Key Points
-
Induced protein degradation is an emerging drug discovery platform with the potential to reduce drug exposure requirements, counteract compensatory protein expression and target the 'undruggable proteome'.
-
Selective oestrogen receptor degraders (such as fulvestrant) demonstrated some of the advantages of induced protein degradation over the traditional inhibitor-based approach.
-
Hydrophobic tagging attempts to mimic partially unfolded proteins to induce target protein degradation, although the exact mechanism of action remains unknown.
-
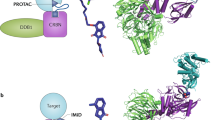
Proteolysis-targeting chimaeras (PROTACs) are a promising platform technology that is modular, catalytic and specific for its protein target with picomolar potencies in cell culture and demonstrated activity in mouse models.
-
An added layer of specificity over the traditional inhibition approach can be accomplished with PROTACs by utilizing different E3 ligases and conditional recruiting ligands.
-
Future development of targeted protein degradation will be focused on targeting the undruggable proteome.
Abstract
Small-molecule drug discovery has traditionally focused on occupancy of a binding site that directly affects protein function, and this approach typically precludes targeting proteins that lack such amenable sites. Furthermore, high systemic drug exposures may be needed to maintain sufficient target inhibition in vivo, increasing the risk of undesirable off-target effects. Induced protein degradation is an alternative approach that is event-driven: upon drug binding, the target protein is tagged for elimination. Emerging technologies based on proteolysis-targeting chimaeras (PROTACs) that exploit cellular quality control machinery to selectively degrade target proteins are attracting considerable attention in the pharmaceutical industry owing to the advantages they could offer over traditional small-molecule strategies. These advantages include the potential to reduce systemic drug exposure, the ability to counteract increased target protein expression that often accompanies inhibition of protein function and the potential ability to target proteins that are not currently therapeutically tractable, such as transcription factors, scaffolding and regulatory proteins.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Campbell, J. et al. Large-scale profiling of kinase dependencies in cancer cell lines. Cell Rep. 14, 2490–2501 (2016).
Cowley, G. S. et al. Parallel genome-scale loss of function screens in 216 cancer cell lines for the identification of context-specific genetic dependencies. Sci. Data 1, 140035 (2014).
Wang, T. et al. Identification and characterization of essential genes in the human genome. Science 350, 1096–1101 (2015).
Barretina, J. et al. The Cancer Cell Line Encyclopedia enables predictive modelling of anticancer drug sensitivity. Nature 483, 603–607 (2012).
Hopkins, A. L. & Groom, C. R. The druggable genome. Nat. Rev. Drug Discov. 1, 727–730 (2002).
Lazo, J. S. & Sharlow, E. R. Drugging undruggable molecular cancer targets. Annu. Rev. Pharmacol. Toxicol. 56, 23–40 (2016).
Jin, L., Wang, W. & Fang, G. Targeting protein–protein interaction by small molecules. Annu. Rev. Pharmacol. Toxicol. 54, 435–456 (2014).
Adjei, A. A. What is the right dose? The elusive optimal biologic dose in phase I clinical trials. J. Clin. Oncol. 24, 4054–4055 (2006).
Cong, L. et al. Multiplex genome engineering using CRISPR/Cas systems. Science 339, 819–823 (2013).
Jinek, M. et al. A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science 337, 816–821 (2012).
Doudna, J. A. & Charpentier, E. The new frontier of genome engineering with CRISPR–Cas9. Science 346, 1258096 (2014).
Hsu, P. D., Lander, E. S. & Zhang, F. Development and applications of CRISPR–Cas9 for genome engineering. Cell 157, 1262–1278 (2014).
Sander, J. D. & Joung, J. K. CRISPR–Cas systems for editing, regulating and targeting genomes. Nat. Biotechnol. 32, 347–355 (2014).
de Smidt, P. C., Le Doan, T., de Falco, S. & van Berkel, T. J. Association of antisense oligonucleotides with lipoproteins prolongs the plasma half-life and modifies the tissue distribution. Nucleic Acids Res. 19, 4695–4700 (1991).
Geary, R. S. et al. Pharmacokinetic properties of 2′-O-(2-methoxyethyl)-modified oligonucleotide analogs in rats. J. Pharmacol. Exp. Ther. 296, 890–897 (2001).
McMahon, B. M. et al. Pharmacokinetics and tissue distribution of a peptide nucleic acid after intravenous administration. Antisense Nucleic Acid Drug Dev. 12, 65–70 (2002).
Marques, J. T. & Williams, B. R. Activation of the mammalian immune system by siRNAs. Nat. Biotechnol. 23, 1399–1405 (2005).
Krieg, A. M. CpG motifs in bacterial DNA and their immune effects. Annu. Rev. Immunol. 20, 709–760 (2002).
Dahlman, J. E., Kauffman, K. J., Langer, R. & Anderson, D. G. Nanotechnology for in vivo targeted siRNA delivery. Adv. Genet. 88, 37–69 (2014).
Whitehead, K. A., Langer, R. & Anderson, D. G. Knocking down barriers: advances in siRNA delivery. Nat. Rev. Drug Discov. 8, 129–138 (2009).
Wilhelm, S. et al. Analysis of nanoparticle delivery to tumours. Nat. Rev. Mater. 1, 16014 (2016).
Peer, D. & Lieberman, J. Special delivery: targeted therapy with small RNAs. Gene Ther. 18, 1127–1133 (2011).
Fedorov, Y. et al. Off-target effects by siRNA can induce toxic phenotype. RNA 12, 1188–1196 (2006).
Jackson, A. L. et al. Expression profiling reveals off-target gene regulation by RNAi. Nat. Biotechnol. 21, 635–637 (2003).
Qiu, S., Adema, C. M. & Lane, T. A computational study of off-target effects of RNA interference. Nucleic Acids Res. 33, 1834–1847 (2005).
Burnett, J. C. & Rossi, J. J. RNA-based therapeutics: current progress and future prospects. Chem. Biol. 19, 60–71 (2012).
Melnikova, I. RNA-based therapies. Nat. Rev. Drug Discov. 6, 863–864 (2007).
