Key Points
-
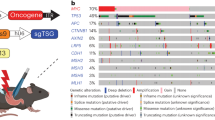
Genetically engineered mouse models (GEMMs) have been invaluable in advancing our knowledge of tumour biology. However, accelerated cancer gene discovery through large-scale cancer genomics and an increasing desire to use GEMMs for preclinical therapeutic studies have strained the capacity of germline GEMMs.
-
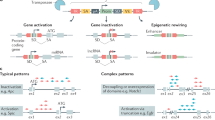
Non-germline genetic engineering approaches allow for accelerated and flexible genetic manipulation of models.
-
Chimeric models develop tumours in the context of normal stroma, with reduced timelines and mouse housing cost.
-
Transplantation models allow flexible and speedy manipulation of tissue stem and/or progenitor cells with multiple genetic tools (such as knock out, transgenes and RNA interference).
-
Human donor tissue models (or human in mouse models) allow the de novo development of primary human tumours in mouse stroma by manipulating primary human cells.
-
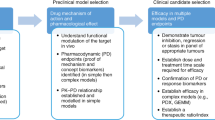
Therapeutic studies in vivo benefit from the wealth of complex GEMM and non-GEMM models to guide drug discovery.
Abstract
Genetically engineered mouse models (GEMMs) of cancer have affected virtually all areas of cancer research. However, the accelerated discovery of new cancer genes emerging from large-scale cancer genomics and new chemical entities pouring from the drug discovery pipeline have strained the capacity of traditional germline mouse models to provide crucial insights. This Review introduces new approaches to modelling cancer, with emphasis on a growing collection of non-germline GEMMs (nGEMMs). These offer flexibility, speed and uniformity at reduced costs, thus paving the way for much needed throughput and practical preclinical therapeutic testing models.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Sharpless, N. E. & Depinho, R. A. The mighty mouse: genetically engineered mouse models in cancer drug development. Nature Rev. Drug Discov. 5, 741–754 (2006).
Huettner, C. S., Zhang, P., Van Etten, R. A. & Tenen, D. G. Reversibility of acute B-cell leukaemia induced by BCR-ABL1. Nature Genet. 24, 57–60 (2000).
Roberts, R. B. et al. Importance of epidermal growth factor receptor signaling in establishment of adenomas and maintenance of carcinomas during intestinal tumorigenesis. Proc. Natl Acad. Sci. USA 99, 1521–1526 (2002).
Jonkers, J. & Berns, A. Conditional mouse models of sporadic cancer. Nature Rev. Cancer 2, 251–265 (2002).
Chin, L. et al. Essential role for oncogenic Ras in tumour maintenance. Nature 400, 468–472 (1999). The first conditional transgenic mouse model to demonstrate solid tumour regression on extinction of the initiating oncogene.
Su, L. K. et al. Multiple intestinal neoplasia caused by a mutation in the murine homolog of the APC gene. Science 256, 668–670 (1992).
Corpet, D. E. & Pierre, F. How good are rodent models of carcinogenesis in predicting efficacy in humans? A systematic review and meta-analysis of colon chemoprevention in rats, mice and men. Eur. J. Cancer 41, 1911–1922 (2005).
Jimeno, A. et al. A direct pancreatic cancer xenograft model as a platform for cancer stem cell therapeutic development. Mol. Cancer Ther. 8, 310–314 (2009).
Edelmann, W. et al. Mutation in the mismatch repair gene Msh6 causes cancer susceptibility. Cell 91, 467–477 (1997).
Meuwissen, R., Linn, S. C., van der Valk, M., Mooi, W. J. & Berns, A. Mouse model for lung tumorigenesis through Cre/lox controlled sporadic activation of the K-Ras oncogene. Oncogene 20, 6551–6558 (2001).
Tuveson, D. A. et al. Endogenous oncogenic K-rasG12D stimulates proliferation and widespread neoplastic and developmental defects. Cancer Cell 5, 375–387 (2004).
Dinulescu, D. M. et al. Role of K-ras and Pten in the development of mouse models of endometriosis and endometrioid ovarian cancer. Nature Med. 11, 63–70 (2005).
