Key Points
-
The metabolic ecology of tumours enables component cells to generate ATP, maintain redox balance, and undertake biosynthesis, which in turn support tumour progression
-
Tumours share features of complex ecosystems, with cancer cells inducing nutrient enrichment; however, the requirement for a tight nutrient balance might be a vulnerability of tumours that can be exploited therapeutically
-
Compared with their normal counterparts, tumour cells require higher rates of catabolite uptake, transfer, and utilization; hence, catabolite-deprivation might be a selective and effective anticancer treatment strategy
-
Targeting glycolysis and mitochondrial metabolism with drug combinations holds promise as another strategy to disrupt the diverse metabolic compartments within tumours
-
Measuring glucose, lactate, pyruvate, β-hydroxybutyrate, and glutamine levels in different tumour compartments and their intercompartmental transfer is needed in clinical trials that examine the efficacy of drugs targeting tumour metabolism
-
Normal tissues frequently have activation of metabolic pathways that are upregulated in cancer and, therefore, dose-limiting toxicity is a challenge in the development of drugs targeting these pathways
Abstract
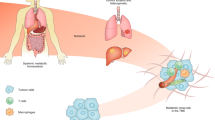
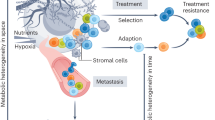
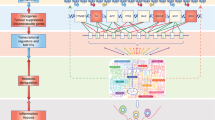
Awareness that the metabolic phenotype of cells within tumours is heterogeneous — and distinct from that of their normal counterparts — is growing. In general, tumour cells metabolize glucose, lactate, pyruvate, hydroxybutyrate, acetate, glutamine, and fatty acids at much higher rates than their nontumour equivalents; however, the metabolic ecology of tumours is complex because they contain multiple metabolic compartments, which are linked by the transfer of these catabolites. This metabolic variability and flexibility enables tumour cells to generate ATP as an energy source, while maintaining the reduction–oxidation (redox) balance and committing resources to biosynthesis — processes that are essential for cell survival, growth, and proliferation. Importantly, experimental evidence indicates that metabolic coupling between cell populations with different, complementary metabolic profiles can induce cancer progression. Thus, targeting the metabolic differences between tumour and normal cells holds promise as a novel anticancer strategy. In this Review, we discuss how cancer cells reprogramme their metabolism and that of other cells within the tumour microenvironment in order to survive and propagate, thus driving disease progression; in particular, we highlight potential metabolic vulnerabilities that might be targeted therapeutically.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Change history
17 January 2017
Owing to a typesetting error, the final line of text in Box 3, and the abbreviation lists for Tables 2 and 3 were omitted from the print and the online pdf versions of this article; for Table 3, the abbrevation list was also omitted from the online html version. These errors have now been corrected in the online pdf and html versions of the manuscript.
References
Ahn, C. S. & Metallo, C. M. Mitochondria as biosynthetic factories for cancer proliferation. Cancer Metab. 3, 1 (2015).
Vander Heiden, M. G., Cantley, L. C. & Thompson, C. B. Understanding the Warburg effect: the metabolic requirements of cell proliferation. Science 324, 1029–1033 (2009).
Pfeiffer, T., Schuster, S. & Bonhoeffer, S. Cooperation and competition in the evolution of ATP-producing pathways. Science 292, 504–507 (2001).
Cox, E. & Bonner, J. The advantages of togetherness. Science 292, 448–449 (2001).
Zu, X. L. & Guppy, M. Cancer metabolism: facts, fantasy, and fiction. Biochem. Biophys. Res. Commun. 313, 459–465 (2004).
Martinez-Outschoorn, U. E., Lisanti, M. P. & Sotgia, F. Catabolic cancer-associated fibroblasts transfer energy and biomass to anabolic cancer cells, fueling tumor growth. Semin. Cancer Biol. 25, 47–60 (2014).
Sonveaux, P. et al. Targeting lactate-fueled respiration selectively kills hypoxic tumor cells in mice. J. Clin. Invest. 118, 3930–3942 (2008).
Whitaker-Menezes, D. et al. Hyperactivation of oxidative mitochondrial metabolism in epithelial cancer cells in situ: visualizing the therapeutic effects of metformin in tumor tissue. Cell Cycle 10, 4047–4064 (2011).
Goodwin, M. L. et al. Modeling alveolar soft part sarcomagenesis in the mouse: a role for lactate in the tumor microenvironment. Cancer Cell 26, 851–862 (2014).
Doherty, J. R. & Cleveland, J. L. Targeting lactate metabolism for cancer therapeutics. J. Clin. Invest. 123, 3685–3692 (2013).
Pietrocola, F., Galluzzi, L., Bravo- San Pedro, J. M., Madeo, F. & Kroemer, G. Acetyl coenzyme A: a central metabolite and second messenger. Cell Metab. 21, 805–821 (2015).
Hanahan, D. & Weinberg, R. A. Hallmarks of cancer: the next generation. Cell 144, 646–674 (2011).
Viale, A. et al. Oncogene ablation-resistant pancreatic cancer cells depend on mitochondrial function. Nature 514, 628–632 (2014).
Wallace, D. C. Mitochondria and cancer Nat. Rev.Cancer 12, 685–698 (2012).
Pavlides, S. et al. The reverse Warburg effect: aerobic glycolysis in cancer associated fibroblasts and the tumor stroma. Cell Cycle 8, 3984–4001 (2009).
Nieman, K. M. et al. Adipocytes promote ovarian cancer metastasis and provide energy for rapid tumor growth. Nat. Med. 17, 1498–1503 (2011).
Fiaschi, T. et al. Reciprocal metabolic reprogramming through lactate shuttle coordinately influences tumor–stroma interplay. Cancer Res. 72, 5130–5140 (2012).
Boroughs, L. K. & DeBerardinis, R. J. Metabolic pathways promoting cancer cell survival and growth. Nat. Cell Biol. 17, 351–359 (2015).
