Key Points
-
All RNA-targeted therapeutic technologies exploit oligonucleotides that bind to target RNA, but they differ in their mechanism of action and produce different effects.
-
Small interfering RNAs, antisense oligonucleotides and external guide sequences lead to enzyme-dependent degradation of targeted mRNA. Drugs involving these approaches are designed to reduce the level of harmful gene products such as viral or bacterial proteins or disease-promoting cellular proteins. They could be useful against cancer as well as viral and bacterial infections, or used to prevent the accumulation of high levels of cholesterol in the bloodstream.
-
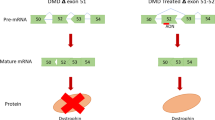
Steric-blocking oligonucleotides block the access of cellular machinery to pre-mRNA and mRNA without degrading RNA. Splice-switching oligonucleotides are discussed in detail in this Review; these oligonucleotides redirect alternative splicing, repair defective RNA or restore the production of proteins that are missing because of genetic defects. Splice-switching oligonucleotide-based drugs should be useful for the treatment of genetic diseases such as Duchenne muscular dystrophy, spinal muscular atrophy and β-thalassaemia.
-
Compared to classical small-molecule drugs, it is much more difficult to achieve intracellular delivery with oligonucleotides; this is still a major issue for this class of drugs. The advantage of oligonucleotides is their high specificity, which results from sequence-specific base pairing to target RNA.
-
The oligonucleotide-based drug fomivirsen was approved by the US Food and Drug Administration in 1998 for the treatment of viral retinitis in patients with AIDS. Oligonucleotide-based drugs are now in advanced clinical trials for the treatment of cancer and Duchenne muscular dystrophy as well as for lowering high cholesterol levels.
Abstract
Here, we discuss three RNA-based therapeutic technologies exploiting various oligonucleotides that bind to RNA by base pairing in a sequence-specific manner yet have different mechanisms of action and effects. RNA interference and antisense oligonucleotides downregulate gene expression by inducing enzyme-dependent degradation of targeted mRNA. Steric-blocking oligonucleotides block the access of cellular machinery to pre-mRNA and mRNA without degrading the RNA. Through this mechanism, steric-blocking oligonucleotides can redirect alternative splicing, repair defective RNA, restore protein production or downregulate gene expression. Moreover, they can be extensively chemically modified to acquire more drug-like properties. The ability of RNA-blocking oligonucleotides to restore gene function makes them best suited for the treatment of genetic disorders. Positive results from clinical trials for the treatment of Duchenne muscular dystrophy show that this technology is close to achieving its clinical potential.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Davidson, B. L. & McCray, P. B. Jr. Current prospects for RNA interference-based therapies. Nature Rev. Genet. 12, 329–340 (2011).
Goodchild, J. Therapeutic oligonucleotides. Methods Mol. Biol. 764, 1–15 (2011).
Crooke, S. T. Vitravene — another piece in the mosaic. Antisense Nucleic Acid Drug Dev. 8, vii–viii (1998). This is a paper on the first — and so far the only — antisense oligonucleotide drug approved for use as a treatment for cytomegalovirus-induced retinitis in patients with advanced AIDS.
Moshfeghi, A. A. & Puliafito, C. A. Pegaptanib sodium for the treatment of neovascular age-related macular degeneration. Expert Opin. Investig. Drugs 14, 671–682 (2005).
Fire, A. et al. Potent and specific genetic interference by double-stranded RNA in Caenorhabditis elegans. Nature 391, 806–811 (1998).
de Fougerolles, A., Vornlocher, H. P., Maraganore, J. & Lieberman, J. Interfering with disease: a progress report on siRNA-based therapeutics. Nature Rev. Drug Discov. 6, 443–453 (2007).
Elbashir, S. M. et al. Duplexes of 21-nucleotide RNAs mediate RNA interference in cultured mammalian cells. Nature 411, 494–498 (2001). This was the first demonstration that short double-stranded RNAs, now known as siRNAs can induce mRNA degradation via the RNAi pathway in human cells.
Song, E. et al. RNA interference targeting Fas protects mice from fulminant hepatitis. Nature Med. 9, 347–351 (2003).
Watts, J. K., Deleavey, G. F. & Damha, M. J. Chemically modified siRNA: tools and applications. Drug Discov. Today 13, 842–855 (2008).