Flesken-Nikitin, A., Choi, K. C., Eng, J. P., Shmidt, E. N. & Nikitin, A. Y. Induction of carcinogenesis by concurrent inactivation of p53 and Rb1 in the mouse ovarian surface epithelium. Cancer Res. 63, 3459–3463 (2003).
Engelman, J. A. et al. Effective use of PI3K and MEK inhibitors to treat mutant Kras G12D and PIK3CA H1047R murine lung cancers. Nature Med. 14, 1351–1356 (2008).
Indra, A. K. et al. Temporally-controlled site-specific mutagenesis in the basal layer of the epidermis: comparison of the recombinase activity of the tamoxifen-inducible Cre-ER(T) and Cre-ER(T2) recombinases. Nucleic Acids Res. 27, 4324–4327 (1999).
Bosenberg, M. et al. Characterization of melanocyte-specific inducible Cre recombinase transgenic mice. Genesis 44, 262–267 (2006).
Dankort, D. et al. Braf(V600E) cooperates with Pten loss to induce metastatic melanoma. Nature Genet. 41, 544–552 (2009).
Dhomen, N. et al. Oncogenic Braf induces melanocyte senescence and melanoma in mice. Cancer Cell 15, 294–303 (2009).
Schmidt-Supprian, M., Wunderlich, F. T. & Rajewsky, K. Excision of the Frt-flanked neoR cassette from the CD19cre knock-in transgene reduces Cre-mediated recombination. Transgenic Res. 16, 657–660 (2007).
Gossen, M. et al. Transcriptional activation by tetracyclines in mammalian cells. Science 268, 1766–1769 (1995).
Felsher, D. W. & Bishop, J. M. Reversible tumorigenesis by MYC in hematopoietic lineages. Mol. Cell 4, 199–207 (1999).
Jain, M. et al. Sustained loss of a neoplastic phenotype by brief inactivation of MYC. Science 297, 102–104 (2002).
D'Cruz, C. M. et al. c-MYC induces mammary tumorigenesis by means of a preferred pathway involving spontaneous Kras2 mutations. Nature Med. 7, 235–239 (2001).
Rao, G. et al. Sonic hedgehog and insulin-like growth factor signaling synergize to induce medulloblastoma formation from nestin-expressing neural progenitors in mice. Oncogene 23, 6156–6162 (2004).
Shih, A. H. et al. Dose-dependent effects of platelet-derived growth factor-B on glial tumorigenesis. Cancer Res. 64, 4783–4789 (2004).
Momota, H., Nerio, E. & Holland, E. C. Perifosine inhibits multiple signaling pathways in glial progenitors and cooperates with temozolomide to arrest cell proliferation in gliomas in vivo. Cancer Res. 65, 7429–7435 (2005).
Pao, W., Klimstra, D. S., Fisher, G. H. & Varmus, H. E. Use of avian retroviral vectors to introduce transcriptional regulators into mammalian cells for analyses of tumor maintenance. Proc. Natl Acad. Sci. USA 100, 8764–8769 (2003).
Regales, L. et al. Dual targeting of EGFR can overcome a major drug resistance mutation in mouse models of EGFR mutant lung cancer. J. Clin. Invest. 119, 3000–3010 (2009).
Marumoto, T. et al. Development of a novel mouse glioma model using lentiviral vectors. Nature Med. 15, 110–116 (2009).
Singh, M et al. Modelling therapeutic responses in Kras mutant cancers using genetically engineered mouse models. Nature Biotechnol. 23 May 2010 (doi: 10.103a8/nbt.1640).
Zhou, Y. et al. Chimeric mouse tumor models reveal differences in pathway activation between ERBB family- and KRAS-dependent lung adenocarcinomas. Nature Biotechnol. 28, 71–78 (2010). EGFR- but not KRAS-driven chimeric lung adenocarcinomas generated from engineered ESCs show near-complete tumour regression on treatment with an EGFR inhibitor.
Wu, M. et al. Dissecting genetic requirements of human breast tumorigenesis in a tissue transgenic model of human breast cancer in mice. Proc. Natl Acad. Sci. USA 106, 7022–7027 (2009). Normal breast cancer cells grown as organoids produce orthotopic tumours in mice with histopathological features that more closely resemble the primary tumour than the features of tumours generated from cells cultured on plastic.
