Abstract
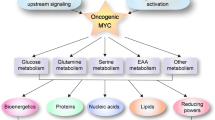
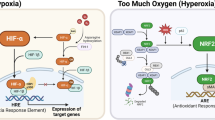
The interaction of MYC and hypoxia inducible factors (HIFs) under physiological, non-tumorigenic conditions provides insights into normal homeostatic cellular responses to low oxygen levels (hypoxia). Many tumours contain genetic alterations, such as MYC activation, that can collaborate with HIF to confer metabolic advantages to tumour cells, which tend to exist in a hypoxic microenvironment. This Perspective emphasizes the differences between the transcriptional network that operates under normal homeostatic conditions and the network in a tumorigenic milieu.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Shchors, K. & Evan, G. Tumor angiogenesis: cause or consequence of cancer? Cancer Res. 67, 7059–7061 (2007).
Adhikary, S. & Eilers, M. Transcriptional regulation and transformation by Myc proteins. Nature Rev. Mol. Cell Biol. 6, 635–645 (2005).
Gordan, J. D. & Simon, M. C. Hypoxia-inducible factors: central regulators of the tumor phenotype. Curr. Opin. Genet. Dev. 17, 71–77 (2007).
Gordan, J. D., Thompson, C. B. & Simon, M. C. HIF and c-Myc: sibling rivals for control of cancer cell metabolism and proliferation. Cancer Cell 12, 108–113 (2007).
Semenza, G. L. Targeting HIF-1 for cancer therapy. Nature Rev. Cancer 3, 721–732 (2003).
Lawlor, E. R. et al. Reversible kinetic analysis of Myc targets in vivo provides novel insights into Myc-mediated tumorigenesis. Cancer Res. 66, 4591–4601 (2006).
Dang, C. V. et al. The c-Myc target gene network. Semin. Cancer Biol. 16, 253–264 (2006).
Liu, J. & Levens, D. Making myc. Curr. Top. Microbiol. Immunol. 302, 1–32 (2006).
Chung, H. J. & Levens, D. c-myc expression: keep the noise down! Mol. Cells 20, 157–166 (2005).
Grandori, C., Cowley, S. M., James, L. P. & Eisenman, R. N. The Myc/Max/Mad network and the transcriptional control of cell behavior. Annu. Rev. Cell Dev. Biol. 16, 653–699 (2000).
Baudino, T. A. & Cleveland, J. L. The Max network gone mad. Mol. Cell Biol. 21, 691–702 (2001).
Zeller, K. I., Jegga, A. G., Aronow, B. J., O'Donnell, K. A. & Dang, C. V. An integrated database of genes responsive to the Myc oncogenic transcription factor: identification of direct genomic targets. Genome Biol. 4, R69 (2003).
Zeller, K. I. et al. Global mapping of c-Myc binding sites and target gene networks in human B cells. Proc. Natl Acad. Sci. USA 103, 17834–17839 (2006).
Schreiber-Agus, N. et al. Drosophila Myc is oncogenic in mammalian cells and plays a role in the diminutive phenotype. Proc. Natl Acad. Sci. USA 94, 1235–1240 (1997).
Gallant, P., Shiio, Y., Cheng, P. F., Parkhurst, S. M. & Eisenman, R. N. Myc and Max homologs in Drosophila. Science 274, 1523–1527 (1996).
Greasley, P. J., Bonnard, C. & Amati, B. Myc induces the nucleolin and BN51 genes: possible implications in ribosome biogenesis. Nucleic Acids Res. 28, 446–453 (2000).
Fernandez, P. C. et al. Genomic targets of the human c-Myc protein. Genes Dev. 17, 1115–1129 (2003).
Grewal, S. S., Li, L., Orian, A., Eisenman, R. N. & Edgar, B. A. Myc-dependent regulation of ribosomal RNA synthesis during Drosophila development. Nature Cell Biol. 7, 295–302 (2005).
Grandori, C. et al. c-Myc binds to human ribosomal DNA and stimulates transcription of rRNA genes by RNA polymerase I. Nature Cell Biol. 7, 311–318 (2005).