DeVincenzo, J. et al. Evaluation of the safety, tolerability and pharmacokinetics of ALN-RSV01, a novel RNAi antiviral therapeutic directed against respiratory syncytial virus (RSV). Antiviral Res. 77, 225–231 (2008).
Semple, S. C. et al. Rational design of cationic lipids for siRNA delivery. Nature Biotech. 28, 172–176 (2010).
Jackson, A. L. & Linsley, P. S. Recognizing and avoiding siRNA off-target effects for target identification and therapeutic application. Nature Rev. Drug Discov. 9, 57–67 (2010).
Bonetta, L. RNA-based therapeutics: ready for delivery? Cell 136, 581–584 (2009).
Couzin-Frankel, J. Drug research. Roche exits RNAi field, cuts 4800 jobs. Science 330, 1163 (2010).
Krieg, A. M. Is RNAi dead? Mol. Ther. 19, 1001–1002 (2011).
Stein, C. A. Does antisense exist? Nature Med. 1, 1119–1121 (1995).
Zamecnik, P. C. & Stephenson, M. L. Inhibition of Rous sarcoma virus replication and cell transformation by a specific oligodeoxynucleotide. Proc. Natl Acad. Sci. USA 75, 280–284 (1978).
Lima, W., Wu, H. & Crooke, S. T. in Antisense Drug Technology: Principles, Strategies, and Applications 2nd edn (ed. Crooke, S. T.) 47–74 (CRC Press, Boca Raton, Florida, 2007).
Raal, F. J. et al. Mipomersen, an apolipoprotein B synthesis inhibitor, for lowering of LDL cholesterol concentrations in patients with homozygous familial hypercholesterolemia: a randomized, double-blind, placebo-controlled trial. Lancet 375, 998–1006 (2010).
Chi, K. N. et al. Randomized Phase II study of docetaxel and prednisone with or without OGX-011 in patients with metastatic castration-resistant prostate cancer. J. Clin. Oncol. 28, 4247–4254 (2010).
Chia, S. et al. Phase II trial of OGX-011 in combination with docetaxel in metastatic breast cancer. Clin. Cancer Res. 15, 708–713 (2009).
Smith, C. C., Aurelian, L., Reddy, M. P., Miller, P. S. & Ts'o, P. O. Antiviral effect of an oligo(nucleoside methylphosphonate) complementary to the splice junction of herpes simplex virus type 1 immediate early pre-mRNAs 4 and 5. Proc. Natl Acad. Sci. USA 83, 2787–2791 (1986).
Stirchak, E. P., Summerton, J. E. & Weller, D. D. Uncharged stereoregular nucleic acid analogs: 2. Morpholino nucleoside oligomers with carbamate internucleoside linkages. Nucleic Acids Res. 17, 6129–6141 (1989).
Ozsolak, F. & Milos, P. M. RNA sequencing: advances, challenges and opportunities. Nature Rev. Genet. 12, 87–98 (2011).
Sazani, P., Graziewicz, M. & Kole, R. in Antisense Drug Technology: Principles, Strategies, and Applications 2nd edn (ed. Crooke, S. T.) 89–114 (CRC Press, Boca Raton, 2007).
Lu, Q. L. et al. Systemic delivery of antisense oligoribonucleotide restores dystrophin expression in body-wide skeletal muscles. Proc. Natl Acad. Sci. USA 102, 198–203 (2005).
Sazani, P. et al. Systemically delivered antisense oligomers upregulate gene expression in mouse tissues. Nature Biotech. 20, 1228–1233 (2002).
Dillman, J. et al. Efficient and persistent splice switching by systemically delivered LNA oligonucleotides in mice. Mol. Ther. 14, 471–475 (2006).
van Deutekom, J. C. et al. Local dystrophin restoration with antisense oligonucleotide PRO051. N. Engl. J. Med. 357, 2677–2686 (2007).
Kinali, M. et al. Local restoration of dystrophin expression with the morpholino oligomer AVI-4658 in Duchenne muscular dystrophy: a single-blind, placebo-controlled, dose-escalation, proof-of-concept study. Lancet Neurol. 8, 918–928 (2009).
Goemans, N. M. et al. Systemic administration of PRO051 in Duchenne's muscular dystrophy. N. Engl. J. Med. 364, 1513–1522 (2011).
