Key Points
-
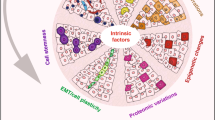
Primary human tumours consist of cells that differ in clinically important phenotypic features. This phenotypic heterogeneity is a result of the interplay between genetic and non-genetic factors that shape cellular phenotypes.
-
Genomic instability, which is frequently observed in human cancers, in combination with the large numbers of cell divisions required for the formation of macroscopic tumours, leads to inevitable genetic diversity in populations of tumour cells.
-
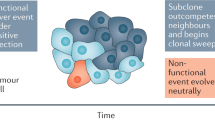
Somatic evolution that drives tumour progression is characterized by complex dynamics arising from the Darwinian nature of the process. As a result, individual tumours have a unique clonal architecture that is spatially and temporally heterogeneous.
-
The cancer stem cell perspective can explain only some of the non-genetic variability in tumour cell phenotypes. A more comprehensive explanation of non-genetic sources of phenotypic heterogeneity necessitates the consideration of mechanisms that underlie cellular phenotypes.
-
Both deterministic and stochastic determinants of cellular phenotypes can be substantially affected during oncogenic transformation and tumour progression, contributing both to abnormal phenotypes and to an increased degree of phenotypic plasticity.
-
Phenotypic and genetic heterogeneity within tumours impedes clinical diagnostics: owing to topological heterogeneity in the distribution of diagnostically important phenotypes even multiple sampling might not provide adequate information. At the same time, given the link between a high degree of genetic heterogeneity and poor prognosis, a measure of heterogeneity by itself may be useful as a prognostic marker.
-
Phenotypic heterogeneity in tumour cell populations that results from both genetic and non-genetic determinants constitutes a major source of therapeutic resistance. Initial phenotypic heterogeneity and changes in cellular phenotypes resulting from adaptation to response and selection for resistant phenotypes need to be accounted for in order to achieve substantial improvements in therapeutic outcomes.
Abstract
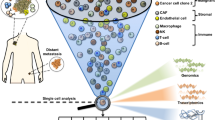
Populations of tumour cells display remarkable variability in almost every discernable phenotypic trait, including clinically important phenotypes such as ability to seed metastases and to survive therapy. This phenotypic diversity results from the integration of both genetic and non-genetic influences. Recent technological advances have improved the molecular understanding of cancers and the identification of targets for therapeutic interventions. However, it has become exceedingly apparent that the utility of profiles based on the analysis of tumours en masse is limited by intra-tumour genetic and epigenetic heterogeneity, as characteristics of the most abundant cell type might not necessarily predict the properties of mixed populations. In this Review, we discuss both genetic and non-genetic causes of phenotypic heterogeneity of tumour cells, with an emphasis on heritable phenotypes that serve as a substrate for clonal selection. We discuss the implications of intra-tumour heterogeneity in diagnostics and the development of therapeutic resistance.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Fidler, I. J. Tumor heterogeneity and the biology of cancer invasion and metastasis. Cancer Res. 38, 2651–2660 (1978).
Gerlinger, M. et al. Intratumor heterogeneity and branched evolution revealed by multiregion sequencing. N. Engl. J. Med. 366, 883–892 (2012). This paper demonstrated substantial spatial genetic heterogeneity within tumours, including synonymous driving mutations independently arising in distinct clones. Furthermore, it reported distinct diagnostic signatures from different biopsies of the same tumour.
Anderson, K. et al. Genetic variegation of clonal architecture and propagating cells in leukaemia. Nature 469, 356–361 (2011). This paper demonstrated substantial genetic variability in putative driver mutations between leukaemia stem cells, suggesting complex patterns of clonal evolution.
Campbell, L. L. & Polyak, K. Breast tumor heterogeneity: cancer stem cells or clonal evolution? Cell Cycle 6, 2332–2338 (2007).
Shackleton, M., Quintana, E., Fearon, E. R. & Morrison, S. J. Heterogeneity in cancer: cancer stem cells versus clonal evolution. Cell 138, 822–829 (2009).
