Key Points
-
Molecular de novo design, which involves incremental construction of a ligand model within a model of the receptor or enzyme active site, produces novel molecular structures with desired pharmacological properties from scratch.
-
De novo molecule-design software is confronted with a virtually infinite search space. As such a large space prohibits exhaustive searching, navigation in the de novo design process relies on the principle of local optimization.
-
Basically, three questions have to be addressed by a de novo design program: how to assemble the candidate compounds; how to evaluate their potential quality; and how to sample the search space effectively.
-
This review gives an overview of computer-based molecular de novo design methods on a conceptual level, considering these three questions, and focusing on the design of small, drug-like molecules. Successful examples of de novo design in the hit- and lead-finding stages of the drug discovery process are used to show that de novo design provides a method for lead identification.
-
De novo design can therefore be regarded as a complement to other virtual techniques, such as database searching, and non-virtual techniques such as high-throughput screening. We also accentuate strengths and weaknesses of current de novo design approaches.
Abstract
Ever since the first automated de novo design techniques were conceived only 15 years ago, the computer-based design of hit and lead structure candidates has emerged as a complementary approach to high-throughput screening. Although many challenges remain, de novo design supports drug discovery projects by generating novel pharmaceutically active agents with desired properties in a cost- and time-efficient manner. In this review, we outline the various design concepts and highlight current developments in computer-based de novo design.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Accession codes
References
Dobson, C. M. Chemical space and biology. Nature 432, 824–828 (2004).
Lipinski, C. & Hopkins, A. Navigating chemical space for biology and medicine. Nature 432, 855–861 (2004).
Schneider, G. Trends in virtual combinatorial library design. Curr. Med. Chem. 9, 2095–2101 (2002).
Richardson, J. S. & Richardson, D. C. The de novo design of protein structures. Trends Biochem. Sci. 14, 304–309 (1989).
Richardson, J. S. et al. Looking at proteins: representations, folding, packing, and design. Biophys. J. 63, 1185–1209 (1992).
Moon, J. B. & Howe, W. J. Computer design of bioactive molecules: a method for receptor-based de novo ligand design. Proteins 11, 314–328 (1991).
Schneider, G. & Wrede, P. The rational design of amino acid sequences by artificial neural networks and simulated molecular evolution: de novo design of an idealized leader peptidase cleavage site. Biophys. J. 66, 335–344 (1994).
Schneider, G. et al. Peptide design by artificial neural networks and computer-based evolutionary search. Proc. Natl Acad. Sci. USA 95, 12179–12184 (1998).
Venkatasubramanian, V., Chan, K. & Caruthers, J. M. Computer-aided molecular design using genetic algorithms. Computers Chem. Eng. 18, 833–844 (1994).
Venkatasubramanian, V., Sundaram, A., Chan, K. & Caruthers, J. M. in Genetic Algorithms in Molecular Modelling (ed. Devillers, J.) 271–302 (Academic, London, 1996).
Sundaram, A. & Venkatasubramanian, V. Parametric sensitivity and search-space characterization studies of genetic algorithms for computer-aided polymer design. J. Chem. Inf. Comput. Sci. 38, 1177–1191 (1998).
Danziger, D. J. & Dean, P. M. Automated site-directed drug design: a general algorithm for knowledge acquisition about hydrogen-bonding regions at protein surfaces. Proc R Soc Lond B Biol Sci B 236, 101–113 (1989). First work about interaction site derivation from a receptor structure tailored for the use in automated de novo design.
Böhm, H. -J. The computer program LUDI: a new simple method for the de-novo design of enzyme inhibitors. J. Comput. Aided Mol. Des. 6, 61–78 (1992).
Böhm, H. -J. LUDI: rule-based automatic design of new substituents for enzyme inhibitor leads. J. Comput. Aided Mol. Des. 6, 593–606 (1992).
Clark, D. E. et al. PRO LIGAND: an approach to de novo molecular design. 1. Application to the design of organic molecules. J. Comput. Aided Mol. Des. 9, 13–32 (1995). A comprehensive approach that adopts a lot of earlier ideas and provides new concepts.