Cirak, S. et al. Exon skipping and dystrophin restoration in patients with Duchenne muscular dystrophy after systemic phosphorodiamidate morpholino oligomer treatment: an open-label, Phase 2, dose-escalation study. Lancet 378, 595–605 (2011). These two studies showed encouraging results in clinical trials applying two SSO variants, 2′-OMe phosphorothioate (reference 31) and PMO (reference 32), for the systemic treatment of Duchenne muscular dystrophy.
Van Ommen, G. J., van Deutekom, J. & Aaartsma-Rus, A. The therapeutic potential of antisense-mediated exon skipping. Curr. Opin. Mol. Ther. 10, 140–149 (2008).
Muntoni, F., Torelli, S. & Ferlini, A. Dystrophin and mutations: one gene, several proteins, multiple phenotypes. Lancet Neurol. 2, 731–740 (2003).
Matsuo, M. et al. Exon skipping during splicing of dystrophin mRNA precursor due to an intraexon deletion in the dystrophin gene of Duchenne muscular dystrophy kobe. J. Clin. Invest. 87, 2127–2131 (1991).
Aartsma-Rus, A. et al. Theoretic applicability of antisense-mediated exon skipping for Duchenne muscular dystrophy mutations. Hum. Mutat. 30, 293–299 (2009).
Wilton, S. D. et al. Specific removal of the nonsense mutation from the mdx dystrophin mRNA using antisense oligonucleotides. Neuromuscul. Disord. 9, 330–338 (1999).
Bulfield, G., Siller, W., Wight, P. & Moore, K. X chromosome-linked muscular dystrophy (mdx) in the mouse. Proc. Natl Acad. Sci. USA 81, 1189–1192 (1984).
Lu, Q. L. et al. Functional amounts of dystrophin produced by skipping the mutated exon in the mdx dystrophic mouse. Nature Med. 9, 1009–1014 (2003).
Takeshima, Y. et al. Oligonucleotides against a splicing enhancer sequence led to dystrophin production in muscle cells from a Duchenne muscular dystrophy patient. Brain Dev. 23, 788–790 (2001).
van Deutekom, J. C. et al. Antisense-induced exon skipping restores dystrophin expression in DMD patient derived muscle cells. Hum. Mol. Genet. 10, 1547–1554 (2001).
Heemskerk, H. A. et al. In vivo comparison of 2′-O-methyl phosphorothioate and morpholino antisense oligonucleotides for Duchenne muscular dystrophy exon skipping. J. Gene Med. 11, 257–266 (2009).
Alter, J. et al. Systemic delivery of morpholino oligonucleotide restores dystrophin expression bodywide and improves dystrophic pathology. Nature Med. 12, 175–177 (2006).
Yokota, T. Efficacy of systemic morpholino exon-skipping in Duchenne dystrophy dogs. Ann. Neurol. 65, 667–676 (2009).
Muntoni, F. & Wood, M. J. Targeting RNA to treat neuromuscular disease. Nature Rev. Drug Discov. 10, 621–637 (2011).
Sazani, P., Weller, D. L. & Shrewsbury, S. B. Safety pharmacology and genotoxicity evaluation of AVI-4658. Int. J. Toxicol. 29, 143–156 (2010).
Wu, B. et al. Dose-dependent restoration of dystrophin expression in cardiac muscle of dystrophic mice by systemically delivered morpholino. Gene Ther. 17, 132–140 (2010).
Heemskerk, H. et al. Preclinical PK and PD studies on 2′-O-methyl-phosphorothioate RNA antisense oligonucleotides in the mdx mouse model. Mol. Ther. 18, 1210–1217 (2010).
Jearawiriyapaisarn, N. et al. Sustained dystrophin expression induced by peptide-conjugated morpholino oligomers in the muscles of mdx mice. Mol. Ther. 16, 1624–1629 (2008).
Jearawiriyapaisarn, N., Moulton, H. M., Sazani, P., Kole, R. & Willis, M. S. Long-term improvement in mdx cardiomyopathy after therapy with peptide-conjugated morpholino oligomers. Cardiovasc. Res. 85, 444–453 (2010).
Wu, B. et al. Effective rescue of dystrophin improves cardiac function in dystrophin-deficient mice by a modified morpholino oligomer. Proc. Natl Acad. Sci. USA 105, 14814–14819 (2008).