Notta, F. et al. Evolution of human BCR-ABL1 lymphoblastic leukaemia-initiating cells. Nature 469, 362–367 (2011). This paper demonstrated substantial genetic diversity between leukaemia-initiating cells, suggesting complex clonal architecture.
Shipitsin, M. et al. Molecular definition of breast tumor heterogeneity. Cancer Cell 11, 259–273 (2007). This paper demonstrated that presumed stem cell and non-stem cell subpopulations in breast tumours can have different patterns of mutations.
Vogelstein, B. & Kinzler, K. W. Cancer genes and the pathways they control. Nature Med. 10, 789–799 (2004).
Rangarajan, A., Hong, S. J., Gifford, A. & Weinberg, R. A. Species- and cell type-specific requirements for cellular transformation. Cancer Cell 6, 171–183 (2004).
Ruiz, C. et al. Advancing a clinically relevant perspective of the clonal nature of cancer. Proc. Natl Acad. Sci. USA 108, 12054–12059 (2011).
Ashworth, A., Lord, C. J. & Reis-Filho, J. S. Genetic interactions in cancer progression and treatment. Cell 145, 30–38 (2011).
Raj, A. & van Oudenaarden, A. Nature, nurture, or chance: stochastic gene expression and its consequences. Cell 135, 216–226 (2008).
Niepel, M., Spencer, S. L. & Sorger, P. K. Non-genetic cell-to-cell variability and the consequences for pharmacology. Curr. Opin. Chem. Biol. 13, 556–561 (2009).
Altschuler, S. J. & Wu, L. F. Cellular heterogeneity: do differences make a difference? Cell 141, 559–563 (2010).
Huang, S. Non-genetic heterogeneity of cells in development: more than just noise. Development 136, 3853–3862 (2009).
Eldar, A. & Elowitz, M. B. Functional roles for noise in genetic circuits. Nature 467, 167–173 (2010).
Dalerba, P. et al. Single-cell dissection of transcriptional heterogeneity in human colon tumors. Nature Biotechnol. 29, 1120–1127 (2011).
Clevers, H. The cancer stem cell: premises, promises and challenges. Nature Med. 17, 313–319 (2011).
Rosen, J. M. & Jordan, C. T. The increasing complexity of the cancer stem cell paradigm. Science 324, 1670–1673 (2009).
Chaffer, C. L. et al. Normal and neoplastic nonstem cells can spontaneously convert to a stem-like state. Proc. Natl Acad. Sci. USA. 108, 7950–7955 (2011).
Quintana, E. et al. Efficient tumour formation by single human melanoma cells. Nature 456, 593–598 (2008).
Balazsi, G., van Oudenaarden, A. & Collins, J. J. Cellular decision making and biological noise: from microbes to mammals. Cell 144, 910–925 (2011).
Arias, A. M. & Hayward, P. Filtering transcriptional noise during development: concepts and mechanisms. Nature Rev. Genet. 7, 34–44 (2006).
Hughes, T. R. et al. Functional discovery via a compendium of expression profiles. Cell 102, 109–126 (2000).
Luo, J., Solimini, N. L. & Elledge, S. J. Principles of cancer therapy: oncogene and non-oncogene addiction. Cell 136, 823–837 (2009).
Whitesell, L. & Lindquist, S. L. HSP90 and the chaperoning of cancer. Nature Rev. Cancer 5, 761–772 (2005).
Egeblad, M., Nakasone, E. S. & Werb, Z. Tumors as organs: complex tissues that interface with the entire organism. Dev. Cell 18, 884–901 (2010).
Allinen, M. et al. Molecular characterization of the tumor microenvironment in breast cancer. Cancer Cell 6, 17–32 (2004).
Tlsty, T. D. & Coussens, L. M. Tumor stroma and regulation of cancer development. Annu. Rev. Pathol. 1, 119–150 (2006).
Bissell, M. J., Kenny, P. A. & Radisky, D. C. Microenvironmental regulators of tissue structure and function also regulate tumor induction and progression: the role of extracellular matrix and its degrading enzymes. Cold Spring Harb. Symp. Quant. Biol. 70, 343–356 (2005).
