Key Points
-
Modelling and simulation are now being recognized as important aids to the drug development process. An important element of this is the ability to predict in vivo pharmacokinetic behaviour from in vitro data.
-
Most attempts at in vitro–in vivo extrapolation (IVIVE) have focused on predicting mean values of drug clearance and the impact of drug–drug interactions in the 'average' subject. This review considers methodology to assess outcomes in populations of virtual patients, thereby adding the capacity to identify the characteristics of individuals at particular risk of inadequate or excessive drug exposure.
-
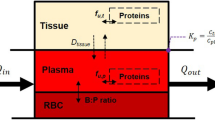
Factors influencing variability in hepatic drug clearance are considered in detail. These include the abundance of different hepatic drug metabolizing enzymes, hepatic microsomal protein and hepatocellularity, liver weight, hepatic blood flow, plasma binding and haematocrit.
-
Based on Monte Carlo simulation, the incorporation of variability in pharmacokinetic factors in mechanistic, physiologically based models, together with demographic data and specific information on the genetic variability of enzymes, allows prediction of the net exposure and time-course of drugs in the body across patient populations.
-
Applications of the approach in predicting drug clearance in adults from different ethnic backgrounds and in neonates, infants and children are described, as well as its utility in the assessment of inter-individual variability in metabolically based drug–drug interactions.
-
The authors conclude that considerable progress has been made towards predicting pharmacokinetic behaviour from in vitro information on the interaction between drug molecules and enzymes. Further challenges are indicated in the area of enzyme-transporter interplay, and in the linkage of pharmacokinetic predictions to models of pharmacodynamic response.
Abstract
The perceived failure of new drug development has been blamed on deficiencies in in vivo studies of drug efficacy and safety. Prior simulation of the potential exposure of different individuals to a given dose might help to improve the design of such studies. This should also help researchers to focus on the characteristics of individuals who present with extreme reactions to therapy. An effort to build virtual populations using extensive demographic, physiological, genomic and in vitro biochemical data to simulate and predict drug disposition from routinely collected in vitro data is outlined.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Uehling, M.D. Model Patient. Bio-IT World web site [online], (2003).
Lin, J. H. Sense and nonsense in the prediction of drug–drug interactions. Curr. Drug Metab. 1, 305–331 (2000).
Clarke, S. E. & Jeffrey, P. Utility of metabolic stability screening: comparison of in vitro and in vivo clearance. Xenobiotica 31, 591–598 (2001).
Andersson, T. B., Bredberg, E., Ericsson, H. & Sjoberg, H. An evaluation of the in vitro metabolism data for predicting the clearance and drug–drug interaction potential of CYP2C9 substrates. Drug Metab. Dispos. 32, 715–721 (2004).
Rostami-Hodjegan, A. & Tucker, G. T. 'In silico' simulations to assess the 'in vivo' consequences of 'in vitro' metabolic drug–drug interactions. Drug Discov. Today Technol. 1, 441–448 (2004). A complementary article to this review that places emphasis on issues related to metabolic drug–drug interactions.
Maxwell, C., Domenet, J. G. & Joyce, C. R. Instant experience in clinical trials: a novel aid to teaching by simulation. J. Clin. Pharmacol. New Drugs 11, 323–331 (1971).
Holford, N. H., Kimko, H. C., Monteleone, J. P. & Peck, C. C. Simulation of clinical trials. Annu. Rev. Pharmacol. Toxicol. 40, 209–234 (2000). Essential reading for those interested in clinical trial simulations.
Marsh, B. T. The planning of a clinical trials workshop. Br. J. Clin. Pharmacol. 2, 455–461 (1975).
Bland, J. M. Computer simulation of a clinical trial as an aid to teaching the concept of statistical significance. Stat. Med. 5, 193–197 (1986).
Madsen, B. W. et al. Clinical trial experience by simulation: a workshop report. BMJ 2, 1333–1335 (1978).
Jackson, P. R., Tucker, G. T., Lennard, M. S. & Woods, H. F. Polymorphic drug oxidation: pharmacokinetic basis and comparison of experimental indices. Br. J. Clin. Pharmacol. 22, 541–550 (1986).
Jackson, P. R. & Tucker, G. T. Pharmacokinetic–pharmacogenetic modelling in the detection of polymorphisms in xenobiotic metabolism. Ann. Occup. Hyg. 34, 653–662 (1990).
Jackson, P. R., Tucker, G. T. & Woods, H. F. Backtracking booze with Bayes — the retrospective interpretation of blood alcohol data. Br. J. Clin. Pharmacol. 31, 55–63 (1991). A practical example illustrating the importance of attention to variability.
