticancer therapy: carpet bombing with proteasome inhibitors versus surgical strikes with E1, E2, E3, or DUB inhibitors. Biochim. Biophys. Acta 1823, 2014–2021 (2012).
Schulman, B. A. & Harper, J. W. Ubiquitin-like protein activation by E1 enzymes: the apex for downstream signalling pathways. Nature Rev. Mol. Cell Biol. 10, 319–331 (2009).
Berndsen, C. E. & Wolberger, C. New insights into ubiquitin E3 ligase mechanism. Nature Struct. Mol. Biol. 21, 301–307 (2014).
Metzger, M. B., Hristova, V. A. & Weissman, A. M. HECT and RING finger families of E3 ubiquitin ligases at a glance. J. Cell Sci. 125, 531–537 (2012).
Deshaies, R. J. & Joazeiro, C. A. RING domain E3 ubiquitin ligases. Annu. Rev. Biochem. 78, 399–434 (2009).
Lydeard, J. R., Schulman, B. A. & Harper, J. W. Building and remodelling Cullin-RING E3 ubiquitin ligases. EMBO Rep. 14, 1050–1061 (2013).
Wei, N., Serino, G. & Deng, X. W. The COP9 signalosome: more than a protease. Trends Biochem. Sci. 33, 592–600 (2008).
Jin, J. et al. Systematic analysis and nomenclature of mammalian F-box proteins. Genes Dev. 18, 2573–2580 (2004).
Skaar, J. R., D'Angiolella, V., Pagan, J. K. & Pagano, M. SnapShot: F box proteins II. Cell 137, 1358.e1–1358.e2 (2009).
Sutterluty, H. et al. p45SKP2 promotes p27Kip1 degradation and induces S phase in quiescent cells. Nature Cell Biol. 1, 207–214 (1999).
Carrano, A. C., Eytan, E., Hershko, A. & Pagano, M. SKP2 is required for ubiquitin-mediated degradation of the CDK inhibitor p27. Nature Cell Biol. 1, 193–199 (1999).
Ganoth, D. et al. The cell-cycle regulatory protein Cks1 is required for SCF(Skp2)-mediated ubiquitinylation of p27. Nature Cell Biol. 3, 321–324 (2001).
Spruck, C. et al. A CDK-independent function of mammalian Cks1: targeting of SCF(Skp2) to the CDK inhibitor p27Kip1. Mol. Cell 7, 639–650 (2001).
Abbas, T. et al. CRL1-FBXO11 promotes Cdt2 ubiquitylation and degradation and regulates Pr-Set7/Set8-mediated cellular migration. Mol. Cell 49, 1147–1158 (2013).
Rossi, M. et al. Regulation of the CRL4(Cdt2) ubiquitin ligase and cell-cycle exit by the SCF(Fbxo11) ubiquitin ligase. Mol. Cell 49, 1159–1166 (2013).
Kuchay, S. et al. FBXL2- and PTPL1-mediated degradation of p110-free p85β regulatory subunit controls the PI(3)K signalling cascade. Nature Cell Biol. 15, 472–480 (2013).
Pitts, T. M., Davis, S. L., Eckhardt, S. G. & Bradshaw-Pierce, E. L. Targeting nuclear kinases in cancer: development of cell cycle kinase inhibitors. Pharmacol. Ther. 142, 258–269 (2014).
Zhang, J., Yang, P. L. & Gray, N. S. Targeting cancer with small molecule kinase inhibitors. Nature Rev. Cancer 9, 28–39 (2009).
Davies, S. P., Reddy, H., Caivano, M. & Cohen, P. Specificity and mechanism of action of some commonly used protein kinase inhibitors. Biochem. J. 351, 95–105 (2000).
Wilhelm, S. et al. Discovery and development of sorafenib: a multikinase inhibitor for treating cancer. Nature Rev. Drug Discov. 5, 835–844 (2006).
Cardozo, T. & Pagano, M. The SCF ubiquitin ligase: insights into a molecular machine. Nature Rev. Mol. Cell Biol. 5, 739–751 (2004).
Vassilev, L. T. MDM2 inhibitors for cancer therapy. Trends Mol. Med. 13, 23–31 (2007).
Vassilev, L. T. et al. In vivo activation of the p53 pathway by small-molecule antagonists of MDM2. Science 303, 844–848 (2004).
Van Maerken, T. et al. Pharmacologic activation of wild-type p53 by nutlin therapy in childhood cancer. Cancer Lett. 344, 157–165 (2014).
Soucy, T. A. et al. An inhibitor of NEDD8-activating enzyme as a new approach to treat cancer. Nature 458, 732–736 (2009).