Gomez-Roman, N., Grandori, C., Eisenman, R. N. & White, R. J. Direct activation of RNA polymerase III transcription by c-Myc. Nature 421, 290–294 (2003).
Osthus, R. C. et al. Deregulation of glucose transporter 1 and glycolytic gene expression by c-Myc. J. Biol. Chem. 275, 21797–21800 (2000).
Shim, H. et al. c-Myc transactivation of LDH-A: implications for tumor metabolism and growth. Proc. Natl Acad. Sci. USA 94, 6658–6663 (1997).
Kim, J. W. et al. Evaluation of myc E-box phylogenetic footprints in glycolytic genes by chromatin immunoprecipitation assays. Mol. Cell Biol. 24, 5923–5936 (2004).
Li, F. et al. Myc stimulates nuclearly encoded mitochondrial genes and mitochondrial biogenesis. Mol. Cell Biol. 25, 6225–6234 (2005).
Zhang, H. et al. HIF-1 inhibits mitochondrial biogenesis and cellular respiration in VHL-deficient renal cell carcinoma by repression of C-MYC activity. Cancer Cell 11, 407–420 (2007).
Brahimi-Horn, M. C., Chiche, J. & Pouyssegur, J. Hypoxia signalling controls metabolic demand. Curr. Opin. Cell Biol. 19, 223–229 (2007).
Brunelle, J. K. et al. Oxygen sensing requires mitochondrial ROS but not oxidative phosphorylation. Cell Metab. 1, 409–414 (2005).
Guzy, R. D. et al. Mitochondrial complex III is required for hypoxia-induced ROS production and cellular oxygen sensing. Cell Metab. 1, 401–408 (2005).
Kaelin, W. G. Jr. ROS: really involved in oxygen sensing. Cell Metab. 1, 357–358 (2005).
Lu, H. et al. Reversible inactivation of HIF-1 prolyl hydroxylases allows cell metabolism to control basal HIF-1. J. Biol. Chem. 280, 41928–41939 (2005).
Semenza, G. L. et al. Hypoxia response elements in the aldolase A, enolase 1, and lactate dehydrogenase A gene promoters contain essential binding sites for hypoxia-inducible factor 1. J. Biol. Chem. 271, 32529–32537 (1996).
Semenza, G. L., Roth, P. H., Fang, H. M. & Wang, G. L. Transcriptional regulation of genes encoding glycolytic enzymes by hypoxia-inducible factor 1. J. Biol. Chem. 269, 23757–23763 (1994).
Papandreou, I., Cairns, R. A., Fontana, L., Lim, A. L. & Denko, N. C. HIF-1 mediates adaptation to hypoxia by actively downregulating mitochondrial oxygen consumption. Cell Metab. 3, 187–197 (2006).
Kim, J. W., Tchernyshyov, I., Semenza, G. L. & Dang, C. V. HIF-1-mediated expression of pyruvate dehydrogenase kinase: a metabolic switch required for cellular adaptation to hypoxia. Cell Metab. 3, 177–185 (2006).
Fukuda, R. et al. HIF-1 regulates cytochrome oxidase subunit composition to optimize efficiency of respiration in hypoxic cells. Cell 129, 111–122 (2007)
Goda, N. et al. Hypoxia-inducible factor 1α is essential for cell cycle arrest during hypoxia. Mol. Cell Biol. 23, 359–369 (2003).
Green, S. L., Freiberg, R. A. & Giaccia, A. J. p21(Cip1) and p27(Kip1) regulate cell cycle reentry after hypoxic stress but are not necessary for hypoxia-induced arrest. Mol. Cell Biol. 21, 1196–1206 (2001).
Gardner, L. B. et al. Hypoxia inhibits G1/S transition through regulation of p27 expression. J. Biol. Chem. 276, 7919–7926 (2001).
Koshiji, M. et al. HIF-1α induces cell cycle arrest by functionally counteracting Myc. EMBO J. 23, 1949–1956 (2004).
Kleine-Kohlbrecher, D., Adhikary, S. & Eilers, M. Mechanisms of transcriptional repression by Myc. Curr. Top. Microbiol. Immunol. 302, 51–62 (2006).