Galluzzi, L., Kepp, O., Vander Heiden, M. G. & Kroemer, G. Metabolic targets for cancer therapy. Nat. Rev. Drug Discov. 12, 829–846 (2013).
Marchiq, I. & Pouyssegur, J. Hypoxia, cancer metabolism and the therapeutic benefit of targeting lactate/H+ symporters. J. Mol. Med. (Berl.) 94, 155–171 (2016).
Deep, G. & Agarwal, R. Targeting tumor microenvironment with silibinin: promise and potential for a translational cancer chemopreventive strategy. Curr. Cancer Drug Targets 13, 486–499 (2013).
Ooi, A. T. & Gomperts, B. N. Molecular pathways: targeting cellular energy metabolism in cancer via inhibition of SLC2A1 and LDHA. Clin. Cancer Res. 21, 2440–2444 (2015).
Vander Heiden, M. G. et al. Identification of small molecule inhibitors of pyruvate kinase M2. Biochem. Pharmacol. 79, 1118–1124 (2010).
Cortes-Cros, M. et al. M2 isoform of pyruvate kinase is dispensable for tumor maintenance and growth. Proc. Natl Acad. Sci. USA 110, 489–494 (2013).
Maschek, G. et al. 2-deoxy-d-glucose increases the efficacy of adriamycin and paclitaxel in human osteosarcoma and non-small cell lung cancers in vivo. Cancer Res. 64, 31–34 (2004).
Goldin, N. et al. Methyl jasmonate binds to and detaches mitochondria-bound hexokinase. Oncogene 27, 4636–4643 (2008).
Dwarakanath, B. S. et al. Clinical studies for improving radiotherapy with 2-deoxy-D-glucose: present status and future prospects. J. Cancer Res. Ther. 5 (Suppl. 1), S21–S26 (2009).
Papaldo, P. et al. Addition of either lonidamine or granulocyte colony-stimulating factor does not improve survival in early breast cancer patients treated with high-dose epirubicin and cyclophosphamide. J. Clin. Oncol. 21, 3462–3468 (2003).
DeBerardinis, R. J. & Cheng, T. Q's next: the diverse functions of glutamine in metabolism, cell biology and cancer. Oncogene 29, 313–324 (2009).
Marin-Valencia, I. et al. Analysis of tumor metabolism reveals mitochondrial glucose oxidation in genetically diverse human glioblastomas in the mouse brain in vivo. Cell Metab. 15, 827–837 (2012).
Lai, H. S., Lee, J. C., Lee, P. H., Wang, S. T. & Chen, W. J. Plasma free amino acid profile in cancer patients. Semin. Cancer Biol. 15, 267–276 (2005).
Fan, J. et al. Glutamine-driven oxidative phosphorylation is a major ATP source in transformed mammalian cells in both normoxia and hypoxia. Mol. Syst. Biol. 9, 712 (2013).
Metallo, C. M. et al. Reductive glutamine metabolism by IDH1 mediates lipogenesis under hypoxia. Nature 481, 380–384 (2011).
Hensley, C. T., Wasti, A. T. & DeBerardinis, R. J. Glutamine and cancer: cell biology, physiology, and clinical opportunities. J. Clin. Invest. 123, 3678–3684 (2013).
Mullen, A. R. et al. Reductive carboxylation supports growth in tumour cells with defective mitochondria. Nature 481, 385–388 (2011).
Diehn, M. et al. Association of reactive oxygen species levels and radioresistance in cancer stem cells. Nature 458, 780–783 (2009).
Xiang, Y. et al. Targeted inhibition of tumor-specific glutaminase diminishes cell-autonomous tumorigenesis. J. Clin. Invest. 125, 2293–2306 (2015).
Gross, M. I. et al. Antitumor activity of the glutaminase inhibitor CB-839 in triple-negative breast cancer. Mol. Cancer Ther. 13, 890–901 (2014).
Weinberg, F. et al. Mitochondrial metabolism and ROS generation are essential for Kras-mediated tumorigenicity. Proc. Natl Acad. Sci. USA 107, 8788–8793 (2010).
Filipp, F. V. et al. Glutamine-fueled mitochondrial metabolism is decoupled from glycolysis in melanoma. Pigment Cell Melanoma Res. 25, 732–739 (2012).
Hirayama, A. et al. Quantitative metabolome profiling of colon and stomach cancer microenvironment by capillary electrophoresis time-of-flight mass spectrometry. Cancer Res. 69, 4918–4925 (2009).
Ogawa, T., Washio, J., Takahashi, T., Echigo, S. & Takahashi, N. Glucose and glutamine metabolism in oral squamous cell carcinoma: insight from a quantitative metabolomic approach. Oral Surg. Oral Med. Oral Pathol. Oral Radiol. 118, 218–225 (2014).
Hirschhaeuser, F., Sattler, U. G. & Mueller-Klieser, W. Lactate: a metabolic key player in cancer. Cancer Res. 71, 6921–6925 (2011).
Kennedy, K. M. et al. Catabolism of exogenous lactate reveals it as a legitimate metabolic substrate in breast cancer. PLoS ONE 8, e75154 (2013).
Walenta, S., Schroeder, T. & Mueller-Klieser, W. Lactate in solid malignant tumors: potential basis of a metabolic classification in clinical oncology. Curr. Med. Chem. 11, 2195–2204 (2004).
Joyce, J. A. & Fearon, D. T. T cell exclusion, immune privilege, and the tumor microenvironment. Science 348, 74–80 (2015).
Colegio, O. R. et al. Functional polarization of tumour-associated macrophages by tumour-derived lactic acid. Nature 513, 559–563 (2013).
Le, A. et al. Inhibition of lactate dehydrogenase A induces oxidative stress and inhibits tumor progression. Proc. Natl Acad. Sci. USA 107, 2037–2042 (2010).
Doherty, J. R. et al. Blocking lactate export by inhibiting the Myc target MCT1 disables glycolysis and glutathione synthesis. Cancer Res. 74, 908–920 (2014).