Anderson, A. R., Weaver, A. M., Cummings, P. T. & Quaranta, V. Tumor morphology and phenotypic evolution driven by selective pressure from the microenvironment. Cell 127, 905–915 (2006).
Chen, J., Sprouffske, K., Huang, Q. & Maley, C. C. Solving the puzzle of metastasis: the evolution of cell migration in neoplasms. PloS ONE 6, e17933 (2011).
Polyak, K. & Weinberg, R. A. Transitions between epithelial and mesenchymal states: acquisition of malignant and stem cell traits. Nature Rev. Cancer 9, 265–273 (2009).
Hendrix, M. J., Seftor, E. A., Hess, A. R. & Seftor, R. E. Vasculogenic mimicry and tumour-cell plasticity: lessons from melanoma. Nature Rev. Cancer 3, 411–421 (2003).
Radisky, D. C. & Bissell, M. J. Matrix metalloproteinase-induced genomic instability. Curr. Opin. Genet. Dev. 16, 45–50 (2006).
Goel, S. et al. Normalization of the vasculature for treatment of cancer and other diseases. Physiol. Rev. 91, 1071–1121 (2011).
Farmer, P. et al. A stroma-related gene signature predicts resistance to neoadjuvant chemotherapy in breast cancer. Nature Med. 15, 68–74 (2009).
Finak, G. et al. Stromal gene expression predicts clinical outcome in breast cancer. Nature Med. 14, 518–527 (2008).
Nowell, P. C. The clonal evolution of tumor cell populations. Science 194, 23–28 (1976).
Merlo, L. M., Pepper, J. W., Reid, B. J. & Maley, C. C. Cancer as an evolutionary and ecological process. Nature Rev. Cancer 6, 924–935 (2006).
Jones, P. A. & Baylin, S. B. The epigenomics of cancer. Cell 128, 683–692 (2007).
Brock, A., Chang, H. & Huang, S. Non-genetic heterogeneity--a mutation-independent driving force for the somatic evolution of tumours. Nature Rev. Genetics 10, 336–342 (2009).
Weinberg, R. A. The Biology of Cancer (Garland Science, 2007).
Barrick, J. E. & Lenski, R. E. Genome-wide mutational diversity in an evolving population of Escherichia coli. Cold Spring Harb. Symp. Quant. Biol. 74, 119–129 (2009).
Desai, M. M., Fisher, D. S. & Murray, A. W. The speed of evolution and maintenance of variation in asexual populations. Curr. Biol. 17, 385–394 (2007).
Marusyk, A. & Polyak, K. Tumor heterogeneity: causes and consequences. Biochim. Biophys. Acta 1805, 105–117 (2010).
Navin, N. et al. Tumour evolution inferred by single-cell sequencing. Nature 472, 90–94 (2011). The first report that interrogated genomic-wide diversity within tumours on a single-cell level.
Tsao, J. L. et al. Colorectal adenoma and cancer divergence. Evidence of multilineage progression. Am. J. Pathol. 154, 1815–1824 (1999).
Gonzalez-Garcia, I., Sole, R. V. & Costa, J. Metapopulation dynamics and spatial heterogeneity in cancer. Proc. Natl Acad. Sci. USA 99, 13085–13089 (2002).
Snuderl, M. et al. Mosaic amplification of multiple receptor tyrosine kinase genes in glioblastoma. Cancer Cell 20, 810–817 (2011).
Stoecklein, N. H. & Klein, C. A. Genetic disparity between primary tumours, disseminated tumour cells, and manifest metastasis. Int. J. Cancer. 126, 589–598 (2010).
Fearon, E. R. & Vogelstein, B. A genetic model for colorectal tumorigenesis. Cell 61, 759–767 (1990).
Ramaswamy, S., Ross, K. N., Lander, E. S. & Golub, T. R. A molecular signature of metastasis in primary solid tumors. Nature Genet. 33, 49–54 (2003).