Bennett, C. F. & Swayze, E. E. RNA targeting therapeutics: molecular mechanisms of antisense oligonucleotides as a therapeutic platform. Annu. Rev. Pharmacol. Toxicol. 50, 259–293 (2010).
Perry, C. M. & Balfour, J. A. B. Fomivirsen. Drugs 57, 375–380 (1999).
Wong, E. & Goldberg, T. Mipomersen (Kynamro): a novel antisense oligonucleotide inhibitor for the management of homozygous familial hypercholesterolemia. Pharmacy Ther. 39, 119–122 (2014).
Sinha, G. Antisense battles small molecule for slice of rare lipid disorder market. Nat. Biotechnol. 31, 179–180 (2013).
Crews, C. M. Targeting the undruggable proteome: the small molecules of my dreams. Chem. Biol. 17, 551–555 (2010).
Overington, J. P., Al-Lazikani, B. & Hopkins, A. L. How many drug targets are there? Nat. Rev. Drug Discov. 5, 993–996 (2006).
Duncan, J. S. et al. Dynamic reprogramming of the kinome in response to targeted MEK inhibition in triple-negative breast cancer. Cell 149, 307–321 (2012).
Visakorpi, T. et al. In vivo amplification of the androgen receptor gene and progression of human prostate cancer. Nat. Genet. 9, 401–406 (1995).
Hatzivassiliou, G. et al. RAF inhibitors prime wild-type RAF to activate the MAPK pathway and enhance growth. Nature 464, 431–435 (2010).
Heidorn, S. J. et al. Kinase-dead BRAF and oncogenic RAS cooperate to drive tumor progression through CRAF. Cell 140, 209–221 (2010).
Poulikakos, P. I., Zhang, C., Bollag, G., Shokat, K. M. & Rosen, N. RAF inhibitors transactivate RAF dimers and ERK signalling in cells with wild-type BRAF. Nature 464, 427–430 (2010).
Rauch, J., Volinsky, N., Romano, D. & Kolch, W. The secret life of kinases: functions beyond catalysis. Cell Commun. Signal. 9, 23 (2011). An excellent review of kinase-independent functions that are unlikely to be affected by traditional kinase inhibitors but would be amendable to protein degradation.
Tan, X., Thapa, N., Sun, Y. & Anderson, R. A. A kinase-independent role for EGF receptor in autophagy initiation. Cell 160, 145–160 (2015).
Vivanco, I. et al. A kinase-independent function of AKT promotes cancer cell survival. eLife 3, e03751 (2014).
Weihua, Z. et al. Survival of cancer cells is maintained by EGFR independent of its kinase activity. Cancer Cell 13, 385–393 (2008).
Moulick, K. et al. Affinity-based proteomics reveal cancer-specific networks coordinated by Hsp90. Nat. Chem. Biol. 7, 818–826 (2011).
Kamal, A. et al. A high-affinity conformation of Hsp90 confers tumour selectivity on Hsp90 inhibitors. Nature 425, 407–410 (2003).
Grenert, J. P. et al. The amino-terminal domain of heat shock protein 90 (hsp90) that binds geldanamycin is an ATP/ADP switch domain that regulates hsp90 conformation. J. Biol. Chem. 272, 23843–23850 (1997).
Stebbins, C. E. et al. Crystal structure of an Hsp90–geldanamycin complex: targeting of a protein chaperone by an antitumor agent. Cell 89, 239–250 (1997).
Prodromou, C. et al. Identification and structural characterization of the ATP/ADP-binding site in the Hsp90 molecular chaperone. Cell 90, 65–75 (1997).
Schneider, C. et al. Pharmacologic shifting of a balance between protein refolding and degradation mediated by Hsp90. Proc. Natl Acad. Sci. USA 93, 14536–14541 (1996).
Mimnaugh, E. G., Chavany, C. & Neckers, L. Polyubiquitination and proteasomal degradation of the p185c-erbB-2 receptor protein-tyrosine kinase induced by geldanamycin. J. Biol. Chem. 271, 22796–22801 (1996).
Samuni, Y. et al. Reactive oxygen species mediate hepatotoxicity induced by the Hsp90 inhibitor geldanamycin and its analogs. Free Radic. Biol. Med. 48, 1559–1563 (2010).
Banerji, U. et al. Phase I pharmacokinetic and pharmacodynamic study of 17-allylamino, 17-demethoxygeldanamycin in patients with advanced malignancies. J. Clin. Oncol. 23, 4152–4161 (2005).
Taldone, T., Gozman, A., Maharaj, R. & Chiosis, G. Targeting Hsp90: small-molecule inhibitors and their clinical development. Curr. Opin. Pharmacol. 8, 370–374 (2008).
McClellan, A. J. et al. Diverse cellular functions of the Hsp90 molecular chaperone uncovered using systems approaches. Cell 131, 121–135 (2007).
Taipale, M., Jarosz, D. F. & Lindquist, S. HSP90 at the hub of protein homeostasis: emerging mechanistic insights. Nat. Rev. Mol. Cell Biol. 11, 515–528 (2010).
Taipale, M. et al. Quantitative analysis of Hsp90-client interactions reveals principles of substrate recognition. Cell 150, 987–1001 (2012).
Kuduk, S. D. et al. Synthesis and evaluation of geldanamycin–testosterone hybrids. Bioorg. Med. Chem. Lett. 10, 1303–1306 (2000).
Kuduk, S. D., Zheng, F. F., Sepp-Lorenzino, L., Rosen, N. & Danishefsky, S. J. Synthesis and evaluation of geldanamycin–estradiol hybrids. Bioorg. Med. Chem. Lett. 9, 1233–1238 (1999).
Katerina, S. & Evangelia, P. HSP90 Inhibitors: current development and potential in cancer therapy. Recent Pat. Anticancer Drug Discov. 9, 1–20 (2014).
Jego, G., Hazoume, A., Seigneuric, R. & Garrido, C. Targeting heat shock proteins in cancer. Cancer Lett. 332, 275–285 (2013).