Goyenvalle, A. et al. Prevention of dystrophic pathology in severely affected dystrophin/utrophin-deficient mice by morpholino-oligomer-mediated exon-skipping. Mol. Ther. 18, 198–205 (2010).
Yin, H. et al. Pip5 transduction peptides direct high efficiency oligonucleotide-mediated dystrophin exon skipping in heart and phenotypic correction in mdx mice. Mol. Ther. 19, 1295–1303 (2011).
Youngblood, D. S., Hatlevig, S. A., Hassinger, J. N., Iversen, P. L. & Moulton, H. M. Stability of cell-penetrating peptide–morpholino oligomer conjugates in human serum and in cells. Bioconjug. Chem. 18, 50–60 (2007).
Lefebvre, S. et al. Identification and characterization of a spinal muscular atrophy-determining gene. Cell 80, 155–165 (1995).
Lorson, C. L., Hahnen, E., Androphy, E. J. & Wirth, B. A single nucleotide in the SMN gene regulates splicing and is responsible for spinal muscular atrophy. Proc. Natl Acad. Sci. USA 96, 6307–6311 (1999).
Hua, Y., Vickers, T. A., Baker, B. F., Bennett, C. F. & Krainer, A. R. Enhancement of SMN2 exon 7 inclusion by antisense oligonucleotides targeting the exon. PLoS Biol. 5, e73 (2007).
Singh, N. K., Singh, N. N., Androphy, E. J. & Singh, R. N. Splicing of a critical exon of human survival motor neuron is regulated by a unique silencer element located in the last intron. Mol. Cell Biol. 26, 1333–1346 (2006).
Hua, Y., Vickers, T. A., Okunola, H. L., Bennett, C. F. & Krainer, A. R. Antisense masking of an hnRNP A1/A2 intronic splicing silencer corrects SMN2 splicing in transgenic mice. Am. J. Hum. Genet. 82, 834–848 (2008).
Williams, J. H. et al. Oligonucleotide-mediated survival of motor neuron protein expression in CNS improves phenotype in a mouse model of spinal muscular atrophy. J. Neurosci. 29, 7633–7638 (2009).
Hua, Y. et al. Antisense correction of SMN2 splicing in the CNS rescues necrosis in a type III SMA mouse model. Genes Dev. 24, 1634–1644 (2010).
Passini, M. A. et al. Antisense oligonucleotides delivered to the mouse CNS ameliorate symptoms of severe spinal muscular atrophy. Sci. Transl. Med. 3, 72ra18 (2011).
Hua, Y. et al. Peripheral SMN restoration is essential for long-term rescue of a severe spinal muscular atrophy mouse model. Nature 478, 123–126 (2011). This paper reported the unexpected discovery that in a mouse model of spinal muscular atrophy the phenotype can be reversed not only by intrathecal but also by systemic peripheral delivery of an SSO that promotes the correct expression of the SMN2 gene.
Gibbons, R., Higgs, D. R., Olivieri, N. F. & Wood, W. G. in The Thalassaemia Syndromes 4th edn Ch. 7 (eds Weatherall, D. J. & Clegg, J. B.) 287–356 (Blackwell Science, Oxford, 2001).
Dominski, Z. & Kole, R. Restoration of correct splicing in thalassemic pre-mRNA by antisense oligonucleotides. Proc. Natl Acad. Sci. USA 90, 8673–8677 (1993). This was the first report of oligonucleotide-mediated RNA repair of thalassaemic pre-mRNA by SSOs.
Lewis, J. et al. A common human β globin splicing mutation modeled in mice. Blood 91, 2152–2156 (1998).
Svasti, S. et al. RNA repair restores hemoglobin expression in IVS2-654 thalassemic mice. Proc. Natl Acad. Sci. USA 106, 1205–1210 (2009).
Xie, S. Y. et al. Correction of β654-thalassaemia mice using direct intravenous injection of siRNA and antisense RNA vectors. Int. J. Hematol. 93, 301–310 (2011).
Cao, A. & Galanello, R. β-thalassemia. Genet. Med. 12, 61–76 (2010).
Taylor, J. K., Zhang, Q. Q., Wyatt, J. R. & Dean, N. M. Induction of endogenous Bcl-xS through the control of Bcl-x pre-mRNA splicing by antisense oligonucleotides. Nature Biotech. 17, 1097–1100 (1999).