Li, L. et al. Overactivated neddylation pathway as a therapeutic target in lung cancer. J. Natl Cancer Inst. 106, dju083 (2014).
Li, H. et al. Inactivation of SAG/RBX2 E3 ubiquitin ligase suppresses KrasG12D-driven lung tumorigenesis. J. Clin. Invest. 124, 835–846 (2014).
Gu, Y. et al. MLN4924, an NAE inhibitor, suppresses AKT and mTOR signaling via upregulation of REDD1 in human myeloma cells. Blood 123, 3269–3276 (2014).
Milhollen, M. A. et al. MLN4924, a NEDD8-activating enzyme inhibitor, is active in diffuse large B-cell lymphoma models: rationale for treatment of NF-κB-dependent lymphoma. Blood 116, 1515–1523 (2010).
Pan, W. W. et al. Ubiquitin E3 ligase CRL4(CDT2/DCAF2) as a potential chemotherapeutic target for ovarian surface epithelial cancer. J. Biol. Chem. 288, 29680–29691 (2013).
Mackintosh, C. et al. WEE1 accumulation and deregulation of S-phase proteins mediate MLN4924 potent inhibitory effect on Ewing sarcoma cells. Oncogene 32, 1441–1451 (2013).
Wei, D. et al. Radiosensitization of human pancreatic cancer cells by MLN4924, an investigational NEDD8-activating enzyme inhibitor. Cancer Res. 72, 282–293 (2012).
Smith, M. A. et al. Initial testing of the investigational NEDD8-activating enzyme inhibitor MLN4924 by the pediatric preclinical testing program. Pediatr. Blood Cancer 59, 246–253 (2012).
Xu, G. W. et al. Mutations in UBA3 confer resistance to the NEDD8-activating enzyme inhibitor MLN4924 in human leukemic cells. PLoS ONE 9, e93530 (2014).
Blank, J. L. et al. Novel DNA damage checkpoints mediating cell death induced by the NEDD8-activating enzyme inhibitor MLN4924. Cancer Res. 73, 225–234 (2013).
Lin, J. J., Milhollen, M. A., Smith, P. G., Narayanan, U. & Dutta, A. NEDD8-targeting drug MLN4924 elicits DNA rereplication by stabilizing Cdt1 in S phase, triggering checkpoint activation, apoptosis, and senescence in cancer cells. Cancer Res. 70, 10310–10320 (2010).
Soucy, T. A., Dick, L. R., Smith, P. G., Milhollen, M. A. & Brownell, J. E. The NEDD8 conjugation pathway and its relevance in cancer biology and therapy. Genes Cancer 1, 708–716 (2010).
Wiley, D. J. et al. Yeast Augmented Network Analysis (YANA): a new systems approach to identify therapeutic targets for human genetic diseases. F1000Res 3, 121 (2014).
Watson, I. R., Irwin, M. S. & Ohh, M. NEDD8 pathways in cancer, Sine Quibus Non. Cancer Cell 19, 168–176 (2011).
Toth, J. I., Yang, L., Dahl, R. & Petroski, M. D. A gatekeeper residue for NEDD8-activating enzyme inhibition by MLN4924. Cell Rep. 1, 309–316 (2012).
Milhollen, M. A. et al. Treatment-emergent mutations in NAEbeta confer resistance to the NEDD8-activating enzyme inhibitor MLN4924. Cancer Cell 21, 388–401 (2012).
Wang, C. et al. Identification of FBL2 as a geranylgeranylated cellular protein required for hepatitis C virus RNA replication. Mol. Cell 18, 425–434 (2005).
Sandberg, J. K., Andersson, S. K., Bachle, S. M., Nixon, D. F. & Moll, M. HIV-1 Vpu interference with innate cell-mediated immune mechanisms. Curr. HIV Res. 10, 327–333 (2012).
Zhang, E. E. et al. Cryptochrome mediates circadian regulation of cAMP signaling and hepatic gluconeogenesis. Nature Med. 16, 1152–1156 (2010).
Price, C. T. et al. Molecular mimicry by an F-box effector of Legionella pneumophila hijacks a conserved polyubiquitination machinery within macrophages and protozoa. PLoS Pathog. 5, e1000704 (2009).
Zhang, L., Villa, N. Y. & McFadden, G. Interplay between poxviruses and the cellular ubiquitin/ubiquitin-like pathways. FEBS Lett. 583, 607–614 (2009).
Dupuis, J. et al. New genetic loci implicated in fasting glucose homeostasis and their impact on type 2 diabetes risk. Nature Genet. 42, 105–116 (2010).