Schmitt, C. A., McCurrach, M. E., de Stanchina, E., Wallace-Brodeur, R. R. & Lowe, S. W. INK4a/ARF mutations accelerate lymphomagenesis and promote chemoresistance by disabling p53. Genes Dev. 13, 2670–2677 (1999).
Zender, L. et al. Identification and validation of oncogenes in liver cancer using an integrative oncogenomic approach. Cell 125, 1253–1267 (2006). Liver progenitor cells harvested from embryonic mice and engineered with various oncogenes produce carcinomas orthotopically in syngeneic hosts and can be used to identify new oncogenes.
Pear, W. S. et al. Efficient and rapid induction of a chronic myelogenous leukemia-like myeloproliferative disease in mice receiving P210 bcr/abl-transduced bone marrow. Blood 92, 3780–3792 (1998).
Le, L. Q., Shipman, T., Burns, D. K. & Parada, L. F. Cell of origin and microenvironment contribution for NF1-associated dermal neurofibromas. Cell Stem Cell 4, 453–463 (2009).
Trimboli, A. J. et al. Pten in stromal fibroblasts suppresses mammary epithelial tumours. Nature 461, 1084–1091 (2009).
Paik, J. H. et al. FoxOs are lineage-restricted redundant tumor suppressors and regulate endothelial cell homeostasis. Cell 128, 309–323 (2007).
Jongsma, J. et al. A conditional mouse model for malignant mesothelioma. Cancer Cell 13, 261–271 (2008).
Watters, J. W. et al. De novo discovery of a γ-secretase inhibitor response signature using a novel in vivo breast tumor model. Cancer Res. 69, 8949–8957 (2009).
Hemann, M. T. et al. An epi-allelic series of p53 hypomorphs created by stable RNAi produces distinct tumor phenotypes in vivo. Nature Genet. 33, 396–400 (2003).
Takada, A. & Takada, Y. Proliferation of donor hematopoietic cells in lethally irradiated host mice. Transplantation 13, 276–280 (1972).
Schmitt, C. A. et al. Dissecting p53 tumor suppressor functions in vivo. Cancer Cell 1, 289–298 (2002).
Schwaller, J. et al. Transformation of hematopoietic cell lines to growth-factor independence and induction of a fatal myelo- and lymphoproliferative disease in mice by retrovirally transduced TEL/JAK2 fusion genes. EMBO J. 17, 5321–5333 (1998).
Fridman, J. S. et al. Tumor promotion by Mdm2 splice variants unable to bind p53. Cancer Res. 63, 5703–5706 (2003).
Wendel, H. G. et al. Dissecting eIF4E action in tumorigenesis. Genes Dev. 21, 3232–3237 (2007).
Phan, V. T. et al. Cooperation of cytokine signaling with chimeric transcription factors in leukemogenesis: PML-retinoic acid receptor alpha blocks growth factor-mediated differentiation. Mol. Cell. Biol. 23, 4573–4585 (2003).
Hemann, M. T. et al. Evasion of the p53 tumour surveillance network by tumour-derived MYC mutants. Nature 436, 807–811 (2005).
Schmitt, C. A., Rosenthal, C. T. & Lowe, S. W. Genetic analysis of chemoresistance in primary murine lymphomas. Nature Med. 6, 1029–1035 (2000).
Zuber, J. et al. Mouse models of human AML accurately predict chemotherapy response. Genes Dev. 23, 877–889 (2009). nGEMM leukaemia models driven by an MLLT1 gene fusion, but not ones driven by a RUNX1 gene fusion, are refractory to chemotherapy, mirroring the prognostic course of therapy in patients harbouring either of the respective fusions.
Wendel, H. G. et al. Survival signalling by Akt and eIF4E in oncogenesis and cancer therapy. Nature 428, 332–337 (2004).
Mullighan, C. G., Williams, R. T., Downing, J. R. & Sherr, C. J. Failure of CDKN2A/B (INK4A/B-ARF)-mediated tumor suppression and resistance to targeted therapy in acute lymphoblastic leukemia induced by BCR-ABL. Genes Dev. 22, 1411–1415 (2008).