Mercatante, D. R., Bortner, C. D., Cidlowski, J. A. & Kole, R. Modification of alternative splicing of Bcl-x pre-mRNA in prostate and breast cancer cells. analysis of apoptosis and cell death. J. Biol. Chem. 276, 16411–16417 (2001).
Graziewicz, M. A. et al. An endogenous TNF-α antagonist induced by splice-switching oligonucleotides reduces inflammation in hepatitis and arthritis mouse models. Mol. Ther. 16, 1316–1322 (2008).
Paquet, J. et al. Alternative for anti-TNF antibodies for arthritis treatment. Mol. Ther. 19, 1887–1895 (2011).
Miller, J. W. et al. Recruitment of human muscleblind proteins to (CUG)n expansions associated with myotonic dystrophy. EMBO J. 19, 4439–4448 (2000).
Lee, J. E. & Cooper, T. A. Pathogenic mechanisms of myotonic dystrophy. Biochem. Soc. Trans. 37, 1281–1286 (2009).
Wheeler, T. M. et al. Reversal of RNA dominance by displacement of protein sequestered on triplet repeat RNA. Science 325, 336–339 (2009). This paper describes the in vivo application of SSO-displacing splicing factors bound to an expanded triplet repeat as a potential treatment for myotonic dystrophy.
Mulders, S. A. et al. Triplet-repeat oligonucleotide mediated reversal of RNA toxicity in myotonic dystrophy. Proc. Natl Acad. Sci. USA 106, 13915–13920 (2009).
Nakamori, M., Gourdon, G. & Thornton, C. A. Stabilization of expanded (CTG)•(CAG) repeats by antisense oligonucleotides. Mol. Ther. 19, 2222–2227 (2011).
Geller, B. L., Deere, J., Tilley, L. & Iversen, P. L. Antisense phosphorodiamidate morpholino oligomer inhibits viability of Escherichia coli in pure culture and in mouse peritonitis. J. Antimicrob. Chemother. 55, 983–988 (2005).
Mellbye, B. L., Puckett, S. E., Tilley, L. D., Iversen, P. L. & Geller, B. L. Variations in amino acid composition of antisense peptide-phosphorodiamidate morpholino oligomer affect potency against Escherichia coli in vitro and in vivo. Antimicrob. Agents Chemother. 53, 525–530 (2009).
Mellbye, B. L. et al. Cationic phosphorodiamidate morpholino oligomers efficiently prevent growth of Escherichia coli in vitro and in vivo. J. Antimicrob. Chemother. 65, 98–106 (2010).
Shen, N. et al. Inactivation of expression of several genes in a variety of bacterial species by EGS technology. Proc. Natl Acad. Sci. USA 106, 8163–8168 (2009).
Lundblad, E. W. & Altman, S. Inhibition of gene expression by RNase P. Nature Biotech. 27, 212–221 (2010).
Wesolowski, D. et al. Basicpeptide–morpholino oligomer conjugate that is very effective in killing bacteria by gene-specific and nonspecific modes. Proc. Natl Acad. Sci. USA 108, 16582–16587 (2011). This paper describes the application of EGS technology using PPMOs as a potential treatment for drug-resistant bacterial infections.
Jiang, X. et al. Engineered external guide sequences effectively block viral gene expression and replication in cultured cells. J. Biol. Chem. 286, 322–330 (2011).
Warren, T. K. et al. Advanced antisense therapies for postexposure protection against lethal filovirus infections. Nature Med. 16, 991–994 (2010). This study showed that PMO-based translation-suppressing oligomers protect monkeys from lethal haemorrhagic infection by Ebola and Marburg viruses.
Lai, S. H. et al. Inhibition of respiratory syncytial virus infections with morpholino oligomers in cell cultures and in mice. Mol. Ther. 16, 1120–1128 (2008).
Eide, K. et al. Reduction of herpes simplex virus type-2 replication in cell cultures and in rodent models with peptide-conjugated morpholino oligomers. Antivir. Ther. 15, 1141–1149 (2010).
Stone, J. K. et al. A morpholino oligomer targeting highly conserved internal ribosome entry site sequence is able to inhibit multiple species of picornavirus. Antimicrob. Agents Chemother. 52, 1970–1981 (2008).