Di Fonzo, A. et al. FBXO7 mutations cause autosomal recessive, early-onset parkinsonian-pyramidal syndrome. Neurology 72, 240–245 (2009).
Toh, K. L. et al. An hPer2 phosphorylation site mutation in familial advanced sleep phase syndrome. Science 291, 1040–1043 (2001).
Welcker, M. & Clurman, B. E. FBW7 ubiquitin ligase: a tumour suppressor at the crossroads of cell division, growth and differentiation. Nature Rev. Cancer 8, 83–93 (2008).
Wang, Z., Liu, P., Inuzuka, H. & Wei, W. Roles of F-box proteins in cancer. Nature Rev. Cancer 14, 233–247 (2014).
Kossatz, U. et al. Skp2-dependent degradation of p27kip1 is essential for cell cycle progression. Genes Dev. 18, 2602–2607 (2004).
Nakayama, K. et al. Targeted disruption of Skp2 results in accumulation of cyclin E and p27(Kip1), polyploidy and centrosome overduplication. EMBO J. 19, 2069–2081 (2000).
Nakayama, K. et al. Skp2-mediated degradation of p27 regulates progression into mitosis. Dev. Cell 6, 661–672 (2004).
Tsvetkov, L. M., Yeh, K. H. & Lee, S. J., Sun, H. & Zhang, H. p27Kip1 ubiquitination and degradation is regulated by the SCFSkp2 complex through phosphorylated Thr187 in p27. Curr. Biol. 9, 661–664 (1999).
Hao, B. et al. Structural basis of the Cks1-dependent recognition of p27Kip1 by the SCFSkp2 ubiquitin ligase. Mol. Cell 20, 9–19 (2005).
Wu, L. et al. Specific small molecule inhibitors of Skp2-mediated p27 degradation. Chem. Biol. 19, 1515–1524 (2012).
Pavlides, S. C. et al. Inhibitors of SCF-Skp2/Cks1 E3 ligase block estrogen-induced growth stimulation and degradation of nuclear p27kip1: therapeutic potential for endometrial cancer. Endocrinology 154, 4030–4045 (2013).
Chan, C. H. et al. Pharmacological inactivation of Skp2 SCF ubiquitin ligase restricts cancer stem cell traits and cancer progression. Cell 154, 556–568 (2013).
Aghajan, M. et al. Chemical genetics screen for enhancers of rapamycin identifies a specific inhibitor of an SCF family E3 ubiquitin ligase. Nature Biotech. 28, 738–742 (2010).
Orlicky, S. et al. An allosteric inhibitor of substrate recognition by the SCFCdc4 ubiquitin ligase. Nature Biotech. 28, 733–737 (2010).
Chen, Q. et al. Targeting the p27 E3 ligase SCFSkp2 results in p27- and Skp2-mediated cell-cycle arrest and activation of autophagy. Blood 111, 4690–4699 (2008).
Huang, K. S. & Vassilev, L. T. High-throughput screening for inhibitors of the Cks1-Skp2 interaction. Methods Enzymol. 399, 717–728 (2005).
Rico-Bautista, E., Yang, C. C., Lu, L., Roth, G. P. & Wolf, D. A. Chemical genetics approach to restoring p27Kip1 reveals novel compounds with antiproliferative activity in prostate cancer cells. BMC Biol. 8, 153 (2010).
Zhang, G. J. et al. Bioluminescent imaging of Cdk2 inhibition in vivo. Nature Med. 10, 643–648 (2004).
Busino, L. et al. Fbxw7α- and GSK3-mediated degradation of p100 is a pro-survival mechanism in multiple myeloma. Nature Cell Biol. 14, 375–385 (2012).
Busino, L. et al. SCFFbxl3 controls the oscillation of the circadian clock by directing the degradation of cryptochrome proteins. Science 316, 900–904 (2007).
Godinho, S. I. et al. The after-hours mutant reveals a role for Fbxl3 in determining mammalian circadian period. Science 316, 897–900 (2007).
Siepka, S. M. et al. Circadian mutant Overtime reveals F-box protein FBXL3 regulation of cryptochrome and period gene expression. Cell 129, 1011–1023 (2007).
Xing, W. et al. SCFFBXL3 ubiquitin ligase targets cryptochromes at their cofactor pocket. Nature 496, 64–68 (2013).
Hirota, T. et al. Identification of small molecule activators of cryptochrome. Science 337, 1094–1097 (2012).