Trepel, J., Mollapour, M., Giaccone, G. & Neckers, L. Targeting the dynamic HSP90 complex in cancer. Nat. Rev. Cancer 10, 537–549 (2010).
Dauvois, S., Danielian, P. S., White, R. & Parker, M. G. Antiestrogen ICI 164,384 reduces cellular estrogen receptor content by increasing its turnover. Proc. Natl Acad. Sci. USA 89, 4037–4041 (1992). The study illustrates the seminal finding that fulvestrant (ICI 164,384) induces the degradation of the ER.
Giannetti, A. M. From experimental design to validated hits a comprehensive walk-through of fragment lead identification using surface plasmon resonance. Methods Enzymol. 493, 169–218 (2011).
Shuker, S. B., Hajduk, P. J., Meadows, R. P. & Fesik, S. W. Discovering high-affinity ligands for proteins: SAR by NMR. Science 274, 1531–1534 (1996).
Mashalidis, E. H., Sledz, P., Lang, S. & Abell, C. A three-stage biophysical screening cascade for fragment-based drug discovery. Nat. Protoc. 8, 2309–2324 (2013).
Erlanson, D. A., Fesik, S. W., Hubbard, R. E., Jahnke, W. & Jhoti, H. Twenty years on: the impact of fragments on drug discovery. Nat. Rev. Drug Discov. 15, 605–619 (2016).
Renaud, J.-P. et al. Biophysics in drug discovery: impact, challenges and opportunities. Nat. Rev. Drug Discov. 15, 679–698 (2016).
Bleicher, K. H., Bohm, H.-J., Muller, K. & Alanine, A. I. Hit and lead generation: beyond high-throughput screening. Nat. Rev. Drug Discov. 2, 369–378 (2003).
Macarron, R. et al. Impact of high-throughput screening in biomedical research. Nat. Rev. Drug Discov. 10, 188–195 (2011).
Kola, I. & Landis, J. Can the pharmaceutical industry reduce attrition rates? Nat. Rev. Drug Discov. 3, 711–716 (2004).
Bondeson, D. P. et al. Catalytic in vivo protein knockdown by small-molecule PROTACs. Nat. Chem. Biol. 11, 611–617 (2015). This paper demonstrates the first all-small-molecule VHL-based PROTAC capable of degrading ERRα and RIPK2, and it shows that PROTACs remain specific for their targets and are active in vivo.
Lai, A. C. et al. Modular PROTAC design for the degradation of oncogenic BCR–ABL. Angew. Chem. Int. Ed. 55, 807–810 (2016). This study explores VHL- versus CRBN-based PROTACs targeting the oncogenic tyrosine kinase BCR–ABL.
Winter, G. E. et al. Phthalimide conjugation as a strategy for in vivo target protein degradation. Science 348, 1376–1381 (2015).
Heldring, N. et al. Estrogen receptors: how do they signal and what are their targets. Physiol. Rev. 87, 905–931 (2007).
Nilsson, S. et al. Mechanisms of estrogen action. Physiol. Rev. 81, 1535–1565 (2001).
Jordan, V. C. Tamoxifen: a most unlikely pioneering medicine. Nat. Rev. Drug Discov. 2, 205–213 (2003).
Early Breast Cancer Trialists' Collaborative Group. Tamoxifen for early breast cancer: an overview of the randomised trials. Lancet 351, 1451–1467 (1998).
Martin, L. A. et al. Enhanced estrogen receptor (ER) α, ERBB2, and MAPK signal transduction pathways operate during the adaptation of MCF-7 cells to long term estrogen deprivation. J. Biol. Chem. 278, 30458–30468 (2003).
Wakeling, A. E., Dukes, M. & Bowler, J. A potent specific pure antiestrogen with clinical potential. Cancer Res. 51, 3867–3873 (1991).
Buzdar, A. U. & Robertson, J. F. Fulvestrant: pharmacologic profile versus existing endocrine agents for the treatment of breast cancer. Ann. Pharmacother. 40, 1572–1583 (2006).
Wu, Y. L. et al. Structural basis for an unexpected mode of SERM-mediated ER antagonism. Mol. Cell 18, 413–424 (2005). This paper proposes a mechanism of action for SERDs by suggesting that increased surface hydrophobicity upon drug binding leads to the degradation of the ER.
Wittmann, B. M., Sherk, A. & McDonnell, D. P. Definition of functionally important mechanistic differences among selective estrogen receptor down-regulators. Cancer Res. 67, 9549–9560 (2007).
Kato, S. et al. Activation of the estrogen receptor through phosphorylation by mitogen-activated protein kinase. Science 270, 1491–1494 (1995).
Connor, C. E. et al. Circumventing tamoxifen resistance in breast cancers using antiestrogens that induce unique conformational changes in the estrogen receptor. Cancer Res. 61, 2917–2922 (2001).
Osborne, C. K., Wakeling, A. & Nicholson, R. I. Fulvestrant: an oestrogen receptor antagonist with a novel mechanism of action. Br. J. Cancer 90, S2–S6 (2004).
Vergote, I. & Abram, P. Fulvestrant, a new treatment option for advanced breast cancer: tolerability versus existing agents. Ann. Oncol. 17, 200–204 (2006).
Di Leo, A. et al. Results of the CONFIRM phase III trial comparing fulvestrant 250 mg with fulvestrant 500 mg in postmenopausal women with estrogen receptor-positive advanced breast cancer. J. Clin. Oncol. 28, 4594–4600 (2010).
van Kruchten, M. et al. Measuring residual estrogen receptor availability during fulvestrant therapy in patients with metastatic breast cancer. Cancer Discov. 5, 72–81 (2015).
Bross, P. F. et al. Fulvestrant in postmenopausal women with advanced breast cancer. Clin. Cancer Res. 9, 4309–4317 (2003).
Govek, S. P. et al. Optimization of an indazole series of selective estrogen receptor degraders: tumor regression in a tamoxifen-resistant breast cancer xenograft. Bioorg. Med. Chem. Lett. 25, 5163–5167 (2015).