Gabriel, G., Nordmann, A., Stein, D. A., Iversen, P. L. & Klenk, H. D. Morpholino oligomers targeting the PB1 and NP genes enhance the survival of mice infected with highly pathogenic influenza A H7N7 virus. J. Gen. Virol. 89, 939–948 (2008).
Lupfer, C. et al. Inhibition of influenza A H3N8 virus infections in mice by morpholino oligomers. Arch. Virol. 153, 929–937 (2008).
Paessler, S. et al. Inhibition of alphavirus infection in cell culture and in mice with antisense morpholino oligomers. Virology 376, 357–370 (2008).
Lipinski, C. A. Drug-like properties and the causes of poor solubility and poor permeability. J. Pharmacol. Toxicol. Methods 44, 235–249 (2000). This paper describes desirable properties for an effective drug, and leads the reader to the conclusion that oligonucleotides lack such criteria.
Köhler, G. & Milstein, C. Continuous cultures of fused cells secreting antibody of predefined specificity. Nature 256, 495–497 (1975).
Waldmann, T. A. Immunotherapy: past, present and future. Nature Med. 9, 269–277 (2003).
Stephenson, M. L. & Zamecnik, P. C. Inhibition of Rous sarcoma viral RNA translation by a specific oligodeoxyribonucleotide. Proc. Natl Acad. Sci. USA 75, 285–288 (1978). This paper describes the discovery of an oligonucleotide that acts as an antisense compound that is capable of preventing viral replication.
Matsukura, M. et al. Phosphorothioate analogs of oligodeoxynucleotides: inhibitors of replication and cytopathic effects of human immunodeficiency virus. Proc. Natl Acad. Sci. USA 84, 7706–7710 (1987).
Agrawal, S., Mayrand, S. H., Zamecnik, P. C. & Pederson, T. Site-specific excision from RNA by RNase H and mixed-phosphate-backbone oligodeoxynucleotides. Proc. Natl Acad. Sci. USA 87, 1401–1405 (1990).
Geary, R. S., Yu, R. Z. & Levin, A. A. in Antisense Drug Technologies: Principles, Strategies, and Applications 2nd edn (ed. Crooke, S. T.) 183–217 (CRC Press, Boca Raton, Florida, 2007).
Krieg, A. M. Therapeutic potential of Toll-like receptor 9 activation. Nature Rev. Drug Discov. 5, 471–484 (2006).
Agarwala, S. S. et al. LDH correlation with survival in advanced melanoma from two large, randomised trials (Oblimersen GM301 and EORTC 18951). Eur. J. Cancer 45, 1807–1814 (2009).
Chanan-Khan, A. A. et al. Phase III randomised study of dexamethasone with or without oblimersen sodium for patients with advanced multiple myeloma. Leuk. Lymphoma 50, 559–565 (2009).
Morrison, T. Genta plunges on failed Phase III survival analysis for Genasense. BioWorld website [online], (2011).
Berget, S. M., Moore, C. & Sharp, P. A. Spliced segments at the 5′ terminus of adenovirus 2 late mRNA. Proc. Natl Acad. Sci. USA 74, 3171–3175 (1977).
Chow, L. T., Gelinas, R. E., Broker, T. R. & Roberts, R. J. An amazing sequence arrangement at the 5′ ends of adenovirus 2 messenger RNA. Cell 12, 1–8 (1977).
Spritz, R. A. et al. Base substitution in an intervening sequence of a beta+-thalassemic human globin gene. Proc. Natl Acad. Sci. USA 78, 2455–2459 (1981).
Busslinger, M., Moschonas, N. & Flavell, R. A. β+ thalassemia: aberrant splicing results from a single point mutation in an intron. Cell 27, 289–298 (1981).
Tazi, J., Bakkour, N. & Stamm, S. Alternative splicing and disease. Biochim. Biophys. Acta 1792, 14–26 (2009).
Acknowledgements
R.K. would like to thank his past and present colleagues at AVI BioPharma for helpful comments on this article. A.R.K. thanks his collaborators at Cold Spring Harbor Laboratory and Isis Pharmaceuticals for helpful discussions.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
Ryszard Kole is an employee and shareholder of AVI BioPharma.