Nangle, S., Xing, W. & Zheng, N. Crystal structure of mammalian cryptochrome in complex with a small molecule competitor of its ubiquitin ligase. Cell Res. 23, 1417–1419 (2013).
Ye, J. et al. Disruption of hepatitis C virus RNA replication through inhibition of host protein geranylgeranylation. Proc. Natl Acad. Sci. USA 100, 15865–15870 (2003).
Feeney, E. R. & Chung, R. T. Antiviral treatment of hepatitis C. BMJ 349, g3308 (2014).
Pawlotsky, J. M. New hepatitis C therapies: the toolbox, strategies, and challenges. Gastroenterology 146, 1176–1192 (2014).
Chen, B. B., Coon, T. A., Glasser, J. R. & Mallampalli, R. K. Calmodulin antagonizes a calcium-activated SCF ubiquitin E3 ligase subunit, FBXL2, to regulate surfactant homeostasis. Mol. Cell. Biol. 31, 1905–1920 (2011).
Chen, B. B. et al. A combinatorial F box protein directed pathway controls TRAF adaptor stability to regulate inflammation. Nature Immunol. 14, 470–479 (2013).
Chen, B. B., Glasser, J. R., Coon, T. A. & Mallampalli, R. K. Skp-cullin-F box E3 ligase component FBXL2 ubiquitinates Aurora B to inhibit tumorigenesis. Cell Death Dis. 4, e759 (2013).
Chen, B. B., Glasser, J. R., Coon, T. A. & Mallampalli, R. K. F-box protein FBXL2 exerts human lung tumor suppressor-like activity by ubiquitin-mediated degradation of cyclin D3 resulting in cell cycle arrest. Oncogene 31, 2566–2579 (2012).
Chen, B. B. et al. F-box protein FBXL2 targets cyclin D2 for ubiquitination and degradation to inhibit leukemic cell proliferation. Blood 119, 3132–3141 (2012).
Matsuoka, S. et al. Fbxw7 acts as a critical fail-safe against premature loss of hematopoietic stem cells and development of T-ALL. Genes Dev. 22, 986–991 (2008).
Onoyama, I. et al. Conditional inactivation of Fbxw7 impairs cell-cycle exit during T cell differentiation and results in lymphomatogenesis. J. Exp. Med. 204, 2875–2888 (2007).
Thompson, B. J. et al. Control of hematopoietic stem cell quiescence by the E3 ubiquitin ligase Fbw7. J. Exp. Med. 205, 1395–1408 (2008).
Crusio, K. M., King, B., Reavie, L. B. & Aifantis, I. The ubiquitous nature of cancer: the role of the SCFFbw7 complex in development and transformation. Oncogene 29, 4865–4873 (2010).
Akhoondi, S. et al. FBXW7/hCDC4 is a general tumor suppressor in human cancer. Cancer Res. 67, 9006–9012 (2007).
Maser, R. S. et al. Chromosomally unstable mouse tumours have genomic alterations similar to diverse human cancers. Nature 447, 966–971 (2007).
Lawrence, M. S. et al. Discovery and saturation analysis of cancer genes across 21 tumour types. Nature 505, 495–501 (2014).
Annunziata, C. M. et al. Frequent engagement of the classical and alternative NF-κB pathways by diverse genetic abnormalities in multiple myeloma. Cancer Cell 12, 115–130 (2007).
Gregory, M. A. & Hann, S. R. c-Myc proteolysis by the ubiquitin-proteasome pathway: stabilization of c-Myc in Burkitt's lymphoma cells. Mol. Cell. Biol. 20, 2423–2435 (2000).
Bahram, F., von der Lehr, N., Cetinkaya, C. & Larsson, L. G. c-Myc hot spot mutations in lymphomas result in inefficient ubiquitination and decreased proteasome-mediated turnover. Blood 95, 2104–2110 (2000).
Weng, A. P. et al. Activating mutations of NOTCH1 in human T cell acute lymphoblastic leukemia. Science 306, 269–271 (2004).
Sakamoto, K. M. et al. Protacs: chimeric molecules that target proteins to the Skp1-Cullin-F box complex for ubiquitination and degradation. Proc. Natl Acad. Sci. USA 98, 8554–8559 (2001).
Sakamoto, K. M. et al. Development of Protacs to target cancer-promoting proteins for ubiquitination and degradation. Mol. Cell Proteom. 2, 1350–1358 (2003).
Bouchie, A., Allison, M., Webb, S. & Defrancesco, L. Nature Biotechnology's academic spinouts of 2013. Nature Biotech. 32, 229–238 (2014).