Lai, A. et al. Identification of GDC-0810 (ARN-810), an orally bioavailable selective estrogen receptor degrader (SERD) that demonstrates robust activity in tamoxifen-resistant breast cancer xenografts. J. Med. Chem. 58, 4888–4904 (2015).
Garner, F., Shomali, M., Paquin, D., Lyttle, C. R. & Hattersley, G. RAD1901: a novel, orally bioavailable selective estrogen receptor degrader that demonstrates antitumor activity in breast cancer xenograft models. Anticancer Drugs 26, 948–956 (2015).
Weir, H. M. et al. AZD9496: an oral estrogen receptor inhibitor that blocks the growth of ER-positive and ESR1 mutant breast tumours in preclinical models. Cancer Res. http://dx.doi.org/10.1158/0008-5472.can-15-2357 (2016).
Matsumoto, T. et al. The androgen receptor in health and disease. Annu. Rev. Physiol. 75, 201–224 (2013).
Crawford, E. D. et al. A controlled trial of leuprolide with and without flutamide in prostatic carcinoma. N. Engl. J. Med. 321, 419–424 (1989).
Kolvenbag, G. J., Blackledge, G. R. & Gotting-Smith, K. Bicalutamide (Casodex) in the treatment of prostate cancer: history of clinical development. Prostate 34, 61–72 (1998).
Linja, M. J. et al. Amplification and overexpression of androgen receptor gene in hormone-refractory prostate cancer. Cancer Res. 61, 3550–3555 (2001).
Feldman, B. J. & Feldman, D. The development of androgen-independent prostate cancer. Nat. Rev. Cancer 1, 34–45 (2001).
Yap, T. A. et al. Drug discovery in advanced prostate cancer: translating biology into therapy. Nat. Rev. Drug Discov. 15, 699–718 (2016).
Scher, H. I. et al. Increased survival with enzalutamide in prostate cancer after chemotherapy. N. Engl. J. Med. 367, 1187–1197 (2012).
Bradbury, R. H. et al. Small-molecule androgen receptor downregulators as an approach to treatment of advanced prostate cancer. Bioorg. Med. Chem. Lett. 21, 5442–5445 (2011).
Bradbury, R. H. et al. Discovery of AZD3514, a small-molecule androgen receptor downregulator for treatment of advanced prostate cancer. Bioorg. Med. Chem. Lett. 23, 1945–1948 (2013).
Loddick, S. A. et al. AZD3514: a small molecule that modulates androgen receptor signaling and function in vitro and in vivo. Mol. Cancer Ther. 12, 1715–1727 (2013).
Omlin, A. et al. AZD3514, an oral selective androgen receptor down-regulator in patients with castration-resistant prostate cancer — results of two parallel first-in-human phase I studies. Invest. New Drugs 33, 679–690 (2015).
Williams, R. Discontinued drugs in 2012: oncology drugs. Expert Opin. Investig. Drugs 22, 1627–1644 (2013).
Li, H. et al. Characterization of a new class of androgen receptor antagonists with potential therapeutic application in advanced prostate cancer. Mol. Cancer Ther. 12, 2425–2435 (2013).
Neklesa, T. K. & Crews, C. M. Chemical biology: greasy tags for protein removal. Nature 487, 308–309 (2012).
Long, M. J., Gollapalli, D. R. & Hedstrom, L. Inhibitor mediated protein degradation. Chem. Biol. 19, 629–637 (2012). This study describes the first demonstration of the Boc 3 Arg-based HyT technology used against GST and dihydrofolate reductase.
Coffey, R. T. et al. Ubiquilin-mediated small molecule inhibition of mammalian target of rapamycin complex 1 (mTORC1) signaling. J. Biol. Chem. 291, 5221–5233 (2016).
Kubota, H. Quality control against misfolded proteins in the cytosol: a network for cell survival. J. Biochem. 146, 609–616 (2009).
Neklesa, T. K. et al. Small-molecule hydrophobic tagging-induced degradation of HaloTag fusion proteins. Nat. Chem. Biol. 7, 538–543 (2011). The first paper to demonstrate cell culture-based and in vivo efficacy of adamantane-based HyTs for the degradation of HaloTag fusion proteins.
Los, G. V. & Wood, K. The HaloTag: a novel technology for cell imaging and protein analysis. Methods Mol. Biol. 356, 195–208 (2007).
Tae, H. S. et al. Identification of hydrophobic tags for the degradation of stabilized proteins. Chembiochem 13, 538–541 (2012).
Neklesa, T. K. et al. A bidirectional system for the dynamic small molecule control of intracellular fusion proteins. ACS Chem. Biol. 8, 2293–2300 (2013). In contrast to HyT, a ligand separate from the HaloTag was identified in this paper that stabilizes the protein, allowing for bidirectional control of intracellular HaloTag concentrations.
Xie, T. et al. Pharmacological targeting of the pseudokinase Her3. Nat. Chem. Biol. 10, 1006–1012 (2014). This study describes the first demonstration of targeted degradation of a previously undruggable protein by turning a protein ligand into a degrader compound.
Lim, S. M. et al. Development of small molecules targeting the pseudokinase Her3. Bioorg. Med. Chem. Lett. 25, 3382–3389 (2015).
Gustafson, J. L. et al. Small-molecule-mediated degradation of the androgen receptor through hydrophobic tagging. Angew. Chem. Int. Ed. 54, 9659–9662 (2015).
Teutsch, G. et al. Non-steroidal antiandrogens: synthesis and biological profile of high-affinity ligands for the androgen receptor. J. Steroid Biochem. Mol. Biol. 48, 111–119 (1994).
Sakamoto, K. M. et al. Protacs: chimeric molecules that target proteins to the Skp1-cullin-F-box complex for ubiquitination and degradation. Proc. Natl Acad. Sci. USA 98, 8554–8559 (2001). This paper introduces the PROTAC technology by demonstrating proximity-induced ubiquitylation and targeted protein degradation in a Xenopus egg extract.
Yaron, A. et al. Identification of the receptor component of the IκBα-ubiquitin ligase. Nature 396, 590–594 (1998).
Griffith, E. C. et al. Methionine aminopeptidase (type 2) is the common target for angiogenesis inhibitors AGM-1470 and ovalicin. Chem. Biol. 4, 461–471 (1997).