Murray, C. W. et al. PRO_SELECT: combining structure-based drug design and combinatorial chemistry for rapid lead discovery. 1. Technology. J. Comp. Aided Mol. Des. 11, 193–207 (1997).
Gillett, V. J., Myatt, G., Zsoldos, Z. & Johnson, A. P. SPROUT, HIPPO and CAESA: tools for de novo structure generation and estimation of synthetic accessibility. Perspect. Drug Discov. Des. 3, 34–50 (1995).
Rotstein, S. H. & Murcko, M. A. GroupBuild: a fragment-based method for de novo drug design. J. Med. Chem. 36, 1700–1710 (1993).
Goodford, P. J. A computational procedure for determining energetically favorable binding sites on biologically important macromolecules. J. Med. Chem. 28, 849–857 (1985).
Nishibata, Y. & Itai, A. Automatic creation of dug candidate structures based on receptor structure. Starting point for artificial lead generation. Tetrahedron 47, 8985–8990 (1991).
Bohacek, R. S. & McMartin, C. Multiple highly diverse structures complementary to enzyme binding sites: results of extensive application of a de novo design method incorporating combinatorial growth. J. Am. Chem. Soc. 116, 5560–5571 (1994).
Glen, R. C. & Payne, A. W. R. A genetic algorithm for the automated generation of molecules within constraints. J. Comput. Aided. Mol. Des. 9, 181–202 (1995).
Luo, Z., Wang, R. & Lai, L. RASSE: a new method for structure-based drug design. J. Chem. Inf. Comput. Sci. 36, 1187–1194 (1996).
Wang, R., Gao, Y. & Lai, L. LigBuilder: a multi-purpose program for structure-based drug design. J. Mol. Model. 6, 498–516 (2000).
Eisen, M. B., Wiley, D. C., Karplus, M. & Hubbard, R. E. HOOK: a program for finding novel molecular architectures that satisfy the chemical and steric requirements of a macromolecule binding site. Proteins 19, 199–221 (1994).
Miranker, A. & Karplus, M. An automated method for dynamic ligand design. Proteins 23, 472–490 (1995).
Miranker, A. & Karplus, M. Functionality maps of binding sites: a multiple copy simultaneous search method. Proteins 11, 29–34 (1991).
Lewis, R. A. et al. Automated site-directed drug design using molecular lattices. J. Mol. Graphics 10, 66–78 (1992).
Roe, D. C. & Kuntz, I. D. BUILDER v.2: improving the chemistry of a de novo design strategy. J. Comput. Aided Mol. Des. 9, 269–282 (1995).
Tschinke, V. & Cohen, N. C. The NEWLEAD program: a new method for the design of candidate structures from pharmacophoric hypothesis. J. Med. Chem. 36, 3863–3870 (1993).
Lewis, R. A. & Dean, P. M. Automated site-directed drug design: the formation of molecular templates in primary structure generation. Proc R Soc Lond B Biol Sci 236, 141–162 (1989).
Gillett, V. A., Johnson, A. P., Mata, P. & Sike, S. Automated structure design in 3D. Tetrahedron Comput. Method. 3, 681–696 (1990).
Lewis, R. A. Automated site-directed drug design: approaches to the formation of 3D molecular graphs. J. Comput. Aided Mol. Des. 4, 205–210 (1990).
Rotstein, S. H. & Murcko, M. A. GenStar: a method for de novo drug design. J. Comput. Aided. Mol. Des. 7, 23–43 (1993).
Pearlman, D. A. & Murcko, M. A. CONCERTS: dynamic connection of fragments as an approach to de novo ligand design. J. Med. Chem. 39, 1651–1663 (1996). Introduces the concept of consensus molecular dynamics as a method for structure sampling to de novo design.
Liu, H., Duan, Z., Luo, Q. & Shi, Y. Structure-based ligand design by dynamically assembling molecular building blocks at binding site. Proteins 36, 462–470 (1999).
Zhu, J., Yu, H., Fan, H. Liu, H. & Shi, Y. Design of selective inhibitors of cyclooxygenase-2 dynamic assembly of molecular building blocks. J. Comput. Aided Mol. Des. 15, 447–463 (2001).