Koshiji, M. et al. HIF-1α induces genetic instability by transcriptionally downregulating MutSα expression. Mol. Cell 17, 793–803 (2005).
To, K. K., Sedelnikova, O. A., Samons, M., Bonner, W. M. & Huang, L. E. The phosphorylation status of PAS-B distinguishes HIF-1α from HIF-2α in NBS1 repression. EMBO J. 25, 4784–4794 (2006).
Vafa, O. et al. c-Myc can induce DNA damage, increase reactive oxygen species, and mitigate p53 function: a mechanism for oncogene-induced genetic instability. Mol. Cell 9, 1031–1044 (2002).
Karlsson, A. et al. Genomically complex lymphomas undergo sustained tumor regression upon MYC inactivation unless they acquire novel chromosomal translocations. Blood 101, 2797–2803 (2003).
Gardner, L. B., Li, F., Yang, X. & Dang, C. V. Anoxic fibroblasts activate a replication checkpoint that is bypassed by E1a. Mol. Cell Biol. 23, 9032–9045 (2003).
Corn, P. G. et al. Mxi1 is induced by hypoxia in a HIF-1-dependent manner and protects cells from c-Myc-induced apoptosis. Cancer Biol. Ther. 4, 1285–1294 (2005).
O'Hagan, R. C. et al. Gene-target recognition among members of the myc superfamily and implications for oncogenesis. Nature Genet. 24, 113–119 (2000).
Gordan, J. D., Bertout, J. A., Hu, C. J., Diehl, J. A. & Simon, M. C. HIF-2α promotes hypoxic cell proliferation by enhancing c-Myc transcriptional activity. Cancer Cell 11, 335–347 (2007).
Tian, H., McKnight, S. L. & Russell, D. W. Endothelial PAS domain protein 1 (EPAS1), a transcription factor selectively expressed in endothelial cells. Genes Dev. 11, 72–82 (1997).
Dang, C. V. c-Myc target genes involved in cell growth, apoptosis, and metabolism. Mol. Cell Biol. 19, 1–11 (1999).
Oster, S. K., Ho, C. S., Soucie, E. L. & Penn, L. Z. The myc oncogene: MarvelouslY Complex. Adv. Cancer Res. 84, 81–154 (2002).
Smith, D. P., Bath, M. L., Metcalf, D., Harris, A. W. & Cory, S. MYC levels govern hematopoietic tumor type and latency in transgenic mice. Blood 108, 653–661 (2006).
Arvanitis, C. & Felsher, D. W. Conditional transgenic models define how MYC initiates and maintains tumorigenesis. Semin. Cancer Biol. 16, 313–317 (2006).
Alarcon, R. M., Rupnow, B. A., Graeber, T. G., Knox, S. J. & Giaccia, A. J. Modulation of c-Myc activity and apoptosis in vivo. Cancer Res. 56, 4315–4319 (1996).
Rupnow, B. A., Alarcon, R. M., Giaccia, A. J. & Knox, S. J. p53 mediates apoptosis induced by c-Myc activation in hypoxic or gamma irradiated fibroblasts. Cell Death Differ. 5, 141–147 (1998).
Cleveland, J. L. & Sherr, C. J. Antagonism of Myc functions by Arf. Cancer Cell 6, 309–311 (2004).
Gao, P. et al. HIF-dependent anti-tumorigenic effect of anti-oxidants in vivo. Cancer Cell 12, 230–238 (2007).
Blouw, B. et al. The hypoxic response of tumors is dependent on their microenvironment. Cancer Cell 4, 133–146 (2003).
Ryan, H. E. et al. Hypoxia-inducible factor-1α is a positive factor in solid tumor growth. Cancer Res. 60, 4010–4015 (2000).
Kim, J. W., Gao, P., Liu, Y. C., Semenza, G. L. & Dang, C. V. HIF-1 and dysregulated c-Myc cooperatively induces VEGF and metabolic switches, HK2 and PDK1. Mol. Cell Biol. (2007).
Warburg, O. The Metabolism of Tumours (Constable & Co., London, 1930).