Sin, N. et al. The anti-angiogenic agent fumagillin covalently binds and inhibits the methionine aminopeptidase, MetAP-2. Proc. Natl Acad. Sci. USA 94, 6099–6103 (1997).
Sakamoto, K. M. et al. Development of Protacs to target cancer-promoting proteins for ubiquitination and degradation. Mol. Cell Proteomics 2, 1350–1358 (2003).
Hon, W. C. et al. Structural basis for the recognition of hydroxyproline in HIF-1α by pVHL. Nature 417, 975–978 (2002).
Min, J. H. et al. Structure of an HIF-1α -pVHL complex: hydroxyproline recognition in signaling. Science 296, 1886–1889 (2002).
Schneekloth, J. S. Jr. et al. Chemical genetic control of protein levels: selective in vivo targeted degradation. J. Am. Chem. Soc. 126, 3748–3754 (2004).
Bargagna-Mohan, P., Baek, S.-H., Lee, H., Kim, K. & Mohan, R. Use of PROTACS as molecular probes of angiogenesis. Bioorg. Med. Chem. Lett. 15, 2724–2727 (2005).
Rodriguez-Gonzalez, A. et al. Targeting steroid hormone receptors for ubiquitination and degradation in breast and prostate cancer. Oncogene 27, 7201–7211 (2008).
Zhang, D., Baek, S. H., Ho, A. & Kim, K. Degradation of target protein in living cells by small-molecule proteolysis inducer. Bioorg. Med. Chem. Lett. 14, 645–648 (2004).
Henning, R. K. et al. Degradation of Akt using protein-catalyzed capture agents. J. Pept. Sci. 22, 196–200 (2016).
Chu, T.-T. et al. Specific knockdown of endogenous tau protein by peptide-directed ubiquitin-proteasome degradation. Cell Chem. Biol. 23, 453–461 (2016).
Hines, J., Gough, J. D., Corson, T. W. & Crews, C. M. Posttranslational protein knockdown coupled to receptor tyrosine kinase activation with phosphoPROTACs. Proc. Natl Acad. Sci. USA 110, 8942–8947 (2013). By coupling the activation of receptor tyrosine kinases to the degradation of downstream effector proteins, this paper introduces the concept of conditional protein knockdown through PROTAC technology.
Cong, F., Zhang, J., Pao, W., Zhou, P. & Varmus, H. A protein knockdown strategy to study the function of β-catenin in tumorigenesis. BMC Mol. Biol. 4, 10 (2003).
Vassilev, L. T. et al. In vivo activation of the p53 pathway by small-molecule antagonists of MDM2. Science 303, 844–848 (2004).
Schneekloth, A. R., Pucheault, M., Tae, H. S. & Crews, C. M. Targeted intracellular protein degradation induced by a small molecule: en route to chemical proteomics. Bioorg. Med. Chem. Lett. 18, 5904–5908 (2008). The first all-small-molecule PROTAC is introduced in this paper by recruiting the E3 ligase MDM2 to degrade the AR.
Ding, Q. et al. Discovery of RG7388, a potent and selective p53–MDM2 inhibitor in clinical development. J. Med. Chem. 56, 5979–5983 (2013).
Vu, B. et al. Discovery of RG7112: a small-molecule MDM2 inhibitor in clinical development. ACS Med. Chem. Lett. 4, 466–469 (2013).
Andreeff, M. et al. Results of the phase I trial of RG7112, a small-molecule MDM2 antagonist in leukemia. Clin. Cancer Res. 22, 868–876 (2016).
Itoh, Y., Ishikawa, M., Naito, M. & Hashimoto, Y. Protein knockdown using methyl bestatin–ligand hybrid molecules: design and synthesis of inducers of ubiquitination-mediated degradation of cellular retinoic acid-binding proteins. J. Am. Chem. Soc. 132, 5820–5826 (2010). The paper introduces the first cIAP-based PROTAC that uses bestatin-mediated recruitment of cIAP to subsequently degrade CRABPI and CRABPII.
Umezawa, H., Aoyagi, T., Suda, H., Hamada, M. & Takeuchi, T. Bestatin, an inhibitor of aminopeptidase B, produced by actinomycetes. J. Antibiot. 29, 97–99 (1976).
Orning, L., Krivi, G. & Fitzpatrick, F. A. Leukotriene A4 hydrolase. Inhibition by bestatin and intrinsic aminopeptidase activity establish its functional resemblance to metallohydrolase enzymes. J. Biol. Chem. 266, 1375–1378 (1991).
Sekine, K. et al. Small molecules destabilize cIAP1 by activating auto-ubiquitylation. J. Biol. Chem. 283, 8961–8968 (2008).
Itoh, Y. et al. Development of target protein-selective degradation inducer for protein knockdown. Bioorg. Med. Chem. 19, 3229–3241 (2011). This study describes an amide-type cIAP E3 ligase ligand that can recruit cIAP and also reduces autoubiquitylation of cIAP.
Itoh, Y. et al. Double protein knockdown of cIAP1 and CRABP-II using a hybrid molecule consisting of ATRA and IAPs antagonist. Bioorg. Med. Chem. Lett. 22, 4453–4457 (2012).
Demizu, Y. et al. Design and synthesis of estrogen receptor degradation inducer based on a protein knockdown strategy. Bioorg. Med. Chem. Lett. 22, 1793–1796 (2012).
Itoh, Y., Kitaguchi, R., Ishikawa, M., Naito, M. & Hashimoto, Y. Design, synthesis and biological evaluation of nuclear receptor-degradation inducers. Bioorg. Med. Chem. 19, 6768–6778 (2011).
Ohoka, N. et al. Cancer cell death induced by novel small molecules degrading the TACC3 protein via the ubiquitin–proteasome pathway. Cell Death Dis. 5, e1513 (2014).
Buckley, D. L. et al. Targeting the von Hippel–Lindau E3 ubiquitin ligase using small molecules to disrupt the VHL/HIF-1α interaction. J. Am. Chem. Soc. 134, 4465–4468 (2012).