Rostami-Hodjegan, A., Jackson, P. R. & Tucker, G. T. Sensitivity of indirect metrics for assessing 'rate' in bioequivalence studies — moving the 'goalposts' or changing the 'game'. J. Pharm. Sci. 83, 1554–1557 (1994).
Rostami-Hodjegan, A., Nurminen, S., Jackson, P. R. & Tucker, G. T. Caffeine urinary metabolite ratios as markers of enzyme activity: a theoretical assessment. Pharmacogenetics 6, 121–149 (1996).
Bogaards, J. J. et al. Prediction of interindividual variation in drug plasma levels in vivo from individual enzyme kinetic data and physiologically based pharmacokinetic modeling. Eur. J. Pharm. Sci. 12, 117–124 (2000).
Jonsson, F. & Johanson, G. The Bayesian population approach to physiological toxicokinetic–toxicodynamic models — an example using the MCSim software. Toxicol. Lett. 138, 143–150 (2003).
MacDonald, A. J., Rostami-Hodjegan, A., Tucker, G. T. & Linkens, D. A. in Proc. 1999 Health Sci. Simul. Conf. 103–108 (eds Andersen, J. G. & Katzper, M.) (Society Comp. Simul. Int., San Francisco, 1999).
MacDonald, A. J., Rostami-Hodjegan, A., Tucker, G. T. & Linkens, D. A. Analysis of solvent central nervous system toxicity and ethanol interactions using a human population physiologically based kinetic and dynamic model. Regul. Toxicol. Pharmacol. 35, 165–176 (2002).
Nestorov, I., Gueorguieva, I., Jones, H. M., Houston, B. & Rowland, M. Incorporating measures of variability and uncertainty into the prediction of in vivo hepatic clearance from in vitro data. Drug Metab. Dispos. 30, 276–282 (2002). An early indication of the need to incorporate variability in physiologically based pharmacokinetic models, though the clearance component did not include biological sources of variation.
Yang, J., Rostami-Hodjegan, A. & Tucker, G. T. Prediction of ritonavir interaction with sildenafil (Viagra): incorporating population variability. Br. J. Clin. Pharmacol. 53, 438P (2002).
Jonsson, E. N. & Sheiner, L. B. More efficient clinical trials through use of scientific model-based statistical tests. Clin. Pharmacol. Ther. 72, 603–614 (2002).
Ribbing, J. & Jonsson, E. N. Power, selection bias and predictive performance of the Population Pharmacokinetic Covariate model. J. Pharmacokinet. Pharmacodyn. 31, 109–134 (2004).
Lave, T., Coassolo, P. & Reigner, B. Prediction of hepatic metabolic clearance based on interspecies allometric scaling techniques and in vitro–in vivo correlations. Clin. Pharmacokinet. 36, 211–231 (1999).
Zuegge, J., Schneider, G., Coassolo, P. & Lave, T. Prediction of hepatic metabolic clearance: comparison and assessment of prediction models. Clin. Pharmacokinet. 40, 553–563 (2001).
Pang, K. S. & Chiba, M. in Handbook of Experimental Pharmacology: Pharmacokinetics of Drugs Vol 110 (eds Welling, P. G. & Balant, L. P.) 101–187 (Springer, Berlin, 1994).
Wilkinson, G. R. & Shand, D. G. Commentary: a physiological approach to hepatic drug clearance. Clin. Pharmacol. Ther. 18, 377–390 (1975). A classic text for those who wish to understand the link between in vitro and in vivo metabolism in the liver.
Proctor, N. J., Tucker, G. T. & Rostami-Hodjegan, A. Predicting drug clearance from recombinantly expressed CYPs: intersystem extrapolation factors. Xenobiotica 34, 151–178 (2004).
Yang, J., Jamei, M., Yeo, K. R., Tucker, G. T. & Rostami-Hodjegan, A. Kinetic values for mechanism-based enzyme inhibition: assessing the bias introduced by the conventional experimental protocol. Eur. J. Pharm. Sci. 26, 334–340 (2005).
Johnson, T. N., Rostami-Hodjegan, A. & Tucker, G. T. Prediction of the clearance of eleven drugs and associated variability in neonates, infants and children. Clin. Pharmacokinet. 45, 931–956 (2006). An essential reference for anyone seeking information on anatomical and physiological changes from childhood to adulthood. The implications of each element are described with examples. The report also identifies the paucity of data in certain areas.
Rowland-Yeo, K., Rostami-Hodjegan, A. & Tucker, G. T. Abundance of cytochromes P450 in human liver: a meta-analysis. Br. J. Clin. Pharmacol. 57, 687–688 (2004).
Inoue, S. et al. Prediction of in vivo drug clearance from in vitro data. II. Potential inter-ethnic differences. Xenobiotica 36, 499–513 (2006).