Weigelt, B. et al. Molecular portraits and 70-gene prognosis signature are preserved throughout the metastatic process of breast cancer. Cancer Res. 65, 9155–9158 (2005).
Klein, C. A. Parallel progression of primary tumours and metastases. Nature Rev. Cancer 9, 302–312 (2009).
Mardis, E. R. Cancer genomics identifies determinants of tumor biology. Genome Biol. 11, 211 (2010).
Yachida, S. et al. Distant metastasis occurs late during the genetic evolution of pancreatic cancer. Nature 467, 1114–1117 (2010).
Shah, S. P. et al. Mutational evolution in a lobular breast tumour profiled at single nucleotide resolution. Nature 461, 809–813 (2009). The first report that used next-generation sequencing to interrogate clonal diversity and evolution during breast cancer progression.
Ding, L. et al. Genome remodelling in a basal-like breast cancer metastasis and xenograft. Nature 464, 999–1005 (2010).
Perou, C. M., Parker, J. S., Prat, A., Ellis, M. J. & Bernard, P. S. Clinical implementation of the intrinsic subtypes of breast cancer. Lancet Oncol. 11, 718–719 (2010).
Sorlie, T. et al. Gene expression patterns of breast carcinomas distinguish tumor subclasses with clinical implications. Proc. Natl Acad. Sci. USA 98, 10869–10874 (2001).
Komaki, K., Sano, N. & Tangoku, A. Problems in histological grading of malignancy and its clinical significance in patients with operable breast cancer. Breast cancer 13, 249–253 (2006).
Bertos, N. R. & Park, M. Breast cancer - one term, many entities? J. Clin. Invest. 121, 3789–3796 (2011).
Allred, D. C. et al. Ductal carcinoma in situ and the emergence of diversity during breast cancer evolution. Clin. Cancer Res. 14, 370–378 (2008).
Maley, C. C. et al. Genetic clonal diversity predicts progression to esophageal adenocarcinoma. Nature Genet. 38, 468–473 (2006). This paper linked higher extent of genetic clonal diversity with poor clinical outcome.
Merlo, L. M. & Maley, C. C. The role of genetic diversity in cancer. J. Clin. Invest. 120, 401–403 (2010).
Park, S. Y. et al. Heterogeneity for stem cell-related markers according to tumor subtype and histologic stage in breast cancer. Clin. Cancer Res. 16, 876–887 (2010).
Meads, M. B., Gatenby, R. A. & Dalton, W. S. Environment-mediated drug resistance: a major contributor to minimal residual disease. Nature Rev. Cancer 9, 665–674 (2009).
Ebos, J. M. et al. Accelerated metastasis after short-term treatment with a potent inhibitor of tumor angiogenesis. Cancer Cell 15, 232–239 (2009).
Paez-Ribes, M. et al. Antiangiogenic therapy elicits malignant progression of tumors to increased local invasion and distant metastasis. Cancer Cell 15, 220–231 (2009).
Deininger, M. Resistance to imatinib: mechanisms and management. J. Natl Compr. Canc. Netw. 3, 757–768 (2005).
Corless, C. L. & Heinrich, M. C. Molecular pathobiology of gastrointestinal stromal sarcomas. Annu. Rev. Pathol. 3, 557–586 (2008).
Nazarian, R. et al. Melanomas acquire resistance to B-RAF(V600E) inhibition by RTK or N-RAS upregulation. Nature 468, 973–977 (2010).
Engelman, J. A. et al. MET amplification leads to gefitinib resistance in lung cancer by activating ERBB3 signaling. Science 316, 1039–1043 (2007).
Edwards, S. L. et al. Resistance to therapy caused by intragenic deletion in BRCA2. Nature 451, 1111–1115 (2008).
Sakai, W. et al. Secondary mutations as a mechanism of cisplatin resistance in BRCA2-mutated cancers. Nature 451, 1116–1120 (2008).