Buckley, D. L. et al. Small-molecule inhibitors of the interaction between the E3 ligase VHL and HIF1α. Angew. Chem. Int. Ed. 51, 11463–11467 (2012).
Van Molle, I. et al. Dissecting fragment-based lead discovery at the von Hippel–Lindau protein:hypoxia inducible factor 1α protein–protein interface. Chem. Biol. 19, 1300–1312 (2012).
Galdeano, C. et al. Structure-guided design and optimization of small molecules targeting the protein-protein interaction between the von Hippel–Lindau (VHL) E3 ubiquitin ligase and the hypoxia inducible factor (HIF) alpha subunit with in vitro nanomolar affinities. J. Med. Chem. 57, 8657–8663 (2014).
Ito, T. et al. Identification of a primary target of thalidomide teratogenicity. Science 327, 1345–1350 (2010).
Kronke, J. et al. Lenalidomide causes selective degradation of IKZF1 and IKZF3 in multiple myeloma cells. Science 343, 301–305 (2014).
Lu, G. et al. The myeloma drug lenalidomide promotes the cereblon-dependent destruction of Ikaros proteins. Science 343, 305–309 (2014).
Gandhi, A. K. et al. Immunomodulatory agents lenalidomide and pomalidomide co-stimulate T cells by inducing degradation of T cell repressors Ikaros and Aiolos via modulation of the E3 ubiquitin ligase complex CRL4CRBN. Br. J. Haematol. 164, 811–821 (2014).
Fischer, E. S. et al. Structure of the DDB1-CRBN E3 ubiquitin ligase in complex with thalidomide. Nature 512, 49–53 (2014).
Kronke, J. et al. Lenalidomide induces ubiquitination and degradation of CK1α in del(5q) MDS. Nature 523, 183–188 (2015).
Petzold, G., Fischer, E. S. & Thoma, N. H. Structural basis of lenalidomide-induced CK1α degradation by the CRL4CRBN ubiquitin ligase. Nature 532, 127–130 (2016).
Adès, L. & Fenaux, P. Immunomodulating drugs in myelodysplastic syndromes. ASH Educ. Program Book 2011, 556–560 (2011).
Matyskiela, M. E. et al. A novel cereblon modulator recruits GSPT1 to the CRL4CRBN ubiquitin ligase. Nature 535, 252–257 (2016).
Lu, J. et al. Hijacking the E3 ubiquitin ligase cereblon to efficiently target BRD4. Chem. Biol. 22, 755–763 (2015). This paper demonstrates a small-molecule CRBN-based PROTAC capable of degrading BRD4 that is superior to the parent inhibitor OTX-15 in Burkitt's lymphoma cell lines.
Shimamura, T. et al. Efficacy of BET bromodomain inhibition in KRAS-mutant non-small cell lung cancer. Clin. Cancer Res. 19, 6183–6192 (2013).
Raina, K. et al. PROTAC-induced BET protein degradation as a therapy for castration-resistant prostate cancer. Proc. Natl Acad. Sci. USA 113, 7124–7129 (2016).
Zengerle, M., Chan, K.-H. & Ciulli, A. Selective small molecule induced degradation of the BET bromodomain protein BRD4. ACS Chem. Biol. 10, 1770–1777 (2015).
Golas, J. M. et al. SKI-606, a 4-anilino-3-quinolinecarbonitrile dual inhibitor of Src and Abl kinases, is a potent antiproliferative agent against chronic myelogenous leukemia cells in culture and causes regression of K562 xenografts in nude mice. Cancer Res. 63, 375–381 (2003).
O'Hare, T. et al. In vitro activity of Bcr–Abl inhibitors AMN107 and BMS-354825 against clinically relevant imatinib-resistant Abl kinase domain mutants. Cancer Res. 65, 4500–4505 (2005).
Duda, D. M. et al. Structural insights into NEDD8 activation of cullin-RING ligases: conformational control of conjugation. Cell 134, 995–1006 (2008).
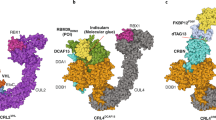
Fischer, E. S. et al. The molecular basis of CRL4DDB2/CSA ubiquitin ligase architecture, targeting, and activation. Cell 147, 1024–1039 (2011). This paper provides a structural rationale to the flexibility of cullin 4-based E3 ligases and explores the ubiquitylation zone of these ligases.
Liu, J. & Nussinov, R. Flexible cullins in cullin-RING E3 ligases allosterically regulate ubiquitination. J. Biol. Chem. 286, 40934–40942 (2011).
Liu, J. & Nussinov, R. The mechanism of ubiquitination in the cullin-RING E3 ligase machinery: conformational control of substrate orientation. PLoS Comput. Biol. 5, e1000527 (2009).
Petroski, M. D. & Deshaies, R. J. Function and regulation of cullin-RING ubiquitin ligases. Nat. Rev. Mol. Cell Biol. 6, 9–20 (2005).
Deshaies, R. J. Protein degradation: prime time for PROTACs. Nat. Chem. Biol. 11, 634–635 (2015).
Toure, M. & Crews, C. M. Small-molecule PROTACS: new approaches to protein degradation. Angew. Chem. Int. Ed. 55, 1966–1973 (2016).
Lou, K.-J. PROTAC the protein. SciBx http://dx.doi.org/10.1038/scibx.2012.514 (2012).
Bouchie, A., Allison, M., Webb, S. & DeFrancesco, L. Nature Biotechnology's academic spinouts of 2013. Nat. Biotechnol. 32, 229–238 (2014).
Chi, K. R. Drug developers delve into the cell's trash-disposal machinery. Nat. Rev. Drug Discov. 15, 295–297 (2016).
Buckley, D. L. & Crews, C. M. Small-molecule control of intracellular protein levels through modulation of the ubiquitin proteasome system. Angew. Chem. Int. Ed. 53, 2312–2330 (2014).
Buckley, D. L. et al. HaloPROTACS: use of small molecule PROTACs to induce degradation of halotag fusion proteins. ACS Chem. Biol. 10, 1831–1837 (2015).