Martinez-Outschoorn, U. E., Sotgia, F. & Lisanti, M. P. Power surge: supporting cells 'fuel' cancer cell mitochondria. Cell Metab. 15, 4–5 (2012).
Sotgia, F. et al. Caveolin-1 and cancer metabolism in the tumor microenvironment: markers, models, and mechanisms. Annu. Rev. Pathol. 7, 423–467 (2012).
Zhang, D. et al. Metabolic reprogramming of cancer-associated fibroblasts by IDH3α downregulation. Cell Rep. 10, 1335–1348 (2015).
Polanski, R. et al. Activity of the monocarboxylate transporter 1 inhibitor AZD3965 in small cell lung cancer. Clin. Cancer Res. 20, 926–937 (2014).
Halestrap, A. P. Monocarboxylic acid transport. Compr. Physiol. 3, 1611–1643 (2013).
Pellerin, L. & Magistretti, P. J. Sweet sixteen for ANLS. J. Cereb. Blood Flow Metab. 32, 1152–1166 (2012).
Scharfman, H. E. Metabolic control of epilepsy. Science 347, 1312–1313 (2015).
Sada, N., Lee, S., Katsu, T., Otsuki, T. & Inoue, T. Targeting LDH enzymes with a stiripentol analog to treat epilepsy. Science 347, 1362–1367 (2015).
Bailey, K. M., Wojtkowiak, J. W., Hashim, A. I. & Gillies, R. J. Targeting the metabolic microenvironment of tumors. Adv. Pharmacol. 65, 63–107 (2012).
Semenza, G. L. Hypoxia-inducible factors: mediators of cancer progression and targets for cancer therapy. Trends Pharmacol. Sci. 33, 207–214 (2012).
Rani, R. & Kumar, V. Recent update on human lactate dehydrogenase enzyme 5 (hLDH5) inhibitors: a promising approach for cancer chemotherapy. J. Med. Chem. 59, 487–496 (2016).
Kanno, T. et al. Lactate dehydrogenase M-subunit deficiency: a new type of hereditary exertional myopathy. Clin. Chim. Acta 173, 89–98 (1988).
Diers, A. R., Broniowska, K. A., Chang, C. F. & Hogg, N. Pyruvate fuels mitochondrial respiration and proliferation of breast cancer cells: effect of monocarboxylate transporter inhibition. Biochem. J. 444, 561–571 (2012).
Hatzivassiliou, G. et al. ATP citrate lyase inhibition can suppress tumor cell growth. Cancer Cell 8, 311–321 (2005).
Kamphorst, J. J. et al. Hypoxic and Ras-transformed cells support growth by scavenging unsaturated fatty acids from lysophospholipids. Proc. Natl Acad. Sci. USA 110, 8882–8887 (2013).
Gaglio, D. et al. Oncogenic K-Ras decouples glucose and glutamine metabolism to support cancer cell growth. Mol. Syst. Biol. 7, 523 (2011).
Porstmann, T. et al. SREBP activity is regulated by mTORC1 and contributes to Akt-dependent cell growth. Cell Metab. 8, 224–236 (2008).
Duvel, K. et al. Activation of a metabolic gene regulatory network downstream of mTOR complex 1. Mol. Cell 39, 171–183 (2010).
Grassian, A. R., Metallo, C. M., Coloff, J. L., Stephanopoulos, G. & Brugge, J. S. Erk regulation of pyruvate dehydrogenase flux through PDK4 modulates cell proliferation. Genes Dev. 25, 1716–1733 (2011).
Schug, Z. T. et al. Acetyl-CoA synthetase 2 promotes acetate utilization and maintains cancer cell growth under metabolic stress. Cancer Cell 27, 57–71 (2015).
Rabinowitz, J. D. & White, E. Autophagy and metabolism. Science 330, 1344–1348 (2010).
Sun, Y. et al. Treatment-induced damage to the tumor microenvironment promotes prostate cancer therapy resistance through WNT16B. Nat. Med. 18, 1359–1368 (2012).
Maslowski, K. M. et al. Regulation of inflammatory responses by gut microbiota and chemoattractant receptor GPR43. Nature 461, 1282–1286 (2009).
Ley, R. E., Turnbaugh, P. J., Klein, S. & Gordon, J. I. Microbial ecology: human gut microbes associated with obesity. Nature 444, 1022–1023 (2006).
Clemente, J. C., Ursell, L. K., Parfrey, L. W. & Knight, R. The impact of the gut microbiota on human health: an integrative view. Cell 148, 1258–1270 (2012).
Abreu, M. T. & Peek, R. M. Jr. Gastrointestinal malignancy and the microbiome. Gastroenterology 146, 1534–1546.e1533 (2014).
Paoli, A., Bosco, G., Camporesi, E. M. & Mangar, D. Ketosis, ketogenic diet and food intake control: a complex relationship. Front. Psychol. 6, 27 (2015).
Kruiswijk, F., Labuschagne, C. F. & Vousden, K. H. p53 in survival, death and metabolic health: a lifeguard with a licence to kill. Nat. Rev. Mol. Cell Biol. 16, 393–405 (2015).
Jeon, S. M., Chandel, N. S. & Hay, N. AMPK regulates NADPH homeostasis to promote tumour cell survival during energy stress. Nature 485, 661–665 (2012).
Parks, S. K., Mazure, N. M., Counillon, L. & Pouyssegur, J. Hypoxia promotes tumor cell survival in acidic conditions by preserving ATP levels. J. Cell. Physiol. 228, 1854–1862 (2013).
Caro, P. et al. Metabolic signatures uncover distinct targets in molecular subsets of diffuse large B cell lymphoma. Cancer Cell 22, 547–560 (2012).
Woyach, J. A. et al. Resistance mechanisms for the Bruton's tyrosine kinase inhibitor ibrutinib. N. Engl. J. Med. 370, 2286–2294 (2014).
Pike, L. S., Smift, A. L., Croteau, N. J., Ferrick, D. A. & Wu, M. Inhibition of fatty acid oxidation by etomoxir impairs NADPH production and increases reactive oxygen species resulting in ATP depletion and cell death in human glioblastoma cells. Biochim. Biophys. Acta 1807, 726–734 (2011).