Warburg, O. On the origin of cancer cells. Science 123, 309–314 (1956).
Helmlinger, G., Yuan, F., Dellian, M. & Jain, R. K. Interstitial pH and pO2 gradients in solid tumors in vivo: high-resolution measurements reveal a lack of correlation. Nature Med. 3, 177–182 (1997).
Plas, D. R. & Thompson, C. B. Akt-dependent transformation: there is more to growth than just surviving. Oncogene 24, 7435–7442 (2005).
Kim, J. W. & Dang, C. V. Cancer's molecular sweet tooth and the Warburg effect. Cancer Res. 66, 8927–8930 (2006).
Dewhirst, M. W. Intermittent hypoxia furthers the rationale for hypoxia-inducible factor-1 targeting. Cancer Res. 67, 854–855 (2007).
Gazit, Y. et al. Fractal characteristics of tumor vascular architecture during tumor growth and regression. Microcirculation 4, 395–402 (1997).
Gatenby, R. A. & Gillies, R. J. Why do cancers have high aerobic glycolysis? Nature Rev. Cancer 4, 891–899 (2004).
Tatsumi, M., Cohade, C., Nakamoto, Y., Fishman, E. K. & Wahl, R. L. Direct comparison of FDG PET and CT findings in patients with lymphoma: initial experience. Radiology 237, 1038–1045 (2005).
Kim, J. W. & Dang, C. V. Multifaceted roles of glycolytic enzymes. Trends Biochem. Sci. 30, 142–150 (2005).
Bonnet, S. et al. A mitochondria-K+ channel axis is suppressed in cancer and its normalization promotes apoptosis and inhibits cancer growth. Cancer Cell 11, 37–51 (2007).
Fantin, V. R., St-Pierre, J. & Leder, P. Attenuation of LDH-A expression uncovers a link between glycolysis, mitochondrial physiology, and tumor maintenance. Cancer Cell 9, 425–434 (2006).
O'Donnell, K. A. et al. Activation of transferrin receptor 1 by c-Myc enhances cellular proliferation and tumorigenesis. Mol. Cell Biol. 26, 2373–2386 (2006).
Tacchini, L., Bianchi, L., Bernelli-Zazzera, A. & Cairo, G. Transferrin receptor induction by hypoxia. HIF-1-mediated transcriptional activation and cell-specific post-transcriptional regulation. J. Biol. Chem. 274, 24142–24146 (1999).
Pandey, M. K. et al. Gambogic acid, a novel ligand for transferrin receptor, potentiates TNF-induced apoptosis through modulation of the nuclear factor-κB signaling pathway. Blood 110, 3517–3525 (2007).
Kasibhatla, S. et al. A role for transferrin receptor in triggering apoptosis when targeted with gambogic acid. Proc. Natl Acad. Sci. USA 102, 12095–12100 (2005).
Louro, I. D. et al. Comparative gene expression profile analysis of GLI and c-MYC in an epithelial model of malignant transformation. Cancer Res. 62, 5867–5873 (2002).
Author information
Authors and Affiliations
Corresponding author
Related links
Rights and permissions
About this article
Cite this article
Dang, C., Kim, Jw., Gao, P. et al. The interplay between MYC and HIF in cancer. Nat Rev Cancer 8, 51–56 (2008). https://doi.org/10.1038/nrc2274
Issue Date:
DOI: https://doi.org/10.1038/nrc2274
This article is cited by
-
Mechanosensitive channel of large conductance enhances the mechanical stretching-induced upregulation of glycolysis and oxidative metabolism in Schwann cells
Cell Communication and Signaling (2024)
-
Transcriptional regulation and post-translational modifications in the glycolytic pathway for targeted cancer therapy
Acta Pharmacologica Sinica (2024)
-
An NF-κB/OVOL2 circuit regulates glucose import and cell survival in non-small cell lung cancer
Cell Communication and Signaling (2022)
-
Histone H2AX promotes metastatic progression by preserving glycolysis via hexokinase-2
Scientific Reports (2022)
-
USP25 promotes pathological HIF-1-driven metabolic reprogramming and is a potential therapeutic target in pancreatic cancer
Nature Communications (2022)