Lipinski, C. A., Lombardo, F., Dominy, B. W. & Feeney, P. J. Experimental and computational approaches to estimate solubility and permeability in drug discovery and development settings. Adv. Drug Deliv. Rev. 46, 3–26 (2001).
Maculins, T. et al. A generic platform for cellular screening against ubiquitin ligases. Sci. Rep. 6, 18940 (2016). This study reports an interesting platform technology that could be used to find novel E3 ligase-recruiting ligands.
Chan, C. H. et al. Pharmacological inactivation of Skp2 SCF ubiquitin ligase restricts cancer stem cell traits and cancer progression. Cell 154, 556–568 (2013).
Orlicky, S. et al. An allosteric inhibitor of substrate recognition by the SCFCdc4 ubiquitin ligase. Nat. Biotechnol. 28, 733–737 (2010).
Aghajan, M. et al. Chemical genetics screen for enhancers of rapamycin identifies a specific inhibitor of an SCF family E3 ubiquitin ligase. Nat. Biotechnol. 28, 738–742 (2010).
Deshaies, R. J. & Joazeiro, C. A. P. RING domain E3 ubiquitin ligases. Annu. Rev. Biochem. 78, 399–434 (2009).
DeBoer, C., Meulman, P. A., Wnuk, R. J. & Peterson, D. H. Geldanamycin, a new antibiotic. J. Antibiot. 23, 442–447 (1970).
Rinehart, K. L., Sasaki, K., Slomp, G., Grostic, M. F. & Olson, E. C. Geldanamycin. I. Structure assignment. J. Am. Chem. Soc. 92, 7591–7593 (1970).
Supko, J. G., Hickman, R. L., Grever, M. R. & Malspeis, L. Preclinical pharmacologic evaluation of geldanamycin as an antitumor agent. Cancer Chemother. Pharmacol. 36, 305–315 (1995).
Ramanathan, R. K. et al. Phase I pharmacokinetic-pharmacodynamic study of 17-(allylamino)-17-demethoxygeldanamycin (17AAG, NSC 330507), a novel inhibitor of heat shock protein 90, in patients with refractory advanced cancers. Clin. Cancer Res. 11, 3385–3391 (2005).
Modi, S. et al. HSP90 inhibition is effective in breast cancer: a phase II trial of tanespimycin (17-AAG) plus trastuzumab in patients with HER2-positive metastatic breast cancer progressing on trastuzumab. Clin. Cancer Res. 17, 5132–5139 (2011).
Demetri, G. D. et al. Final results from a phase III study of IPI-504 (retaspimycin hydrochloride) versus placebo in patients (pts) with gastrointestinal stromal tumors (GIST) following failure of kinase inhibitor therapies. ASCO 2010 Gastrointestinal Cancers Symp. http://meetinglibrary.asco.org/content/2285-72 (2010).
Lundgren, K. et al. BIIB021, an orally available, fully synthetic small-molecule inhibitor of the heat shock protein Hsp90. Mol. Cancer Ther. 8, 921–929 (2009).
Dickson, M. A. et al. Phase II study of the HSP90-inhibitor BIIB021 in gastrointestinal stromal tumors. Ann. Oncol. 24, 252–257 (2013).
Mitchell, P. Biogen Idec restructures, sharpens neurology focus. Nat. Biotechnol. 29, 7–8 (2011).
Eccles, S. A. et al. NVP-AUY922: a novel heat shock protein 90 inhibitor active against xenograft tumor growth, angiogenesis, and metastasis. Cancer Res. 68, 2850–2860 (2008).
Lin, T. Y. et al. The novel HSP90 inhibitor STA-9090 exhibits activity against Kit-dependent and -independent malignant mast cell tumors. Exp. Hematol. 36, 1266–1277 (2008).
Ramalingam, S. et al. A randomized phase II study of ganetespib, a heat shock protein 90 inhibitor, in combination with docetaxel in second-line therapy of advanced non-small cell lung cancer (GALAXY-1). Ann. Oncol. 26, 1741–1748 (2015).
Chatterjee, S., Bhattacharya, S., Socinski, M. A. & Burns, T. F. HSP90 inhibitors in lung cancer: promise still unfulfilled. Clin. Adv. Hematol. Oncol. 14, 346–356 (2016).
Woodhead, A. J. et al. Discovery of (2,4-dihydroxy- 5-isopropylphenyl)-[5-(4-methylpiperazin-1-ylmethyl)-1,3-dihydroisoindol-2-yl]methanone (AT13387), a novel inhibitor of the molecular chaperone Hsp90 by fragment based drug design. J. Med. Chem. 53, 5956–5969 (2010).
Wagner, A. J. et al. Dose-escalation study of a second-generation non-ansamycin HSP90 inhibitor, onalespib (AT13387), in combination with imatinib in patients with metastatic gastrointestinal stromal tumour. Eur. J. Cancer 61, 94–101 (2016).
Rakhit, R., Navarro, R. & Wandless, T. J. Chemical biology strategies for posttranslational control of protein function. Chem. Biol. 21, 1238–1252 (2014).
Raina, K. et al. Targeted protein destabilization reveals an estrogen-mediated ER stress response. Nat. Chem. Biol. 10, 957–962 (2014).
Lytton, J., Westlin, M. & Hanley, M. R. Thapsigargin inhibits the sarcoplasmic or endoplasmic reticulum Ca-ATPase family of calcium pumps. J. Biol. Chem. 266, 17067–17071 (1991).
Takatsuki, A., Arima, K. & Tamura, G. Tunicamycin, a new antibiotic. I. Isolation and characterization of tunicamycin. J. Antibiot. 24, 215–223 (1971).
Miyazaki, Y., Chen, L. C., Chu, B. W., Swigut, T. & Wandless, T. J. Distinct transcriptional responses elicited by unfolded nuclear or cytoplasmic protein in mammalian cells. eLife 4, e07687 (2015).
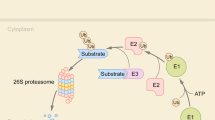
Schulman, B. A. & Harper, J. W. Ubiquitin-like protein activation by E1 enzymes: the apex for downstream signalling pathways. Nat. Rev. Mol. Cell Biol. 10, 319–331 (2009).