Adrian R. Krainer is a consultant for, and has ongoing collaborations with, Isis Pharmaceuticals.
Sidney Altman declares no competing financial interests.
Related links
Glossary
- RNA interference
-
(RNAi). A form of post-transcriptional gene silencing in which the expression or transfection of double-stranded RNA induces degradation by nucleases of the homologous endogenous transcripts, resulting in the reduction or loss of gene activity.
- Antisense oligonucleotides
-
Oligonucleotides that bind to complementary mRNA by base pairing and induce cleavage of targeted mRNA by ribonuclease H, an enzyme that degrades RNA in RNA–DNA duplexes.
- Small interfering RNAs
-
(siRNAs). Synthetic, short, 21–22-nucleotide-long double-stranded RNAs with chemical modifications designed to increase their stability and cellular uptake. One strand of siRNA hybridizes to targeted mRNA and allows mRNA degradation.
- RNA-induced silencing complex
-
(RISC). A multiprotein complex that, when combined with small interfering RNA (siRNA), affects mRNA degradation. A key component of RISC is an endonuclease, argonaute 2, which cleaves the targeted mRNA within the siRNA–mRNA duplex.
- Translation-suppressing oligonucleotides
-
(TSOs). Modified oligonucleotides that block mRNA sequences near the initiation of the translation codon (AUG), interfere with the binding of ribosomes to mRNA and inhibit the translation of undesirable proteins. TSO–mRNA duplexes are not recognized by ribonuclease H or RNA-induced silencing complex, and the mRNA is therefore not cleaved.
- External guide sequence
-
(EGS). A short RNA sequence designed to bind to targeted mRNA and form a structure that is recognized by a tRNA-processing ribozyme, ribonuclease P. Ribonuclease P cleaves mRNA and thereby downregulates the function of a targeted gene.
- Splice-switching oligonucleotides
-
(SSOs). Chemically modified oligonucleotides that block sequences in pre-mRNA that are involved in pre-mRNA splicing, and redirect mRNA splicing pathways. SSO–pre-mRNA duplexes are not recognized by ribonuclease H or the RNA-induced silencing complex, and the pre-mRNA is thus not cleaved.
- Alternative splicing
-
Splicing of pre-mRNA to yield more than one kind of mRNA — that is, different splice variants — by frequently including or excluding an exon.
- Translational reading frame
-
Arrangement of mRNA nucleotides into triplets (codons) that, when read by the ribosome, are translated into one amino acid per codon. The reading frame usually starts with a translation initiation codon, AUG — for example, AUG UUU ACA GCA. Deletion of a nucleotide, such as the third uridylic in the second codon, changes the reading frame to AUG 'UUA CAG CA', thus preventing the translation of the desired protein.
- Exonic splicing silencers
-
Sequences present in exons in pre-mRNA that contribute to the modulation of splicing. They can inhibit the inclusion of a given exon into mature mRNA during splicing. Exonic splicing enhancers also exist.
- Intronic splicing silencers
-
Sequences present in introns in pre-mRNA that contribute to the modulation of splicing. They can inhibit the inclusion of a given intron into mature mRNA during splicing. Intronic splicing enhancers also exist.
- Cryptic splice site
-
A splice site that is not functional under normal conditions but becomes activated if a mutation modifies its sequence to resemble a functional splice site or if an adjacent normal splice site is inactivated by a mutation.
Rights and permissions
About this article
Cite this article
Kole, R., Krainer, A. & Altman, S. RNA therapeutics: beyond RNA interference and antisense oligonucleotides. Nat Rev Drug Discov 11, 125–140 (2012). https://doi.org/10.1038/nrd3625
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrd3625
This article is cited by
-
Splice-Modulating Antisense Oligonucleotides as Therapeutics for Inherited Metabolic Diseases
BioDrugs (2024)
-
Cardiovascular complications of diabetes: role of non-coding RNAs in the crosstalk between immune and cardiovascular systems
Cardiovascular Diabetology (2023)
-
Derivatization with fatty acids in peptide and protein drug discovery
Nature Reviews Drug Discovery (2023)
-
Convergent synthesis of phosphorodiamidate morpholino oligonucleotides (PMOs) by the H-phosphonate approach
Scientific Reports (2023)
-
Construction and Application of DNAzyme-based Nanodevices
Chemical Research in Chinese Universities (2023)