Schneekloth, A. R., Pucheault, M., Tae, H. S. & Crews, C. M. Targeted intracellular protein degradation induced by a small molecule: en route to chemical proteomics. Bioorg. Med. Chem. Lett. 18, 5904–5908 (2008).
Kronke, J. et al. Lenalidomide causes selective degradation of IKZF1 and IKZF3 in multiple myeloma cells. Science 343, 301–305 (2014).
Lu, G. et al. The myeloma drug lenalidomide promotes the cereblon-dependent destruction of Ikaros proteins. Science 343, 305–309 (2014).
Fischer, E. S. et al. Structure of the DDB1–CRBN E3 ubiquitin ligase in complex with thalidomide. Nature 512, 49–53 (2014).
Ito, T., Ando, H. & Handa, H. Teratogenic effects of thalidomide: molecular mechanisms. Cell. Mol. Life Sci. 68, 1569–1579 (2011).
Ito, T. et al. Identification of a primary target of thalidomide teratogenicity. Science 327, 1345–1350 (2010).
Tan, X. et al. Mechanism of auxin perception by the TIR1 ubiquitin ligase. Nature 446, 640–645 (2007).
Nishimura, K., Fukagawa, T., Takisawa, H., Kakimoto, T. & Kanemaki, M. An auxin-based degron system for the rapid depletion of proteins in nonplant cells. Nature Methods 6, 917–922 (2009).
Basse, N. et al. Toward the rational design of p53-stabilizing drugs: probing the surface of the oncogenic Y220C mutant. Chem. Biol. 17, 46–56 (2010).
Boeckler, F. M. et al. Targeted rescue of a destabilized mutant of p53 by an in silico screened drug. Proc. Natl Acad. Sci. USA 105, 10360–10365 (2008).
Duan, S. et al. FBXO11 targets BCL6 for degradation and is inactivated in diffuse large B-cell lymphomas. Nature 481, 90–93 (2012).
Lohr, J. G. et al. Discovery and prioritization of somatic mutations in diffuse large B-cell lymphoma (DLBCL) by whole-exome sequencing. Proc. Natl Acad. Sci. USA 109, 3879–3884 (2012).
Yoshida, K. et al. Frequent pathway mutations of splicing machinery in myelodysplasia. Nature 478, 64–69 (2011).
Stransky, N. et al. The mutational landscape of head and neck squamous cell carcinoma. Science 333, 1157–1160 (2011).
Kan, Z. et al. Diverse somatic mutation patterns and pathway alterations in human cancers. Nature 466, 869–873 (2010).
Richter, J. et al. Recurrent mutation of the ID3 gene in Burkitt lymphoma identified by integrated genome, exome and transcriptome sequencing. Nature Genet. 44, 1316–1320 (2012).
Baraniskin, A. et al. MiR-30a-5p suppresses tumor growth in colon carcinoma by targeting DTL. Carcinogenesis 33, 732–739 (2012).
Pan, H. W. et al. Role of L2DTL, cell cycle-regulated nuclear and centrosome protein, in aggressive hepatocellular carcinoma. Cell Cycle 5, 2676–2687 (2006).
Ueki, T. et al. Involvement of elevated expression of multiple cell-cycle regulator, DTL/RAMP (denticleless/RA-regulated nuclear matrix associated protein), in the growth of breast cancer cells. Oncogene 27, 5672–5683 (2008).
Mackintosh, C. et al. 1q gain and CDT2 overexpression underlie an aggressive and highly proliferative form of Ewing sarcoma. Oncogene 31, 1287–1298 (2012).
Li, J. et al. Identification of retinoic acid-regulated nuclear matrix-associated protein as a novel regulator of gastric cancer. Br. J. Cancer 101, 691–698 (2009).
Mir, A. et al. Truncation of the E3 ubiquitin ligase component FBXO31 causes non-syndromic autosomal recessive intellectual disability in a Pakistani family. Hum. Genet. 133, 975–984 (2014).
Seeley, A. H. et al. Macrocerebellum, epilepsy, intellectual disability, and gut malrotation in a child with a 16q24.1-q24.2 contiguous gene deletion. Am. J. Med. Genet. A 164, 2062–2068 (2014).
Bonnen, P. E. et al. Mutations in FBXL4 cause mitochondrial encephalopathy and a disorder of mitochondrial DNA maintenance. Am. J. Hum. Genet. 93, 471–481 (2013).
Gai, X. et al. Mutations in FBXL4, encoding a mitochondrial protein, cause early-onset mitochondrial encephalomyopathy. Am. J. Hum. Genet. 93, 482–495 (2013).