Ye, Y. & Rape, M. Building ubiquitin chains: E2 enzymes at work. Nat. Rev. Mol. Cell Biol. 10, 755–764 (2009).
Winston, J. T. et al. The SCFβ-TRCP–ubiquitin ligase complex associates specifically with phosphorylated destruction motifs in IκBα and β-catenin and stimulates IκBα ubiquitination in vitro. Genes Dev. 13, 270–283 (1999).
Fang, S., Jensen, J. P., Ludwig, R. L., Vousden, K. H. & Weissman, A. M. Mdm2 is a RING finger-dependent ubiquitin protein ligase for itself and p53. J. Biol. Chem. 275, 8945–8951 (2000).
Wade, M., Li, Y.-C. & Wahl, G. M. MDM2, MDMX and p53 in oncogenesis and cancer therapy. Nat. Rev. Cancer 13, 83–96 (2013).
Chau, V. et al. A multiubiquitin chain is confined to specific lysine in a targeted short-lived protein. Science 243, 1576–1583 (1989).
Lu, Y., Lee, B. H., King, R. W., Finley, D. & Kirschner, M. W. Substrate degradation by the proteasome: a single-molecule kinetic analysis. Science 348, 1250834 (2015).
Komander, D. & Rape, M. The ubiquitin code. Annu. Rev. Biochem. 81, 203–229 (2012).
Husnjak, K. & Dikic, I. Ubiquitin-binding proteins: decoders of ubiquitin-mediated cellular functions. Annu. Rev. Biochem. 81, 291–322 (2012).
MacGurn, J. A., Hsu, P. C. & Emr, S. D. Ubiquitin and membrane protein turnover: from cradle to grave. Annu. Rev. Biochem. 81, 231–259 (2012).
Lee, H., Puppala, D., Choi, E. Y., Swanson, H. & Kim, K. B. Targeted degradation of the aryl hydrocarbon receptor by the PROTAC approach: a useful chemical genetic tool. Chembiochem 8, 2058–2062 (2007).
Tomoshige, S., Naito, M., Hashimoto, Y. & Ishikawa, M. Degradation of HaloTag-fused nuclear proteins using bestatin-HaloTag ligand hybrid molecules. Org. Biomol. Chem. 13, 9746–9750 (2015).
Acknowledgements
The authors thank S. Jamie-Figueroa, T. Neklesa, K. Raina and the rest of the Crews laboratory for their helpful comments. C.M.C. gratefully acknowledges the US National Institutes of Health (NIH) for its support (R35-CA197589). A.C.L acknowledges support from the NIH (MSTP NIH/NIGMS T32GM007205). C.M.C. is founder, consultant and shareholder of Arvinas, LLC.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
C.M.C. is a founder, consultant and shareholder of Arvinas, LLC. A.C.L. declares no competing interests.
Related links
DATABASES
Glossary
- Event-driven pharmacology
-
A drug discovery paradigm based on identifying compounds that lead to the removal of the target protein from the system following drug binding to the target.
- Sub-stoichiometric
-
Pertains to molecular compounds that have a greater effect than the 1:1 activity ratio seen with a stoichiometric agent.
- HaloTag
-
A bacterial dehalogenase engineered by the Promega Corporation to covalently bind to chloroalkanes that has been used primarily for protein purification and imaging-based applications.
- Erythroblastosis oncogene B3
-
(ERBB3). A member of the epidermal growth factor receptor (EGFR) family that has minimal kinase activity and acts primarily as a scaffold for signal transduction.
- NF-κB inhibitor-α
-
(IκBα). Inhibitor of NF-κB that containing a destruction motif that, when phosphorylated, is recognized by the E3 ligase substrate recognition component β-transducin repeat-containing protein.
- von Hippel–Lindau
-
(VHL). The substrate recognition portion of the E3 ligase complex VHL–elongin B/C–cullin 2 (VHL–ELOBC–CUL2) that mediates the transfer of ubiquitin onto hypoxia-inducible factor 1α (HIF1α).
- Poly-D-arginine tag
-
An arginine-containing peptide fragment based on the HIV Tat peptide that facilitates cellular uptake of proteins.
- Occupancy-driven pharmacology
-
A drug discovery paradigm based on identifying compounds that modulate protein function through stoichiometric drug binding and occupation of the binding site to modulate protein function.
- Bromodomain-containing protein 4
-
(BRD4). A member of the bromodomain and extra-terminal (BET) family that recognizes acetylated lysines of histones to modulate the epigenetic code.
- Breakpoint cluster region–Abelson tyrosine kinase
-
(BCR–ABL). An oncogenic fusion of BCR and ABL that can lead to chronic myeloid leukaemia.
- Rule of five
-
The rule of five is a widely used guideline to predict the likelihood of poor absorption or permeability characteristics for small molecules. It is based on upper limits for physicochemical properties of the compounds, including molecular mass (for which 500 Da is the cut-off).
- Degrons
-
Peptide segments recognized by the cellular quality control system that influence the degradation rate of the full-length protein.
Rights and permissions
About this article
Cite this article
Lai, A., Crews, C. Induced protein degradation: an emerging drug discovery paradigm. Nat Rev Drug Discov 16, 101–114 (2017). https://doi.org/10.1038/nrd.2016.211
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrd.2016.211
This article is cited by
-
Tumor-targeted PROTAC prodrug nanoplatform enables precise protein degradation and combination cancer therapy
Acta Pharmacologica Sinica (2024)
-
Development and crystal structures of a potent second-generation dual degrader of BCL-2 and BCL-xL
Nature Communications (2024)
-
Deaggregation of mutant Plasmodium yoelii de-ubiquitinase UBP1 alters MDR1 localization to confer multidrug resistance
Nature Communications (2024)
-
Design, synthesis and biological evaluation of MNK-PROTACs
Molecular Diversity (2024)
-
Inhibition of ADAM9 promotes the selective degradation of KRAS and sensitizes pancreatic cancers to chemotherapy
Nature Cancer (2024)