Schlaepfer, I. R. et al. Hypoxia induces triglycerides accumulation in prostate cancer cells and extracellular vesicles supporting growth and invasiveness following reoxygenation. Oncotarget 6, 22836–22856 (2015).
Zaugg, K. et al. Carnitine palmitoyltransferase 1C promotes cell survival and tumor growth under conditions of metabolic stress. Genes Dev. 25, 1041–1051 (2011).
Holubarsch, C. J. et al. A double-blind randomized multicentre clinical trial to evaluate the efficacy and safety of two doses of etomoxir in comparison with placebo in patients with moderate congestive heart failure: the ERGO (etomoxir for the recovery of glucose oxidation) study. Clin. Sci. (Lond.) 113, 205–212 (2007).
Bertolini, F., Sukhatme, V. P. & Bouche, G. Drug repurposing in oncology — patient and health systems opportunities. Nat. Rev. Clin. Oncol. 12, 732–742 (2015).
Hamanaka, R. B. & Chandel, N. S. Targeting glucose metabolism for cancer therapy. J. Exp. Med. 209, 211–215 (2012).
Wilson, P. M., Danenberg, P. V., Johnston, P. G., Lenz, H. J. & Ladner, R. D. Standing the test of time: targeting thymidylate biosynthesis in cancer therapy. Nat. Rev. Clin. Oncol. 11, 282–298 (2014).
Pieters, R. et al. Pharmacokinetics, pharmacodynamics, efficacy, and safety of a new recombinant asparaginase preparation in children with previously untreated acute lymphoblastic leukemia: a randomized phase 2 clinical trial. Blood 112, 4832–4838 (2008).
Ascierto, P. A. et al. Pegylated arginine deiminase treatment of patients with metastatic melanoma: results from phase I and II studies. J. Clin. Oncol. 23, 7660–7668 (2005).
Glazer, E. S. et al. Phase II study of pegylated arginine deiminase for nonresectable and metastatic hepatocellular carcinoma. J. Clin. Oncol. 28, 2220–2226 (2010).
Zhai, L. et al. Molecular pathways: targeting IDO1 and other tryptophan dioxygenases for cancer immunotherapy. Clin. Cancer Res. 21, 5427–5433 (2015).
Possemato, R. et al. Functional genomics reveal that the serine synthesis pathway is essential in breast cancer. Nature 476, 346–350 (2011).
Locasale, J. W. et al. Phosphoglycerate dehydrogenase diverts glycolytic flux and contributes to oncogenesis. Nat. Genet. 43, 869–874 (2011).
DeNicola, G. M. et al. NRF2 regulates serine biosynthesis in non-small cell lung cancer. Nat. Genet. 47, 1475–1481 (2015).
Li, B. & Simon, M. C. Molecular pathways: targeting MYC-induced metabolic reprogramming and oncogenic stress in cancer. Clin. Cancer Res. 19, 5835–5841 (2013).
Stine, Z. E., Walton, Z. E., Altman, B. J., Hsieh, A. L. & Dang, C. V. MYC, metabolism and cancer. Cancer Discov. 5, 1024–1039 (2015).
Fan, J. et al. Human phosphoglycerate dehydrogenase produces the oncometabolite d-2-hydroxyglutarate. ACS Chem. Biol. 10, 510–516 (2015).
Agrawal, S., Kumar, A., Srivastava, V. & Mishra, B. N. Cloning, expression, activity and folding studies of serine hydroxymethyltransferase: a target enzyme for cancer chemotherapy. J. Mol. Microbiol. Biotechnol. 6, 67–75 (2003).
Fan, J. et al. Quantitative flux analysis reveals folate-dependent NADPH production. Nature 510, 298–302 (2014).
Jain, M. et al. Metabolite profiling identifies a key role for glycine in rapid cancer cell proliferation. Science 336, 1040–1044 (2012).
Bauer, D. E., Hatzivassiliou, G., Zhao, F., Andreadis, C. & Thompson, C. B. ATP citrate lyase is an important component of cell growth and transformation. Oncogene 24, 6314–6322 (2005).
Chajes, V., Cambot, M., Moreau, K., Lenoir, G. M. & Joulin, V. Acetyl-CoA carboxylase α is essential to breast cancer cell survival. Cancer Res. 66, 5287–5294 (2006).
Clem, B. F. et al. A novel small molecule antagonist of choline kinase-alpha that simultaneously suppresses MAPK and PI3K/AKT signaling. Oncogene 30, 3370–3380 (2011).
Flavin, R., Peluso, S., Nguyen, P. L. & Loda, M. Fatty acid synthase as a potential therapeutic target in cancer. Future Oncol. 6, 551–562 (2010).
Mulvihill, M. M. & Nomura, D. K. Therapeutic potential of monoacylglycerol lipase inhibitors. Life Sci. 92, 492–497 (2013).
Gallego-Ortega, D., Gomez del Pulgar, T., Valdes-Mora, F., Cebrian, A. & Lacal, J. C. Involvement of human choline kinase alpha and beta in carcinogenesis: a different role in lipid metabolism and biological functions. Adv. Enzyme Regul. 51, 183–194 (2011).
Glunde, K., Bhujwalla, Z. M. & Ronen, S. M. Choline metabolism in malignant transformation. Nat. Rev. Cancer 11, 835–848 (2011).
Kubatka, P., Kruzliak, P., Rotrekl, V., Jelinkova, S. & Mladosievicova, B. Statins in oncological research: from experimental studies to clinical practice. Crit. Rev. Oncol. Hematol. 92, 296–311 (2014).
Nielsen, S. F., Nordestgaard, B. G. & Bojesen, S. E. Statin use and reduced cancer-related mortality. N. Engl. J. Med. 367, 1792–1802 (2012).
Sriskanthadevan, S. et al. AML cells have low spare reserve capacity in their respiratory chain that renders them susceptible to oxidative metabolic stress. Blood 125, 2120–2130 (2015).