Zhu, J., Fan, H., Liu, H. & Shi, Y. Structure-based ligand design for flexible proteins: application of new F-DycoBlock. J. Comput. Aided Mol. Des. 15, 979–996 (2001).
Pearlman, D. A. & Murcko, M. A. CONCEPTS: new dynamic algorithm for de novo design suggestion. J. Comput. Chem. 14, 1184–1193 (1993).
Eldridge, M. D., Murray, C. W., Auton, T. R., Paolini, G. V. & Mee, R. P. Empirical scoring functions: I. The development of a fast empirical scoring function to estimate the binding affinity of ligands in receptor complexes. J. Comput. Aided Mol. Des. 11, 425–445 (1997).
DeWitte, R. S. & Shakhnovich, E. I. SMoG de novo design method based on simple, fast, and accurate free energy estimates. 1. Methodology and supporting evidence. J. Am. Chem. Soc. 118, 11733–11744 (1996).
Ishchenko, A. V. & Shakhnovich, E. I. SMall Molecule Growth 2001 (SMoG2001): an improved knowledge-based scoring function for protein–ligand interactions. J. Med. Chem. 45, 2770–2780 (2002).
Wise, A., Gearing, K. & Rees, S. Target validation of G-protein coupled receptors. Drug Discov. Today 7, 235–246 (2002).
Waszkowycz, B. et al. PRO LIGAND: an approach to de novo molecular design. 2. design of novel molecules from molecular field analysis (MFA) models and pharmacophores. J. Med. Chem. 37, 3994–4002 (1994).
Nachbar, R. B. Molecular evolution: automated manipulation of hierarchical chemical topology and its application to average molecular structures. Genet. Programming Evolvable Machines 1, 57–94 (2000). Development of genetic operators for graph-based structure sampling and detailed description of the problems that have to be solved.
Pellegrini, E. & Field, M. J. Development and testing of a de novo drug-design algorithm. J. Comp. Aided Mol. Des. 17, 621–641 (2003).
Douguet, D., Thoreau, E. & Grassy, G. A genetic algorithm for the automated generation of small organic molecules: drug design using an evolutionary algorithm. J. Comput. Aided Mol. Des. 14, 449–466 (2000).
Schneider, G., Lee, M. -L., Stahl, M. & Schneider, P. De novo design of molecular architectures by evolutionary assembly of drug-derived building blocks. J. Comput. Aided Mol. Des. 14, 487–494 (2000).
Globus, A., Lawton, J. & Wipke, W. T. Automatic Molecular design using evolutionary algorithms. Nanotechnology 10, 290–299 (1999).
Brown, N., McKay, B., Gilardoni, F. & Gasteiger, J. A graph-based genetic algorithm and its application to the multiobjective evolution of median molecules. J. Chem. Inf. Comput. Sci. 44, 1079–1087 (2004).
Lipinski, C. et al. Experimental and computational approaches to estimate solubility and permeability in drug discovery and development settings. Adv. Drug. Deliv. Rev. 23, 3–25 (1997).
Teague, S. J. et al. The design of leadlike combinatorial libraries. Angew. Chem. Int. Ed. Engl. 38, 3743–3747 (1999).
Aronov, A. M. Predictive in silico modeling for hERG channel blockers. Drug Discov. Today 10, 149–155 (2005).
Lewell, X. O., Budd, D. B., Watson, S. P. & Hann, M. M. RECAP – Retrosynthetic Combinatorial Analysis Procedure: a powerful new technique for identifying privileged molecular fragments with useful applications in combinatorial chemistry. J. Chem. Inf. Comput. Sci. 38, 511–522 (1998).
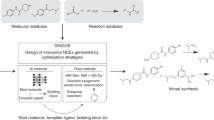
Vinkers, H. M. et al. SYNOPSIS: SYNthesize and OPtimize System in Silico. J. Med. Chem. 46, 2765–2773 (2003).
Honma, T. et al. Structure-based generation of a new class of potent Cdk4 inhibitors: new de novo design strategy and library design. J. Med. Chem. 44, 4615–4627 (2001).
Gillett, V., Johnson, P., Mata, P., Sike, S. & Williams, P. SPROUT: a program for structure generation. J. Comput. Aided Mol. Des. 7, 127–153 (1993).