Roche-Lestienne, C., Lai, J. L., Darre, S., Facon, T. & Preudhomme, C. A mutation conferring resistance to imatinib at the time of diagnosis of chronic myelogenous leukemia. N. Engl. J. Med. 348, 2265–2266 (2003).
Ding, L. et al. Clonal evolution in relapsed acute myeloid leukaemia revealed by whole-genome sequencing. Nature 481, 506–510 (2012).
Mullighan, C. G. et al. Genomic analysis of the clonal origins of relapsed acute lymphoblastic leukemia. Science 322, 1377–1380 (2008).
Glasspool, R. M., Teodoridis, J. M. & Brown, R. Epigenetics as a mechanism driving polygenic clinical drug resistance. Br. J. Cancer 94, 1087–1092 (2006).
Janne, P. A., Gray, N. & Settleman, J. Factors underlying sensitivity of cancers to small-molecule kinase inhibitors. Nature Rev. Drug Discov. 8, 709–723 (2009).
Sharma, S. V. et al. A chromatin-mediated reversible drug-tolerant state in cancer cell subpopulations. Cell 141, 69–80 (2010). This study linked therapy resistance with a distinct epigenetic switch that arises through a spontaneous phenotypic switch. Although this phenotypic state is reversible, it can persist for multiple cell generations and thus serve as a substrate for Darwinian selection.
Cohen, A. A. et al. Dynamic proteomics of individual cancer cells in response to a drug. Science 322, 1511–1516 (2008).
Spencer, S. L., Gaudet, S., Albeck, J. G., Burke, J. M. & Sorger, P. K. Non-genetic origins of cell-to-cell variability in TRAIL-induced apoptosis. Nature 459, 428–432 (2009). This study implicated noise-driven stochastic non-genetic variability in short-term resistance to therapy.
Heppner, G. H. Cancer cell societies and tumor progression. Stem Cells 11, 199–203 (1993).
Bonavia, R., Inda, M. M., Cavenee, W. K. & Furnari, F. B. Heterogeneity maintenance in glioblastoma: a social network. Cancer Res. 71, 4055–4060 (2011).
Park, S. Y., Gönen, M., Kim, H. J., Michor, F. & Polyak, K. Cellular and genetic diversity in the progression of in situ human breast carcinomas to an invasive phenotype. J. Clin. Invest. 120, 636–644 (2010).
Merlo, L. M. et al. A comprehensive survey of clonal diversity measures in Barrett's esophagus as biomarkers of progression to esophageal adenocarcinoma. Cancer Prev. Res. (Phila) 3, 1388–1397 (2010).
Russnes, H. G., Navin, N., Hicks, J. & Borresen-Dale, A. L. Insight into the heterogeneity of breast cancer through next-generation sequencing. J. Clin. Invest. 121, 3810–3818 (2011).
Navin, N. et al. Inferring tumor progression from genomic heterogeneity. Genome Res. 20, 68–80 (2010).
Xu, X. et al. Single-cell exome sequencing reveals single-nucleotide mutation characteristics of a kidney tumor. Cell 148, 886–895 (2012).
Hou, Y. et al. Single-cell exome sequencing and monoclonal evolution of a JAK2-negative myeloproliferative neoplasm. Cell 148, 873–885 (2012).
Miller, B. E., Machemer, T., Lehotan, M. & Heppner, G. H. Tumor subpopulation interactions affecting melphalan sensitivity in palpable mouse mammary tumors. Cancer Res. 51, 4378–4387 (1991).
Calbo, J. et al. A functional role for tumor cell heterogeneity in a mouse model of small cell lung cancer. Cancer Cell 19, 244–256 (2011).
Marks, P. A. & Dokmanovic, M. Histone deacetylase inhibitors: discovery and development as anticancer agents. Expert Opin. Investig. Drugs 14, 1497–1511 (2005).
Gregory, R. I. & Shiekhattar, R. Chromatin modifiers and carcinogenesis. Trends Cell. Biol. 14, 695–702 (2004).
Bissell, M. J. & Hines, W. C. Why don't we get more cancer? A proposed role of the microenvironment in restraining cancer progression. Nature Med. 17, 320–329 (2011).