Funes, J. M. et al. Transformation of human mesenchymal stem cells increases their dependency on oxidative phosphorylation for energy production. Proc. Natl Acad. Sci. USA 104, 6223–6228 (2007).
Fogal, V. et al. Mitochondrial p32 protein is a critical regulator of tumor metabolism via maintenance of oxidative phosphorylation. Mol. Cell. Biol. 30, 1303–1318 (2010).
Curry, J. M. et al. Cancer metabolism, stemness and tumor recurrence: MCT1 and MCT4 are functional biomarkers of metabolic symbiosis in head and neck cancer. Cell Cycle 12, 1371–1384 (2013).
Wurm, C. A. et al. Nanoscale distribution of mitochondrial import receptor Tom20 is adjusted to cellular conditions and exhibits an inner-cellular gradient. Proc. Natl Acad. Sci. USA 108, 13546–13551 (2011).
Gehrke, S. et al. PINK1 and Parkin control localized translation of respiratory chain component mRNAs on mitochondria outer membrane. Cell Metab. 21, 95–108 (2015).
Vazquez, F. et al. PGC1α expression defines a subset of human melanoma tumors with increased mitochondrial capacity and resistance to oxidative stress. Cancer Cell 23, 287–301 (2013).
LeBleu, V. S. et al. PGC-1α mediates mitochondrial biogenesis and oxidative phosphorylation in cancer cells to promote metastasis. Nat. Cell Biol. 16, 992–1003 (2014).
Tan, A. S. et al. Mitochondrial genome acquisition restores respiratory function and tumorigenic potential of cancer cells without mitochondrial DNA. Cell Metab. 21, 81–94 (2015).
Katajisto, P. et al. Asymmetric apportioning of aged mitochondria between daughter cells is required for stemness. Science 348, 340–343 (2015).
Lamb, R. et al. Antibiotics that target mitochondria effectively eradicate cancer stem cells, across multiple tumor types: treating cancer like an infectious disease. Oncotarget 6, 4569–4584 (2015).
Lamb, R. et al. Dissecting tumor metabolic heterogeneity: telomerase and large cell size metabolically define a sub-population of stem-like, mitochondrial-rich, cancer cells. Oncotarget 6, 21892–21905 (2015).
Lo-Coco, F. et al. Retinoic acid and arsenic trioxide for acute promyelocytic leukemia. N. Engl. J. Med. 369, 111–121 (2013).
Owen, M. R., Doran, E. & Halestrap, A. P. Evidence that metformin exerts its anti-diabetic effects through inhibition of complex 1 of the mitochondrial respiratory chain. Biochem. J. 348, 607–614 (2000).
Pelicano, H. et al. Inhibition of mitochondrial respiration: a novel strategy to enhance drug-induced apoptosis in human leukemia cells by a reactive oxygen species-mediated mechanism. J. Biol. Chem. 278, 37832–37839 (2003).
Pollak, M. Overcoming drug development bottlenecks with repurposing: repurposing biguanides to target energy metabolism for cancer treatment. Nat. Med. 20, 591–593 (2014).
Bowker, S. L., Majumdar, S. R., Veugelers, P. & Johnson, J. A. Increased cancer-related mortality for patients with type 2 diabetes who use sulfonylureas or insulin. Diabetes Care 29, 254–258 (2006).
Hirsch, H. A., Iliopoulos, D., Tsichlis, P. N. & Struhl, K. Metformin selectively targets cancer stem cells, and acts together with chemotherapy to block tumor growth and prolong remission. Cancer Res. 69, 7507–7511 (2009).
Hadad, S. et al. Evidence for biological effects of metformin in operable breast cancer: a pre-operative, window-of-opportunity, randomized trial. Breast Cancer Res. Treat. 128, 783–794 (2011).
Menendez, J. A. et al. Metformin is synthetically lethal with glucose withdrawal in cancer cells. Cell Cycle 11, 2782–2792 (2012).
Birsoy, K. et al. Metabolic determinants of cancer cell sensitivity to glucose limitation and biguanides. Nature 508, 108–112 (2014).
Chen, G., Xu, S., Renko, K. & Derwahl, M. Metformin inhibits growth of thyroid carcinoma cells, suppresses self-renewal of derived cancer stem cells, and potentiates the effect of chemotherapeutic agents. J. Clin. Endocrinol. Metab. 97, E510–E520 (2012).
Song, C. W. et al. Metformin kills and radiosensitizes cancer cells and preferentially kills cancer stem cells. Sci. Rep. 2, 362 (2012).
Bao, B. et al. Metformin inhibits cell proliferation, migration and invasion by attenuating CSC function mediated by deregulating miRNAs in pancreatic cancer cells. Cancer Prev. Res. (Phila.) 5, 355–364 (2012).
Foretz, M., Guigas, B., Bertrand, L., Pollak, M. & Viollet, B. Metformin: from mechanisms of action to therapies. Cell Metab. 20, 953–966 (2014).
Kordes, S. et al. Metformin in patients with advanced pancreatic cancer: a double-blind, randomised, placebo-controlled phase 2 trial. Lancet Oncol. 16, 839–847 (2015).
Salani, B. et al. Metformin, cancer and glucose metabolism. Endocr. Relat. Cancer 21, R461–R471 (2014).
Britten, C. D. et al. A phase I and pharmacokinetic study of the mitochondrial-specific rhodacyanine dye analog MKT 077. Clin. Cancer Res. 6, 42–49 (2000).
Propper, D. J. et al. Phase I trial of the selective mitochondrial toxin MKT077 in chemo-resistant solid tumours. Ann. Oncol. 10, 923–927 (1999).
Michelakis, E. D., Webster, L. & Mackey, J. R. Dichloroacetate (DCA) as a potential metabolic-targeting therapy for cancer. Br. J. Cancer 99, 989–994 (2008).
Fu, X. et al. 2-hydroxyglutarate inhibits ATP synthase and mTOR signaling. Cell Metab. 22, 508–515 (2015).