Gillet, V. et al. P. SPROUT: recent developments in the de novo design of molecules. J. Chem. Inf. Comput. Sci. 34, 207–217 (1994).
Mata, P. et al. SPROUT: 3D structure generation using templates. J. Chem. Inf. Comput. Sci. 35, 479–493 (1995).
Ho, C. M. W. & Marshall, G. R. SPLICE: a program to assemble partial query solutions from three-dimensional database searches into novel ligands. J. Comput. Aided Mol. Des. 7, 623–647 (1993).
Gelhaar, D. K. et al. De novo design of enzyme inhibitors by monte carlo ligand generation. J. Med. Chem. 38, 466–472 (1995).
Pierce, A. C., Rao, G. & Bemis, G. W. BREED: generating novel inhibitors through hybridization of known ligands. application to CDK2, P38, and HIV protease. J. Med. Chem. 47, 2768–2775 (2004).
Todorov, N. P. & Dean, P. M. Evaluation of a method for controlling molecular scaffold diversity in de novo ligand design. J. Comput. Aided. Mol. Des. 11, 175–192 (1997).
Todorov, N. P. & Dean, P. M. A branch-and-bound method for optimal atom-type assignment in de novo ligand design. J. Comput. Aided. Mol. Des. 12, 335–350 (1998).
Darwin, C. On the Origin of Species (Facsimile of the First Edition) (Harvard Univ. Press, Cambridge, Massachusetts, 1859/1975).
Weininger, D. SMILES, a chemical language and information system. 1. Introduction to methodology and encoding rules. J. Chem. Inf. Comput. Sci. 28, 31–36 (1988).
Pegg, S. C. -H., Haresco, J. J. & Kuntz, I. D. A genetic algorithm for structure-based de novo design. J. Comput. Aided Mol. Des. 15, 911–933 (2001).
Schneider, G. & Böhm, H. -J. Virtual screening and fast automated docking methods. Drug Discov. Today 7, 64–70 (2002).
Hou, T. & Xu, X. Recent development and application of virtual screening in drug discovery: an overview. Curr. Pharm. Des. 10, 1011–1033 (2004).
Honma, T. Recent advances in de novo design strategy for practical lead identification. Med. Res. Rev. 23, 606–632 (2003).
Ji, H. et al. Structure-based de novo design, synthesis, and biological evaluation of non-azole inhibitors specific for lanosterol 14α-demethylase of fungi. J. Med. Chem. 46, 474–485 (2003).
Perola, E., Walters, W. P. & Charifson, P. S. A detailed comparison of current docking and scoring methods on systems of pharmaceutical relevance. Proteins 56, 235–249 (2004).
Schuffenhauer, A. et al. Molecular diversity management strategies for building and enhancement of diverse and focused lead discovery compound screening collections. Comb. Chem. High Throughput Screen. 7, 771–781 (2004).
Honma, T. et al. A novel approach for the development of selective Cdk4 inhibitors: library design based on locations of Cdk4 specific amino acid residues. J. Med. Chem. 44, 4628–4640 (2001).
Rogers-Evans, M., Alanine, A. I., Bleicher, K. H., Kube, D. & Schneider, G. Identification of novel cannabinoid receptor ligands via evolutionary de novo design and rapid parallel synthesis. QSAR Comb. Sci. 23, 426–430 (2004).
Böhm, H. -J., Banner, D. W. & Weber, L. Combinatorial docking and combinatorial chemistry: design of potent non-peptide thrombin inhibitors. J. Comput. Aided Mol. Des. 13, 51–56 (1999).
Obst, U., Banner, D. W., Weber, L. & Diederich, F. Molecular recognition at the thrombin active site: structure-based design and synthesis of potent and selective thrombin inhibitors and the X-ray crystal structures of two thrombin-inhibitor complexes. Chem. Biol. 4, 287–295 (1997).
Olsen, J. A. et al. A fluorine scan of thrombin inhibitors to map the fluorophilicity/fluorophobicity of an enzyme active site: evidence for C–F...C=O interactions. Angew. Chem. Int. Ed. Eng. 42, 2507–2511 (2003).
Gribbon, P. & Sewing A. High-throughput drug discovery: what can we expect from HTS? Drug Discov. Today 10, 17–22 (2005).