Staudt, L. M. Oncogenic activation of NF-κB. Cold Spring Harb. Perspect. Biol. 2, a000109 (2010).
Yaron, A. et al. Identification of the receptor component of the IκBα-ubiquitin ligase. Nature 396, 590–594 (1998).
Dehan, E. et al. βTrCP- and Rsk1/2-mediated degradation of BimEL inhibits apoptosis. Mol. Cell 33, 109–116 (2009).
Soldatenkov, V. A., Dritschilo, A., Ronai, Z. & Fuchs, S. Y. Inhibition of homologue of Slimb (HOS) function sensitizes human melanoma cells for apoptosis. Cancer Res. 59, 5085–5088 (1999).
Takeishi, S. & Nakayama, K. I. Role of Fbxw7 in the maintenance of normal stem cells and cancer-initiating cells. Br. J. Cancer 111, 1054–1059 (2014).
Takeishi, S. et al. Ablation of Fbxw7 eliminates leukemia-initiating cells by preventing quiescence. Cancer Cell 23, 347–361 (2013).
King, B. et al. The ubiquitin ligase FBXW7 modulates leukemia-initiating cell activity by regulating MYC stability. Cell 153, 1552–1566 (2013).
Reavie, L. et al. Regulation of c-Myc ubiquitination controls chronic myelogenous leukemia initiation and progression. Cancer Cell 23, 362–375 (2013).
Navon, A. & Ciechanover, A. The 26 S proteasome: from basic mechanisms to drug targeting. J. Biol. Chem. 284, 33713–33718 (2009).
Buckley, D. L. & Crews, C. M. Small-molecule control of intracellular protein levels through modulation of the ubiquitin proteasome system. Angew. Chem. Int. Ed Engl. 53, 2312–2330 (2014).
Ye, Y. Diverse functions with a common regulator: ubiquitin takes command of an AAA ATPase. J. Struct. Biol. 156, 29–40 (2006).
Fessart, D., Marza, E., Taouji, S., Delom, F. & Chevet, E. P97/CDC-48: proteostasis control in tumor cell biology. Cancer Lett. 337, 26–34 (2013).
Yen, J. L. et al. Signal-induced disassembly of the SCF ubiquitin ligase complex by Cdc48/p97. Mol. Cell 48, 288–297 (2012).
den Besten, W., Verma, R., Kleiger, G., Oania, R. S. & Deshaies, R. J. NEDD8 links cullin-RING ubiquitin ligase function to the p97 pathway. Nature Struct. Mol. Biol. 19, 511–516 (2012).
Chou, T. F., Li, K., Frankowski, K. J., Schoenen, F. J. & Deshaies, R. J. Structure-activity relationship study reveals ML240 and ML241 as potent and selective inhibitors of p97 ATPase. ChemMedChem. 8, 297–312 (2013).
Magnaghi, P. et al. Covalent and allosteric inhibitors of the ATPase VCP/p97 induce cancer cell death. Nature Chem. Biol. 9, 548–556 (2013).
Groettrup, M., Pelzer, C., Schmidtke, G. & Hofmann, K. Activating the ubiquitin family: UBA6 challenges the field. Trends Biochem. Sci. 33, 230–237 (2008).
Yang, Y. et al. Inhibitors of ubiquitin-activating enzyme (E1), a new class of potential cancer therapeutics. Cancer Res. 67, 9472–9481 (2007).
Chen, J. J. et al. Mechanistic studies of substrate-assisted inhibition of ubiquitin-activating enzyme by adenosine sulfamate analogues. J. Biol. Chem. 286, 40867–40877 (2011).
Ceccarelli, D. F. et al. An allosteric inhibitor of the human Cdc34 ubiquitin-conjugating enzyme. Cell 145, 1075–1087 (2011).
Huang, H. et al. E2 enzyme inhibition by stabilization of a low-affinity interface with ubiquitin. Nature Chem. Biol. 10, 156–163 (2014).
Nicholson, B. & Suresh Kumar, K. G. The multifaceted roles of USP7: new therapeutic opportunities. Cell Biochem. Biophys. 60, 61–68 (2011).
Colland, F. et al. Small-molecule inhibitor of USP7/HAUSP ubiquitin protease stabilizes and activates p53 in cells. Mol. Cancer Ther. 8, 2286–2295 (2009).
Lee, B. H., Finley, D. & King, R. W. A. High-throughput screening method for identification of inhibitors of the deubiquitinating enzyme USP14. Curr. Protoc. Chem. Biol. 4, 311–330 (2012).