Yan, H., Bigner, D. D., Velculescu, V. & Parsons, D. W. Mutant metabolic enzymes are at the origin of gliomas. Cancer Res. 69, 9157–9159 (2009).
Fathi, A. T., Wander, S. A., Faramand, R. & Emadi, A. Biochemical, epigenetic and metabolic approaches to target IDH mutations in acute myeloid leukemia. Semin Hematol. 52, 165–171 (2015).
Sagan, L. On the origin of mitosing cells. J. Theor. Biol. 14, 255–274 (1967).
Beckmann, R. & Herrmann, J. M. Mitoribosome oddities. Science 348, 288–289 (2015).
Greber, B. J. et al. The complete structure of the 55S mammalian mitochondrial ribosome. Science 348, 303–308 (2015).
Amunts, A., Brown, A., Toots, J., Scheres, S. H. & Ramakrishnan, V. The structure of the human mitochondrial ribosome. Science 348, 95–98 (2015).
Amunts, A. et al. Structure of the yeast mitochondrial large ribosomal subunit. Science 343, 1485–1489 (2014).
Brown, A. et al. Structure of the large ribosomal subunit from human mitochondria. Science 346, 718–722 (2014).
Wilson, D. N. Ribosome-targeting antibiotics and mechanisms of bacterial resistance. Nat. Rev. Microbiol. 12, 35–48 (2014).
Carter, A. P. et al. Functional insights from the structure of the 30S ribosomal subunit and its interactions with antibiotics. Nature 407, 340–348 (2000).
Moazed, D. & Noller, H. F. Interaction of antibiotics with functional sites in 16S ribosomal RNA. Nature 327, 389–394 (1987).
Greber, B. J. & Ban, N. Structure and function of the mitochondrial ribosome. Annu. Rev. Biochem. http://dx.doi.org/10.1146/annurevbiochem060815014343 (2016).
Prezant, T. R. et al. Mitochondrial ribosomal RNA mutation associated with both antibiotic-induced and non-syndromic deafness. Nat. Genet. 4, 289–294 (1993).
Bitner-Glindzicz, M. et al. Prevalence of mitochondrial 1555A→G mutation in European children. N. Engl. J. Med. 360, 640–642 (2009).
Soriano, A., Miro, O. & Mensa, J. Mitochondrial toxicity associated with linezolid. N. Engl. J. Med. 353, 2305–2306 (2005).
Ferreri, A. J. et al. Bacteria-eradicating therapy with doxycycline in ocular adnexal MALT lymphoma: a multicenter prospective trial. J. Natl Cancer Inst. 98, 1375–1382 (2006).
Ferreri, A. J. et al. Chlamydophila psittaci eradication with doxycycline as first-line targeted therapy for ocular adnexae lymphoma: final results of an international phase II trial. J. Clin. Oncol. 30, 2988–2994 (2012).
Skrtic, M. et al. Inhibition of mitochondrial translation as a therapeutic strategy for human acute myeloid leukemia. Cancer Cell 20, 674–688 (2011).
Bahrami, F., Morris, D. L. & Pourgholami, M. H. Tetracyclines: drugs with huge therapeutic potential. Mini Rev. Med. Chem. 12, 44–52 (2012).
Yang, M., Soga, T. & Pollard, P. J. Oncometabolites: linking altered metabolism with cancer. J. Clin. Invest. 123, 3652–3658 (2013).
Vazquez, A., Liu, J., Zhou, Y. & Oltvai, Z. N. Catabolic efficiency of aerobic glycolysis: the Warburg effect revisited. BMC Syst. Biol. 4, 58 (2010).
Shlomi, T., Benyamini, T., Gottlieb, E., Sharan, R. & Ruppin, E. Genome-scale metabolic modeling elucidates the role of proliferative adaptation in causing the Warburg effect. PLoS Comput. Biol. 7, e1002018 (2011).
Sena, L. A. & Chandel, N. S. Physiological roles of mitochondrial reactive oxygen species. Mol. Cell 48, 158–167 (2012).
Martinez-Outschoorn, U. E. et al. Oxidative stress in cancer associated fibroblasts drives tumor-stroma co-evolution: a new paradigm for understanding tumor metabolism, the field effect and genomic instability in cancer cells. Cell Cycle 9, 3256–3276 (2010).
Asterholm, I. W., Mundy, D. I., Weng, J., Anderson, R. G. & Scherer, P. E. Altered mitochondrial function and metabolic inflexibility associated with loss of caveolin-1. Cell Metab. 15, 171–185 (2012).
Chen, J. L. et al. The genomic analysis of lactic acidosis and acidosis response in human cancers. PLoS Genet. 4, e1000293 (2008).
Xie, J. et al. Beyond Warburg effect — dual metabolic nature of cancer cells. Sci. Rep. 4, 4927 (2014).
Gorrini, C., Harris, I. S. & Mak, T. W. Modulation of oxidative stress as an anticancer strategy. Nat. Rev. Drug Discov. 12, 931–947 (2013).
Nisoli, E. et al. Mitochondrial biogenesis in mammals: the role of endogenous nitric oxide. Science 299, 896–899 (2003).
Nisoli, E. et al. Calorie restriction promotes mitochondrial biogenesis by inducing the expression of eNOS. Science 310, 314–317 (2005).
Hasmann, M. & Schemainda, I. FK866, a highly specific noncompetitive inhibitor of nicotinamide phosphoribosyltransferase, represents a novel mechanism for induction of tumor cell apoptosis. Cancer Res. 63, 7436–7442 (2003).
Sampath, D., Zabka, T. S., Misner, D. L., O'Brien, T. & Dragovich, P. S. Inhibition of nicotinamide phosphoribosyltransferase (NAMPT) as a therapeutic strategy in cancer. Pharmacol. Ther. 151, 16–31 (2015).
Burgos, E. S. NAMPT in regulated NAD biosynthesis and its pivotal role in human metabolism. Curr. Med. Chem. 18, 1947–1961 (2011).