Lauchle, J. O. et al. Response and resistance to MEK inhibition in leukaemias initiated by hyperactive Ras. Nature 461, 411–414 (2009).
Varticovski, L. et al. Accelerated preclinical testing using transplanted tumors from genetically engineered mouse breast cancer models. Clin. Cancer Res. 13, 2168–2177 (2007).
Rottenberg, S. et al. High sensitivity of BRCA1-deficient mammary tumors to the PARP inhibitor AZD2281 alone and in combination with platinum drugs. Proc. Natl Acad. Sci. USA 105, 17079–17084 (2008).
Rottenberg, S. et al. Selective induction of chemotherapy resistance of mammary tumors in a conditional mouse model for hereditary breast cancer. Proc. Natl Acad. Sci. USA 104, 12117–12122 (2007).
Evers, B. et al. A tissue reconstitution model to study cancer cell-intrinsic and -extrinsic factors in mammary tumourigenesis. J. Pathol. 220, 34–44 (2010).
Bachoo, R. M. et al. Epidermal growth factor receptor and Ink4a/Arf: convergent mechanisms governing terminal differentiation and transformation along the neural stem cell to astrocyte axis. Cancer Cell 1, 269–277 (2002).
Zindy, F. et al. Genetic alterations in mouse medulloblastomas and generation of tumors de novo from primary cerebellar granule neuron precursors. Cancer Res. 67, 2676–2684 (2007).
Xing, D. & Orsulic, S. A mouse model for the molecular characterization of brca1-associated ovarian carcinoma. Cancer Res. 66, 8949–8953 (2006).
Xue, W. et al. Senescence and tumour clearance is triggered by p53 restoration in murine liver carcinomas. Nature 445, 656–660 (2007).
Alcantara Llaguno, S. R., Chen, J. & Parada, L. F. Signaling in malignant astrocytomas: role of neural stem cells and its therapeutic implications. Clin. Cancer Res. 15, 7124–7129 (2009).
Uren, A. G. et al. Large-scale mutagenesis in p19ARF- and p53-deficient mice identifies cancer genes and their collaborative networks. Cell 133, 727–741 (2008).
Starr, T. K. et al. A transposon-based genetic screen in mice identifies genes altered in colorectal cancer. Science 323, 1747–1750 (2009).
Zender, L. et al. An oncogenomics-based in vivo RNAi screen identifies tumor suppressors in liver cancer. Cell 135, 852–864 (2008).
Witt, A. E. et al. Functional proteomics approach to investigate the biological activities of cDNAs implicated in breast cancer. J. Proteome Res. 5, 599–610 (2006).
Bric, A. et al. Functional identification of tumor-suppressor genes through an in vivo RNA interference screen in a mouse lymphoma model. Cancer Cell 16, 324–335 (2009).
Meacham, C. E., Ho, E. E., Dubrovsky, E., Gertler, F. B. & Hemann, M. T. In vivo RNAi screening identifies regulators of actin dynamics as key determinants of lymphoma progression. Nature Genet. 41, 1133–1137 (2009).
Wattel, E. et al. p53 mutations are associated with resistance to chemotherapy and short survival in hematologic malignancies. Blood 84, 3148–3157 (1994).
Jiang, H. et al. The combined status of ATM and p53 link tumor development with therapeutic response. Genes Dev. 23, 1895–1909 (2009).
Hoffman, R. M. Orthotopic metastatic (MetaMouse) models for discovery and development of novel chemotherapy. Methods Mol. Med. 111, 297–322 (2005).
Maser, R. S. et al. Chromosomally unstable mouse tumours have genomic alterations similar to diverse human cancers. Nature 447, 966–971 (2007).
Artandi, S. E. & DePinho, R. A. Telomeres and telomerase in cancer. Carcinogenesis 31, 9–18 (2009).
Rudolph, K. L. Telomeres and telomerase influence the course of human diseases, aging and carcinogenesis. Curr. Mol. Med. 5, 133–134 (2005).
Farazi, P. A., Glickman, J., Horner, J. & Depinho, R. A. Cooperative interactions of p53 mutation, telomere dysfunction, and chronic liver damage in hepatocellular carcinoma progression. Cancer Res. 66, 4766–4773 (2006).