Anderson, A. C. & Wright, D. L. The design and docking of virtual compound libraries to structures of drug targets. Curr. Comp. Aided Drug Des. 1, 103–127 (2005). An excellent overview of current developments in molecular docking and scoring and its relation to de novo design.
Doweyko, A. M. 3D-QSAR illusions. J. Comp. Aided Mol. Des. 18, 587–596 (2004).
Bemis, G. W. & Murcko, M. A. The properties of known drugs. 1. Molecular frameworks. J. Med. Chem. 39, 2887–2893 (1996).
Müller, G. in Chemogenomics in Drug Discovery (eds Kubinyi, H. & Müller, G.) 7–41 (Wiley-VCH, Weinheim, 2004).
Jenkins, J. L., Glick, M. & Davies, J. W. A 3D similarity method for scaffold hopping from known drugs or natural ligands to new chemotypes. J. Med. Chem. 47, 6144–6159 (2004).
Bailey, D. & Brown, D. High-throughput chemistry and structure-based design: survival of the smartest. Drug Discov. Today 6, 57–59 (2001).
Verdonk, M. L. & Hartshorn, M. J. Structure-guided fragment screening for lead discovery. Curr. Opin. Drug Discov. Devel. 7, 404–410 (2004).
Villar, H. O., Yan, J. & Hansen, M. R. Using NMR for ligand discovery and optimization. Curr. Opin. Chem. Biol. 8, 387–391 (2004).
Gillet, V. J., Khatib, W., Willett, P., Fleming, P. J. & Green, D. V. S. Combinatorial library design using a multiobjective genetic algorithm. J. Chem. Inf. Comput. Sci. 42, 375–385 (2002).
Fonseca, C. M. & Fleming, P. J. in Genetic Algorithms: Proceedings of the Fifth International Conference (ed. Forrest, S.) 416–423 (Morgan Kaufmann: San Mateo, CA, 1993).
Handschuh, S., Wagener, M. & Gasteiger, J. Superposition of three-dimensional chemical structures allowing for conformational flexibility by a hybrid method. J. Chem. Inf. Comput. Sci. 38, 220–232 (1998).
Agrafiotis, D. K. Multiobjective optimisation of combinatorial libraries. IBM J. Res. DeV. 45, 545–566 (2001).
Wright, T., Gillet, V. J., Green, D. V. S. & Pickett, S. D. Optimizing the size and configuration of combinatorial libraries. J. Chem. Inf. Comput. Sci. 43, 381–390 (2003).
Babine, R. E. et al. Design, synthesis and X-ray crystallographic studies of novel FKBB-12 ligands. Bioorg. Med. Chem. Lett. 5, 1719–1724 (1995).
Schindler, T. et al. Structural mechanism of STI-571 inhibition of Abelson tyrosine kinase. Science 289, 1938–1942 (2000).
Lewis, R. A. & Dean, P. M. Automated site-directed drug design: the concept of spacer skeletons for primary structure generation. Proc R Soc Lond B Biol Sci B 236, 125–140 (1989). Pioneering theoretical outline to tackle the problem of automated drug design from first principles.
Böhm, H. -J. A novel computational tool for automated structure-based drug design. J. Mol. Recognit. 6, 131–137 (1993). Concise overview of the early developments of the program LUDI.
Böhm, H. -J. The development of a simple empirical scoring function to estimate the binding constant for a protein-ligand complex of known three-dimensional structure. J. Comput. Aided Mol. Des. 8, 243–256 (1994).
Böhm, H. -J. Prediction of binding constants of protein ligands: a fast method for the prioritization of hits obtained from de novo design or 3D database search programs. J. Comput. Aided Mol. Des. 12, 309–323 (1998).
Stultz, C. M. & Karplus, M. Dynamic ligand design and combinatorial optimization: designing inhibitors to endothiapepsin. Proteins 40, 258–289 (2000).
Westhead, D. R. et al. PRO LIGAND: an approach to de novo molecular design. 3. A genetic algorithm for structure refinement. J. Comput. Aided Mol. Des. 9, 139–148 (1995).