Yun, J. et al. Vitamin C selectively kills KRAS and BRAF mutant colorectal cancer cells by targeting GAPDH. Science 350, 1391–1396 (2015).
Porporato, P. E. et al. A mitochondrial switch promotes tumor metastasis. Cell Rep. 8, 754–766 (2014).
DeNicola, G. M. et al. Oncogene-induced Nrf2 transcription promotes ROS detoxification and tumorigenesis. Nature 475, 106–109 (2011).
Wells, G. Peptide and small molecule inhibitors of the Keap1–Nrf2 protein–protein interaction. Biochem. Soc. Trans. 43, 674–679 (2015).
Wang, G. L., Jiang, B. H., Rue, E. A. & Semenza, G. L. Hypoxia-inducible factor 1 is a basic-helix-loop- helix-PAS heterodimer regulated by cellular O2 tension. Proc. Natl Acad. Sci. USA 92, 5510–5514 (1995).
Zhang, H. et al. HIF-1 inhibits mitochondrial biogenesis and cellular respiration in VHL-deficient renal cell carcinoma by repression of C-MYC activity. Cancer Cell 11, 407–420 (2007).
Kim, J. W., Gao, P., Liu, Y. C., Semenza, G. L. & Dang, C. V. Hypoxia-inducible factor 1 and dysregulated c-Myc cooperatively induce vascular endothelial growth factor and metabolic switches hexokinase 2 and pyruvate dehydrogenase kinase 1. Mol. Cell. Biol. 27, 7381–7393 (2007).
Luo, W. et al. Pyruvate kinase M2 is a PHD3-stimulated coactivator for hypoxia-inducible factor 1. Cell 145, 732–744 (2011).
Fukuda, R. et al. HIF-1 regulates cytochrome oxidase subunits to optimize efficiency of respiration in hypoxic cells. Cell 129, 111–122 (2007).
Zhang, H. et al. Mitochondrial autophagy is an HIF-1-dependent adaptive metabolic response to hypoxia. J. Biol. Chem. 283, 10892–10903 (2008).
Semenza, G. L. HIF-1 mediates metabolic responses to intratumoral hypoxia and oncogenic mutations. J. Clin. Invest. 123, 3664–3671 (2013).
Velasco-Hernandez, T., Hyrenius-Wittsten, A., Rehn, M., Bryder, D. & Cammenga, J. HIF-1α can act as a tumor suppressor gene in murine acute myeloid leukemia. Blood 124, 3597–3607 (2014).
Chiavarina, B. et al. HIF1-alpha functions as a tumor promoter in cancer associated fibroblasts, and as a tumor suppressor in breast cancer cells: autophagy drives compartment-specific oncogenesis. Cell Cycle 9, 3534–3551 (2010).
Wilson, W. R. & Hay, M. P. Targeting hypoxia in cancer therapy. Nat. Rev. Cancer 11, 393–410 (2011).
Talekar, M., Boreddy, S. R., Singh, A. & Amiji, M. Tumor aerobic glycolysis: new insights into therapeutic strategies with targeted delivery. Expert Opin. Biol. Ther. 14, 1145–1159 (2014).
Chen, B. J., Wu, Y. L., Tanaka, Y. & Zhang, W. Small molecules targeting c-Myc oncogene: promising anti-cancer therapeutics. Int. J. Biol. Sci. 10, 1084–1096 (2014).
Yuneva, M. O. et al. The metabolic profile of tumors depends on both the responsible genetic lesion and tissue type. Cell Metab. 15, 157–170 (2012).
Valencia, T. et al. Metabolic reprogramming of stromal fibroblasts through p62–mTORC1 signaling promotes inflammation and tumorigenesis. Cancer Cell 26, 121–135 (2014).
Amend, S. R. & Pienta, K. J. Ecology meets cancer biology: the cancer swamp promotes the lethal cancer phenotype. Oncotarget 6, 9669–9678 (2015).
Draoui, N. & Feron, O. Lactate shuttles at a glance: from physiological paradigms to anti-cancer treatments. Dis. Model. Mech. 4, 727–732 (2011).
White, E. Deconvoluting the context-dependent role for autophagy in cancer. Nat. Rev. Cancer 12, 401–410 (2012).
Martinez-Outschoorn, U. E. et al. Autophagy in cancer associated fibroblasts promotes tumor cell survival: role of hypoxia, HIF1 induction and NFκB activation in the tumor stromal microenvironment. Cell Cycle 9, 3515–3533 (2010).
Takeuchi, H. et al. Synergistic augmentation of rapamycin-induced autophagy in malignant glioma cells by phosphatidylinositol 3-kinase/protein kinase B inhibitors. Cancer Res. 65, 3336–3346 (2005).
Chiarini, F., Evangelisti, C., McCubrey, J. A. & Martelli, A. M. Current treatment strategies for inhibiting mTOR in cancer. Trends Pharmacol. Sci. 36, 124–135 (2015).
Kimura, T., Takabatake, Y., Takahashi, A. & Isaka, Y. Chloroquine in cancer therapy: a double-edged sword of autophagy. Cancer Res. 73, 3–7 (2013).
Farber, S. & Diamond, L. K. Temporary remissions in acute leukemia in children produced by folic acid antagonist, 4-aminopteroyl-glutamic acid. N. Engl. J. Med. 238, 787–793 (1948).
Visentin, M., Zhao, R. & Goldman, I. D. The antifolates. Hematol. Oncol. Clin. North Am. 26, 629–648 (2012).
Kaelin, W. G. Jr & McKnight, S. L. Influence of metabolism on epigenetics and disease. Cell 153, 56–69 (2013).
Ye, D., Ma, S., Xiong, Y. & Guan, K. L. R-2-hydroxyglutarate as the key effector of IDH mutations promoting oncogenesis. Cancer Cell 23, 274–276 (2013).
Dang, L. et al. Cancer-associated IDH1 mutations produce 2-hydroxyglutarate. Nature 462, 739–744 (2009).
Losman, J. A. et al. (R)-2-hydroxyglutarate is sufficient to promote leukemogenesis and its effects are reversible. Science 339, 1621–1625 (2013).