Attolini, C. S. & Michor, F. Evolutionary theory of cancer. Ann. NY Acad. Sci. 1168, 23–51 (2009).
DeGregori, J. Evolved tumor suppression: why are we so good at not getting cancer? Cancer Res. 71, 3739–3744 (2011).
Greaves, M. & Maley, C. C. Clonal evolution in cancer. Nature 481, 306–313 (2012).
Gatenby, R. A., Silva, A. S., Gillies, R. J. & Frieden, B. R. Adaptive therapy. Cancer Res. 69, 4894–4903 (2009). This paper proposes 'adaptive therapy', which takes into account clonal competition in order to prevent therapy-resistant cells from taking over the population.
Waddington, C. H. The Strategy of the Genes: a Discussion of Some Aspects of Theoretical Biology (Taylor & Francis, 1957).
Huang, S. & Kauffman, S. A. in Encyclopedia of Complexity and Systems Science (ed. Meyers, R. A.) 1180–1213 (Springer, 2009).
Fisher, A. G. Cellular identity and lineage choice. Nature Rev. Immunol. 2, 977–982 (2002).
Bozic, I. et al. Accumulation of driver and passenger mutations during tumor progression. Proc. Natl Acad. Sci. USA 107, 18545–18550 (2010).
Leedham, S. J. et al. Individual crypt genetic heterogeneity and the origin of metaplastic glandular epithelium in human Barrett's oesophagus. Gut 57, 1041–1948 (2008).
Kim, M. Y. et al. Tumor self-seeding by circulating cancer cells. Cell 139, 1315–1326 (2009).
Acknowledgements
We thank members of our laboratory and F. Michor and S. Itzkovitz for their critical reading of our manuscript and for stimulating discussions. Tumour diversity research in the authors' laboratory is supported by US Army Congressionally Directed Research W81XWH-07-1-0294 (K.P.) and BC087579 (A.M.), US National Cancer Institute PO1 CA80111 (K.P.), Susan G. Komen Foundation (K.P.), Breast Cancer Research Foundation (K.P.) and the Cellex Foundation (V.A.).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Related links
FURTHER INFORMATION
Glossary
- Phenotypic plasticity
-
The ability of cells to change phenotype stochastically or in response to changes in the environment.
- Effective population sizes
-
The numbers of tumour cells capable of passing their genotypes to next generations.
- Allelic imbalances
-
Changes in the copy number of alleles as a result of chromosomal amplification or deletions.
- Heat shock protein response
-
A response to cellular stress involving the activation of molecular chaperones called heat shock proteins (HSPs). HSPs help to maintain cellular homeostasis and to promote survival in the face of stress.
- Clone
-
A group of cells that share a common ancestor and that are genetically identical. Whereas all of the cells in the body originate from a single ancestor, every new mutation creates a new clone (a subclone).
- Clonal diversity
-
The diversity of clones within a tumour; results from a branching pattern of evolution. Increased with larger population size, smaller fitness differences, higher mutation rates and heterogeneity of environments.
Rights and permissions
About this article
Cite this article
Marusyk, A., Almendro, V. & Polyak, K. Intra-tumour heterogeneity: a looking glass for cancer?. Nat Rev Cancer 12, 323–334 (2012). https://doi.org/10.1038/nrc3261
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrc3261
This article is cited by
-
Deciphering the role of neddylation in tumor microenvironment modulation: common outcome of multiple signaling pathways
Biomarker Research (2024)
-
A new function for drug combination dose finding trials
Scientific Reports (2024)
-
MedGAN: optimized generative adversarial network with graph convolutional networks for novel molecule design
Scientific Reports (2024)
-
The loss of B7-H4 expression in breast cancer cells escaping from T cell cytotoxicity contributes to epithelial-to-mesenchymal transition
Breast Cancer Research (2023)
-
Synergistic inter-clonal cooperation involving crosstalk, co-option and co-dependency can enhance the invasiveness of genetically distant cancer clones
BMC Ecology and Evolution (2023)