Lee, B. H. et al. Enhancement of proteasome activity by a small-molecule inhibitor of USP14. Nature 467, 179–184 (2010).
Tian, Z. et al. A novel small molecule inhibitor of deubiquitylating enzyme USP14 and UCHL5 induces apoptosis in multiple myeloma and overcomes bortezomib resistance. Blood 123, 706–716 (2014).
Chauhan, D. et al. A small molecule inhibitor of ubiquitin-specific protease-7 induces apoptosis in multiple myeloma cells and overcomes bortezomib resistance. Cancer Cell 22, 345–358 (2012).
Weinstock, J. et al. Selective dual inhibitors of the cancer-related deubiquitylating proteases USP7 and USP47. ACS Med. Chem. Lett. 3, 789–792 (2012).
Kapuria, V. et al. Deubiquitinase inhibition by small-molecule WP1130 triggers aggresome formation and tumor cell apoptosis. Cancer Res. 70, 9265–9276 (2010).
Chen, J. et al. Selective and cell-active inhibitors of the USP1/ UAF1 deubiquitinase complex reverse cisplatin resistance in non-small cell lung cancer cells. Chem. Biol. 18, 1390–1400 (2011).
Faesen, A. C. et al. Mechanism of USP7/HAUSP activation by its C-terminal ubiquitin-like domain and allosteric regulation by GMP-synthetase. Mol. Cell 44, 147–159 (2011).
Cohn, M. A. et al. A UAF1-containing multisubunit protein complex regulates the Fanconi anemia pathway. Mol. Cell 28, 786–797 (2007).
Villamil, M. A. et al. Serine phosphorylation is critical for the activation of ubiquitin-specific protease 1 and its interaction with WD40-repeat protein UAF1. Biochemistry 51, 9112–9123 (2012).
Villamil, M. A., Chen, J., Liang, Q. & Zhuang, Z. A noncanonical cysteine protease USP1 is activated through active site modulation by USP1-associated factor 1. Biochemistry 51, 2829–2839 (2012).
Cohen, P. & Tcherpakov, M. Will the ubiquitin system furnish as many drug targets as protein kinases? Cell 143, 686–693 (2010).
Knight, Z. A., Lin, H. & Shokat, K. M. Targeting the cancer kinome through polypharmacology. Nature Rev. Cancer 10, 130–137 (2010).
Schwartz, G. K. et al. Phase I study of PD 0332991, a cyclin-dependent kinase inhibitor, administered in 3-week cycles (Schedule 2/1). Br. J. Cancer 104, 1862–1868 (2011).
Rocca, A., Farolfi, A., Bravaccini, S., Schirone, A. & Amadori, D. Palbociclib (PD 0332991): targeting the cell cycle machinery in breast cancer. Expert Opin. Pharmacother. 15, 407–420 (2014).
Fry, D. W. et al. Specific inhibition of cyclin-dependent kinase 4/6 by PD 0332991 and associated antitumor activity in human tumor xenografts. Mol. Cancer Ther. 3, 1427–1438 (2004).
Macias-Perez, I. M. & Flinn, I. W. GS-1101: a delta-specific PI3K inhibitor in chronic lymphocytic leukemia. Curr. Hematol. Malig Rep. 8, 22–27 (2013).
Fruman, D. A. & Rommel, C. PI3Kδ inhibitors in cancer: rationale and serendipity merge in the clinic. Cancer Discov. 1, 562–572 (2011).
Blagosklonny, M. V. Flavopiridol, an inhibitor of transcription: implications, problems and solutions. Cell Cycle 3, 1537–1542 (2004).
Mallampalli, R. K. et al. Targeting F box protein Fbxo3 to control cytokine-driven inflammation. J. Immunol. 191, 5247–5255 (2013).
Nakajima, H., Fujiwara, H., Furuichi, Y., Tanaka, K. & Shimbara, N. A novel small-molecule inhibitor of NF-κB signaling. Biochem. Biophys. Res. Commun. 368, 1007–1013 (2008).
Blees, J. S. et al. Erioflorin stabilizes the tumor suppressor Pdcd4 by inhibiting its interaction with the E3-ligase β-TrCP1. PLoS ONE 7, e46567 (2012).