Rohle, D. et al. An inhibitor of mutant IDH1 delays growth and promotes differentiation of glioma cells. Science 340, 626–630 (2013).
Smolkova, K., Dvorak, A., Zelenka, J., Vitek, L. & Jezek, P. Reductive carboxylation and 2-hydroxyglutarate formation by wild-type IDH2 in breast carcinoma cells. Int. J. Biochem. Cell Biol. 65, 125–133 (2015).
Intlekofer, A. M. et al. Hypoxia induces production of L-2-hydroxyglutarate. Cell Metab. 22, 304–311 (2015).
Carrer, A. & Wellen, K. E. Metabolism and epigenetics: a link cancer cells exploit. Curr. Opin. Biotechnol. 34, 23–29 (2015).
Peiris-Pages, M., Martinez-Outschoorn, U. E., Sotgia, F. & Lisanti, M. P. Metastasis and oxidative stress: are antioxidants a metabolic driver of progression? Cell Metab. 22, 956–958 (2015).
Clem, B. F. et al. Targeting 6-phosphofructo-2-kinase (PFKFB3) as a therapeutic strategy against cancer. Mol. Cancer Ther. 12, 1461–1470 (2013).
Morais-Santos, F. et al. Targeting lactate transport suppresses in vivo breast tumour growth. Oncotarget 6, 19177–19189 (2015).
Fujiwara, S. et al. PDK1 inhibition is a novel therapeutic target in multiple myeloma. Br. J. Cancer 108, 170–178 (2013).
Sellers, K. et al. Pyruvate carboxylase is critical for non-small-cell lung cancer proliferation. J. Clin. Invest. 125, 687–698 (2015).
Pardee, T. S. et al. A phase I study of the first-in-class antimitochondrial metabolism agent, CPI-613, in patients with advanced hematologic malignancies. Clin. Cancer Res. 20, 5255–5264 (2014).
Wang, F. et al. Targeted inhibition of mutant IDH2 in leukemia cells induces cellular differentiation. Science 340, 622–626 (2013).
El-Mir, M. Y. et al. Dimethylbiguanide inhibits cell respiration via an indirect effect targeted on the respiratory chain complex I. J. Biol. Chem. 275, 223–228 (2000).
Jara, J. A. & Lopez-Munoz, R. Metformin and cancer: between the bioenergetic disturbances and the antifolate activity. Pharmacol. Res. 101, 102–108 (2015).
Hitosugi, T. et al. Phosphoglycerate mutase 1 coordinates glycolysis and biosynthesis to promote tumor growth. Cancer Cell 22, 585–600 (2012).
Feun, L. G., Kuo, M. T. & Savaraj, N. Arginine deprivation in cancer therapy. Curr. Opin. Clin. Nutr. Metab. Care 18, 78–82 (2015).
Lamb, R. et al. Doxycycline down-regulates DNA-PK and radiosensitizes tumor initiating cells: implications for more effective radiation therapy. Oncotarget 6, 14005–14025 (2015).
Pham, E. et al. Translational impact of nanoparticle-drug conjugate CRLX101 with or without bevacizumab in advanced ovarian cancer. Clin. Cancer Res. 21, 808–818 (2015).
Galluzzi, L. et al. Autophagy in malignant transformation and cancer progression. EMBO J. 34, 856–880 (2015).
Zadra, G., Batista, J. L. & Loda, M. Dissecting the dual role of AMPK in cancer: from experimental to human studies. Mol. Cancer Res. 13, 1059–1072 (2015).
Sayin, V. I. et al. Antioxidants accelerate lung cancer progression in mice. Sci. Transl. Med. 6, 221ra215 (2014).
Pavlides, S. et al. Warburg meets autophagy: cancer-associated fibroblasts accelerate tumor growth and metastasis via oxidative stress, mitophagy, and aerobic glycolysis. Antioxid. Redox Signal. 16, 1264–1284 (2012).
Acknowledgements
The Sotgia and Lisanti laboratories in the UK have been supported, in part, by funding from the EU (European Research Council Advanced Grant), Breast Cancer Now, The Healthy Life Foundation, and the Manchester Cancer Research Centre (MCRC). The work of Ubaldo E. Martinez-Outschoorn has been supported by the National Cancer Institute (NCI) of the National Institutes of Health (NIH), under Award Number K08-CA175193. Richard G. Pestell's laboratory receives funding from the NIH and the NCI, as well as the Breast Cancer Research Foundation and the Ralph and Marian C. Falk Medical Research Trust.
Author information
Authors and Affiliations
Contributions
U.E.M.-O. and M.P.-P. researched the data for the article and wrote the manuscript. U.E.M.-O., M.P.-P., F.S., and M.P.L. contributed substantially to discussions of content, and U.E.M.-O., R.G.P., F.S., and M.P.L. reviewed and edited the manuscript before submission.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Martinez-Outschoorn, U., Peiris-Pagés, M., Pestell, R. et al. Cancer metabolism: a therapeutic perspective. Nat Rev Clin Oncol 14, 11–31 (2017). https://doi.org/10.1038/nrclinonc.2016.60
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrclinonc.2016.60
This article is cited by
-
The molecular subtyping and precision medicine in triple-negative breast cancer---based on Fudan TNBC classification
Cancer Cell International (2024)
-
HIF-2α/LINC02609/APOL1-mediated lipid storage promotes endoplasmic reticulum homeostasis and regulates tumor progression in clear-cell renal cell carcinoma
Journal of Experimental & Clinical Cancer Research (2024)
-
A systematic pan-cancer study on deep learning-based prediction of multi-omic biomarkers from routine pathology images
Communications Medicine (2024)
-
SIRT5-mediated ME2 desuccinylation promotes cancer growth by enhancing mitochondrial respiration
Cell Death & Differentiation (2024)
-
α-Ketoglutarate supplementation and NAD+ modulation enhance metabolic rewiring and radiosensitization in SLC25A1 inhibited cancer cells
Cell Death Discovery (2024)