Olive, K. P. et al. Mutant p53 gain of function in two mouse models of Li-Fraumeni syndrome. Cell 119, 847–860 (2004).
Fan, H., Oro, A. E., Scott, M. P. & Khavari, P. A. Induction of basal cell carcinoma features in transgenic human skin expressing Sonic Hedgehog. Nature Med. 3, 788–792 (1997). The first in a series of papers using normal human skin cells to orthotopically generate carcinomas and melanomas in mice by grafting with stromal elements.
Chudnovsky, Y., Adams, A. E., Robbins, P. B., Lin, Q. & Khavari, P. A. Use of human tissue to assess the oncogenic activity of melanoma-associated mutations. Nature Genet. 37, 745–749 (2005).
Khavari, P. A. Modelling cancer in human skin tissue. Nature Rev. Cancer 6, 270–280 (2006).
Lazarov, M. et al. CDK4 coexpression with Ras generates malignant human epidermal tumorigenesis. Nature Med. 8, 1105–1114 (2002).
Atillasoy, E. S. et al. UVB induces atypical melanocytic lesions and melanoma in human skin. Am. J. Pathol. 152, 1179–1186 (1998).
Barabe, F., Kennedy, J. A., Hope, K. J. & Dick, J. E. Modeling the initiation and progression of human acute leukemia in mice. Science 316, 600–604 (2007). A HIM leukaemia model using human cord blood transduced with an MLLT1 gene fusion can be used to identify LICs.
Kuperwasser, C. et al. Reconstruction of functionally normal and malignant human breast tissues in mice. Proc. Natl Acad. Sci. USA 101, 4966–4971 (2004).
Rong, S. et al. Tumorigenicity of the met proto-oncogene and the gene for hepatocyte growth factor. Mol. Cell. Biol. 12, 5152–5158 (1992).
Hanahan, D. & Weinberg, R. A. The hallmarks of cancer. Cell 100, 57–70 (2000).
Vogelstein, B. & Kinzler, K. W. Cancer genes and the pathways they control. Nature Med. 10, 789–799 (2004).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
J. H. is an employee of AVEO Pharmaceuticals.
Related links
Glossary
- Germline model
-
Mouse model that carries genetic modifications in its germline and which is maintained through breeding.
- Inducible model
-
Mouse model that activates the expression of a transgene through a transactivator transgene that is, tTA or rtTA.
- Non-germline GEMM (nGEMM)
-
Mouse model that carries genetic modifications in some of its somatic cells but not in the germline cells. Each model has to be individually generated through, for example, transplantation and injection.
- Mosaic model
-
Germline model that acquires modifications of the germline genetic modification in some of its somatic cells.
- Conditional GEMM
-
Model that acquires an activation or inactivation of the original genetic modification in somatic cells through the temporal or spatial expression of a modifier such as Cre.
- Chimeric model
-
Mouse model that has been generated by ESC manipulation followed by the injection of these cells into a pre-implantation embryo. The resulting chimeric animal is the model animal.
- Transplantation model
-
Mouse model in which part of a tissue is modified by transplanting tissue stem cells that carry genetic modifications.
- Human in mouse (HIM) model
-
Transplantation model in which the transplanted cells are human tissue stem cells.
Rights and permissions
About this article
Cite this article
Heyer, J., Kwong, L., Lowe, S. et al. Non-germline genetically engineered mouse models for translational cancer research. Nat Rev Cancer 10, 470–480 (2010). https://doi.org/10.1038/nrc2877
Issue Date:
DOI: https://doi.org/10.1038/nrc2877
This article is cited by
-
Mitochondrial genes as strong molecular markers for species identification
The Nucleus (2023)
-
A new mouse model to study restoration of interleukin-6 (IL-6) expression in a Cre-dependent manner: microglial IL-6 regulation of experimental autoimmune encephalomyelitis
Journal of Neuroinflammation (2020)
-
A novel conditional NPM-ALK-driven model of CD30+ T-cell lymphoma mediated by a translational stop cassette
Oncogene (2020)
-
The duality of human oncoproteins: drivers of cancer and congenital disorders
Nature Reviews Cancer (2020)