Acknowledgements
The authors thank T. Cardozo, C. Crewes, R. Deshaies, R. Kumar, R. Mallampalli, K. Nakayama, N. Wilkie and N. Zheng for critically reading the manuscript and apologise for the omission of many colleagues' work owing to space constraints. J.R.S. is a Special Fellow of The Leukemia & Lymphoma Society. J.K.P. is supported by a fellowship from the Lymphoma Research Foundation. Work in the Pagano laboratory is supported by grants (R37CA076584 and R01GM057587) from the US National Institutes of Health. M.P. is an Investigator of the Howard Hughes Medical Institute.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Related links
FURTHER INFORMATION
Glossary
- Ubiquitin
-
A 76-amino acid protein that can be covalently conjugated to other proteins through amine groups at the amino terminus or on internal lysine residues. Proteins can be monoubiquitylated or polyubiquitylated. Chains assembled via K48 (in most cases) and K11 linkages target substrate proteins to the proteasome for degradation.
- Ubiquitylation
-
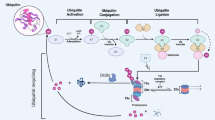
The process of covalently attaching ubiquitin to another protein. Ubiquitylation is accomplished via an enzymatic cascade of an E1 ubiquitin activating enzyme, an E2 ubiquitin conjugating enzyme, and an E3 ubiquitin ligase.
- Proteasome
-
A large protein complex that catalyses the proteolytic destruction of proteins.
- Ubiquitin–proteasome system
-
The ubiquitin–proteasome system (UPS) collectively refers to the proteins regulating ubiquitin-dependent protein degradation by the proteasome.
- RING domains
-
Really interesting new gene (RING) domains coordinate zinc ions and often mediate binding of E2 ubiquitin conjugating enzymes.
- HECT domain
-
Homologous to E6 carboxy-terminus (HECT) domains define a sub-family of E3 ubiquitin ligases. HECT domains recruit the E2 ubiquitin conjugating enzyme and participate directly in substrate ubiquitylation by forming a thioester bond with ubiquitin.
- Cullin (CUL)–RING-ligases
-
(CRLs). A family of multisubunit E3 ubiquitin ligases that share a common architecture. The eight CRLs are each based on a different cullin scaffold protein, and each complex recruits a small RING protein and a unique set of substrate adaptor proteins.
- SCF complexes
-
(SKP1–CUL1–F-box protein complexes; also known as CRL1). A family of multi-subunit E3 ubiquitin ligases. As listed in the name, SCF complexes contain a cullin 1 (CUL1) scaffold, and an S-phase kinase-associated protein 1 (SKP1) that bridges CUL1 to a variable F-box- containing protein, of which there are ~70 in mammals. These complexes also contain the small RING protein RBX1.
- NEDD8
-
A small, ubiquitin-like protein that can be conjugated to other proteins to alter their function. The best-defined and most abundant substrates for neddylation are the cullin family of proteins, which require neddylation for their activity.
- Degrons
-
A small region of a substrate protein that is recognized by the E3 ubiquitin ligase and is required for substrate binding and ubiquitylation. Minimal degrons are often short peptide sequences. A variety of mechanisms, including post-translational modification of residues within the degron, can regulate access and binding of the degron to the E3 ubiquitin ligase.
- Pharmacophores
-
The features of a small-molecule ligand that dictate the interactions with macromolecular targets that are responsible for its biological activities.
- Cryptochrome proteins
-
The two mammalian cryptochrome proteins (cryptochrome 1 (CRY1) and CRY2) function in the circadian clock machinery as transcriptional repressors.
- Hepatitis C virus (HCV) NS5A
-
A multifunctional protein generated from the cleavage of a polyprotein precursor during HCV infection and required for productive HCV infection.
- Proteolysis targeting chimeras
-
(PROTACs). Bi-functional compounds, molecules or proteins that bind an E3 ubiquitin ligase on one end and another protein that is not the normal E3 substrate on the other end. PROTACs retarget the E3 ligase activity to new substrates.
Rights and permissions
About this article
Cite this article
Skaar, J., Pagan, J. & Pagano, M. SCF ubiquitin ligase-targeted therapies. Nat Rev Drug Discov 13, 889–903 (2014). https://doi.org/10.1038/nrd4432
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrd4432
This article is cited by
-
FBXO22 promotes leukemogenesis by targeting BACH1 in MLL-rearranged acute myeloid leukemia
Journal of Hematology & Oncology (2023)
-
NLRP6 potentiates PI3K/AKT signalling by promoting autophagic degradation of p85α to drive tumorigenesis
Nature Communications (2023)
-
FBXW2 suppresses breast tumorigenesis by targeting AKT-Moesin-SKP2 axis
Cell Death & Disease (2023)
-
Functional characterization of FBXL7 as a novel player in human cancers
Cell Death Discovery (2022)
-
FBXO2 targets glycosylated SUN2 for ubiquitination and degradation to promote ovarian cancer development
Cell Death & Disease (2022)