Houston, J. B. & Carlile, D. J. Prediction of hepatic clearance from microsomes, hepatocytes, and liver slices. Drug Metab. Rev. 29, 891–922 (1997). A comprehensive review of the issues involved in extrapolating in vitro data on clearance to clinical studies.
Barter, Z. E. et al. Scaling factors for the extrapolation of in vivo metabolic drug clearance from in vitro data: reaching a consensus on values of human microsomal protein and hepatocellularity per gram of liver. Curr. Drug Metab. 8, 35–45 (2007).
Hakooz, N. et al. determination of a human hepatic microsomal scaling factor for predicting in vivo drug clearance. Pharm. Res. 23, 533–539 (2006).
Johnson, T. N., Tucker, G. T., Tanner, M. S. & Rostami-Hodjegan, A. Changes in liver volume from birth to adulthood: a meta-analysis. Liver Transpl. 11, 1481–1493 (2005).
Guyton, A. C. & Hall J. E. in Textbook of Medical Physiology 9th edn (eds Guyton, A. C. & Hall, J. E.) (WB Saunders Co., Philadelphia,1996).
Mathias, C. J. Effect of food intake on cardiovascular control in patients with impaired autonomic function. J. Neurosci. Methods 34, 193–200 (1990).
Matheson, P. J., Wilson, M. A. & Garrison, R. N. Regulation of intestinal blood flow. J. Surg. Res. 93, 182–196 (2000).
Bernardi, M. et al. The hemodynamic status of preascitic cirrhosis: an evaluation under steady-state conditions and after postural change. Hepatology 16, 341–346 (1992).
Edell, E. S., Cortese, D. A., Krowka, M. J. & Rehder, K. Severe hypoxemia and liver disease. Am. Rev. Respir. Dis. 140, 1631–1635 (1989).
Ruckert, K. H. & Juchems, R. [Hemodynamics in posture changes]. Z. Kreislaufforsch. 59, 685–698 (1970) (in German).
Wong, F., Liu, P., Allidina, Y. & Blendis, L. The effect of posture on central blood volume in patients with preascitic cirrhosis on a sodium-restricted diet. Hepatology 23, 1141–1147 (1996).
Lobell, M. & Sivarajah, V. In silico prediction of aqueous solubility, human plasma protein binding and volume of distribution of compounds from calculated pKa and AlogP98 values. Mol. Divers. 7, 69–87 (2003).
Ekins, S. & Wrighton, S. A. Application of in silico approaches to predicting drug–drug interactions. J. Pharmacol. Toxicol. Methods 45, 65–69 (2001).
Lewis, D. F. Structural characteristics of human P450s involved in drug metabolism: QSARs and lipophilicity profiles. Toxicology 144, 197–203 (2000).
McNamara, P. J. & Alcorn, J. Protein binding predictions in infants. AAPS PharmSci. 4, e4 (2002).
Howgate, E. M., Rowland-Yeo, K., Proctor, N. J., Tucker, G. T. & Rostami-Hodjegan, A. Prediction of in vivo drug clearance from in vitro data. I. Impact of inter-individual variability. Xenobiotica 36, 473–497 (2006).
Shiran, M. R. et al. Prediction of metabolic drug clearance in humans: in vitro–in vivo extrapolation vs allometric scaling. Xenobiotica 36, 567–580 (2006).
Bjornsson, T. D. et al. A review and assessment of potential sources of ethnic differences in drug responsiveness. J. Clin. Pharmacol. 43, 943–967 (2003).
Imaoka, S. et al. Multiple forms of human P450 expressed in Saccharomyces cerevisiae. Systematic characterization and comparison with those of the rat. Biochem. Pharmacol. 51, 1041–1050 (1996).
Nakasa, H. et al. Characterization of human liver microsomal cytochrome P450 involved in the reductive metabolism of zonisamide. Mol. Pharmacol. 44, 216–221 (1993).
Shu, Y. et al. Interindividual variations in levels and activities of cytochrome P-450 in liver microsomes of Chinese subjects. Acta Pharmacol. Sin. 22, 283–288 (2001).
Tateishi, T. et al. A comparison of hepatic cytochrome P450 protein expression between infancy and postinfancy. Life Sci. 61, 2567–2574 (1997).
Tateishi, T. et al. No ethnic difference between Caucasian and Japanese hepatic samples in the expression frequency of CYP3A5 and CYP3A7 proteins. Biochem. Pharmacol. 57, 935–939 (1999).
Yasumori, T., Murayama, N., Yamazoe, Y. & Kato, R. Polymorphism in hydroxylation of mephenytoin and hexobarbital stereoisomers in relation to hepatic P-450 human-2. Clin. Pharmacol. Ther. 47, 313–322 (1990).