Frenkel, D. et al. PRO LIGAND: an approach de novo molecular design. 4. Application to the design of peptides. J. Comput. Aided Mol. Des. 9, 213–225 (1995).
Clark, D. E. & Murray, C. W. PRO LIGAND: an approach to de novo molecular design. 5. Tools for the Analysis of Generated Structures. J. Chem. Inf. Comput. Sci. 35, 914–923 (1995).
Murray, C. W., Clark, D. E., Byrne, D. G. PRO LIGAND: an approach to de novo molecular design. 6. Flexible fitting in the design of peptides. J. Comput. Aided Mol. Des. 9, 381–395 (1995).
Grzybowski, B. A. et al. Combinatorial computational method gives new picomolar ligands for a known enzyme. Proc. Natl Acad. Sci. USA 99, 1270–1273 (2002). Design of a picomolar human carbonic anhydrase II inhibitor, the highest-affinity inhibitor to date, with the program SMoG.
Nachbar, R. B. Molecular evolution: a hierarchical representation for chemical topology and its automated manipulation. Proc. 3rd Ann. Genetic Programming Conf. 246–253 (Univ. of Wisconsin, Madison, Wisconsin 1998).
Wermuth, C. G., Gannelin, C. R., Lindberg, P. and Mitscher, L. A. Glossary of terms used in medicinal chemistry. Pure Appl. Chem. 70, 1129–1143 (1998).
Acknowledgements
H. Kubinyi is thanked for helpful discussion and kind support. This work was supported by the Beilstein-Institut zur Förderung der Chemischen Wissenschaften, Frankfurt am Main. U.F. is thankful for a fellowship granted by Aventis Pharma Deutschland GmbH, a company of the Sanofi-Aventis group.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Related links
Glossary
- DE NOVO DESIGN
-
The design of bioactive compounds by incremental construction of a ligand model within a model of the receptor or enzyme active site, the structure of which is known from X-ray or NMR data106.
- SCAFFOLD HOPPING
-
The identification of isofunctional structures with different backbone architectures.
- PRIMARY TARGET CONSTRAINTS
-
All information that is related to the ligand–receptor interaction — that is, the binding affinity of a ligand to the particular biological target.
- INTERACTION SITE
-
A position in space that is not occupied by the receptor and in which a ligand atom favourably interacts with the receptor.
- PHARMACOPHORE
-
The ensemble of steric and electronic features that is necessary to ensure the optimal supramolecular interactions with a specific biological target structure and to trigger (or to block) its biological response106.
- QUANTITATIVE STRUCTURE–ACTIVITY RELATIONSHIPS
-
(QSAR). Mathematical relationships linking chemical structure and pharmacological activity in a quantitative manner for a series of compounds. Methods that can be used in QSAR include various regression and pattern-recognition techniques106.
- NP-HARD
-
Non-deterministic polynomial-time hard (NP-hard) refers to a class of decision problems of which current knowledge provides no way to obtain or derive a solution time that is less than exponential in problem size.
- HEURISTIC
-
Application of probabilistic rules grounded on knowledge of a particular problem domain to obtain an algorithm that performs 'reasonably well' in many cases, but without proof that it is always fast.
- SECONDARY TARGET CONSTRAINTS
-
Essential drug properties apart from the binding affinity to a biological target — for example, absorption, distribution, metabolism, excretion and toxicity properties, or binding selectivity.
Rights and permissions
About this article
Cite this article
Schneider, G., Fechner, U. Computer-based de novo design of drug-like molecules. Nat Rev Drug Discov 4, 649–663 (2005). https://doi.org/10.1038/nrd1799
Issue Date:
DOI: https://doi.org/10.1038/nrd1799
This article is cited by
-
MedGAN: optimized generative adversarial network with graph convolutional networks for novel molecule design
Scientific Reports (2024)
-
Integrating QSAR modelling and deep learning in drug discovery: the emergence of deep QSAR
Nature Reviews Drug Discovery (2024)
-
A molecule perturbation software library and its application to study the effects of molecular design constraints
Journal of Cheminformatics (2023)
-
Designing molecules with autoencoder networks
Nature Computational Science (2023)
-
Leveraging molecular structure and bioactivity with chemical language models for de novo drug design
Nature Communications (2023)