Yasumori, T. et al. Cytochrome P450 mediated metabolism of diazepam in human and rat: involvement of human CYP2C in N-demethylation in the substrate concentration-dependent manner. Pharmacogenetics 3, 291–301 (1993).
Dickinson, G. L., Lennard, M. S., Tucker, G. T. & Rostami-Hodjegan, A. The use of mechanistic DM–PK–PD modelling to assess the power of pharmacogenetic studies — CYP2C9 and warfarin as an example. Br. J. Clin. Pharmacol. (in the press). A guide to using information on drug metabolizing enzymes to assess the pharmacokinetic and pharmacodynamic consequences of genetic variation.
Ito, K., Brown, H. S. & Houston, J. B. Database analyses for the prediction of in vivo drug–drug interactions from in vitro data. Br. J. Clin. Pharmacol. 57, 473–486 (2004).
Ito, K., Hallifax, D., Obach, R. S. & Houston, J. B. Impact of parallel pathways of drug elimination and multiple cytochrome P450 involvement on drug–drug interactions: CYP2D6 paradigm. Drug Metab. Dispos. 33, 837–844 (2005).
Tucker, G. T. The rational selection of drug interaction studies: implications of recent advances in drug metabolism. Int. J. Clin. Pharmacol. Ther. 30, 550–553 (1992). An early comprehensive report on issues related to assessing drug–drug interactions using in vitro systems.
Food and Drug Administration (FDA). Guidance for industry: drug interaction studies — study design, data analysis, and implications for dosing and labeling. FDA web site [online], (2006). A must-read guide that provides an insight into the current thinking of regulatory authorities on issues related to the use of in vitro systems as an alternative to in vivo studies, or as a framework for deciding the type and design of studies to carry out.
Yang, J. et al. Implications of mechanism-based inhibition of CYP2D6 for the pharmacokinetics and toxicity of MDMA. J. Psychopharmacol. 20, 842–849 (2006).
Howgate, E. M., Rowland Yeo, K., Proctor, N. J., Tucker, G. T. & Rostami Hodjegan, A. Prediction of in vivo drug clearance from in vitro data. I: impact of inter-individual variability. Xenobiotica. 36, 473–497 (2006).
Acknowledgements
The authors wish to thank Ben Meakin for organizing the bibliography for this review.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
A.R.H. and G.T. are founding members of and shareholders in Simcyp, a University of Sheffield spin-out company that develops algorithms for the in vitro prediction of drug metabolism. Simcyp's databases and related software are available free to appropriate non-profit organizations to facilitate research into in vitro–in vivo extrapolation (IVIVE).
Glossary
- Monte Carlo methods
-
A method of generating virtual entities with randomly assigned characteristics.
- FORTRAN
-
A computer programming language, which is not commonly used anymore.
- Quantitative Framework
-
In this context, a quantitative framework is essentially the systems biology of the model relevant to ADME (absorption, distribution, metabolism and excretion). It enables one to rationalize non-linear relationships between different ADME elements in a way that is not possible using non-quantitative or rank order frameworks.
- Haematocrit
-
The proportion of the volume of a sample of blood that is represented by red blood cells.
- Central tendency
-
Central tendency refers to a representation of a widely variable distribution of sets of numbers by their mode, median, average and so on. It is a known phrase in referring to average, but sometimes the average may not be the right measure, so central tendency is wider than the meaning of average but it may include average in many cases.
- Allometric scaling methods
-
Methods that extrapolate pharmacokinetic parameters between different species, or various age groups within the same species, based on the assumption that there is a link between the body size and the magnitude of each pharmacokinetic parameter.
Rights and permissions
About this article
Cite this article
Rostami-Hodjegan, A., Tucker, G. Simulation and prediction of in vivo drug metabolism in human populations from in vitro data. Nat Rev Drug Discov 6, 140–148 (2007). https://doi.org/10.1038/nrd2173
Issue Date:
DOI: https://doi.org/10.1038/nrd2173
This article is cited by
-
Kinetic Characterization of Estradiol Glucuronidation by Liver Microsomes and Expressed UGT Enzymes: The Effects of Organic Solvents
European Journal of Drug Metabolism and Pharmacokinetics (2024)
-
Maternal Ezetimibe Concentrations Measured in Breast Milk and Its Use in Breastfeeding Infant Exposure Predictions
Clinical Pharmacokinetics (2024)
-
Prediction of Drug–Drug Interaction Between Dabrafenib and Irinotecan via UGT1A1-Mediated Glucuronidation
European Journal of Drug Metabolism and Pharmacokinetics (2022)
-
Development and evaluation of physiologically based pharmacokinetic drug-disease models for predicting captopril pharmacokinetics in chronic diseases
Scientific Reports (2021)