Abstract
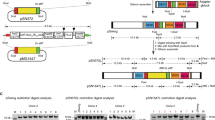
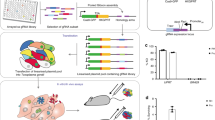
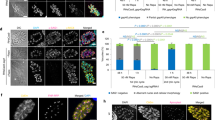
Apicomplexan parasites, such as Toxoplasma gondii, cause extensive morbidity and mortality in humans and livestock, highlighting the need for a deeper understanding of their molecular biology. Although techniques for the generation of targeted gene disruptions have long been available for apicomplexans, such methods are not readily scalable to the entire genome. We recently used CRISPR-Cas9 to disrupt all nuclear protein–coding genes in T. gondii using a pooled format. The method relies on transfection of a guide RNA library into parasites constitutively expressing Cas9. Here, we present the complete workflow of such a screen, including preparation of the guide RNA library, growth and testing of the recipient strain, generation of the mutant population, culture conditions for the screen, preparation of genomic DNA libraries, next-generation sequencing of the guide RNA loci, and analysis to detect fitness-conferring genes. This method can be deployed to study how culture conditions affect the repertoire of genes needed by parasites, which will enable studies of their metabolic needs, host specificity, and drug-resistance mechanisms. In addition, by manipulating the background in which the screen is performed, researchers will be able to investigate genetic interactions, which may help uncover redundancy or epistasis in the parasite genome. Using this method, a genome-wide screen and its analysis can be completed in 3 weeks, after ∼1 month of preparation to generate the library and grow the cells needed, making it a powerful tool for uncovering functionally important genes in apicomplexan parasites.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Accession codes
References
World Health Organization. World Malaria Report http://www.who.int/malaria/publications/world-malaria-report-2015/report/en/ Date: (2015).
Checkley, W. et al. A review of the global burden, novel diagnostics, therapeutics, and vaccine targets for Cryptosporidium. Lancet Infect. Dis. 15, 85–94 (2015).
Torgerson, P.R. & Mastroiacovo, P. The global burden of congenital toxoplasmosis: a systematic review. Bull. World Health Organ. 91, 501–508 (2013).
Animal and Plant Health Inspection Service of the USDA. Toxoplasma on US Sheep Operations https://www.aphis.usda.gov/animal_health/nahms/sheep/downloads/sheep11/Sheep11_is_Toxo.pdf (2014).
Garrison, E. et al. A forward genetic screen reveals that calcium-dependent protein kinase 3 regulates egress in toxoplasma. PLoS Pathog. 8, e1003049 (2012).
Su, C., Howe, D.K., Dubey, J.P., Ajioka, J.W. & Sibley, L.D. Identification of quantitative trait loci controlling acute virulence in Toxoplasma gondii. Proc. Natl. Acad. Sci. USA 99, 10753–10758 (2002).
Gubbels, M.-J. et al. Forward genetic analysis of the apicomplexan cell division cycle in Toxoplasma gondii. PLoS Pathog. 4, e36 (2008).
Singh, U., Brewer, J.L. & Boothroyd, J.C. Genetic analysis of tachyzoite to bradyzoite differentiation mutants in Toxoplasma gondii reveals a hierarchy of gene induction. Mol. Microbiol. 44, 721–733 (2002).
Pfefferkorn, E.R., Borotz, S.E. & Nothnagel, R.F. Toxoplasma gondii: characterization of a mutant resistant to sulfonamides. Exp. Parasitol. 74, 261–270 (1992).
Pfefferkorn, E.R. & Kasper, L.H. Toxoplasma gondii: genetic crosses reveal phenotypic suppression of hydroxyurea resistance by fluorodeoxyuridine resistance. Exp. Parasitol. 55, 207–218 (1983).
Dubey, J.P. & Frenkel, J.K. Cyst-induced toxoplasmosis in cats. J. Protozool. 19, 155–177 (1972).
Behnke, M.S., Dubey, J.P. & Sibley, L.D. Genetic mapping of pathogenesis determinants in Toxoplasma gondii. Annu. Rev. Microbiol. 70, 63–81 (2016).
Pfefferkorn, L.C. & Pfefferkorn, E.R. Toxoplasma gondii: genetic recombination between drug resistant mutants. Exp. Parasitol. 50, 305–316 (1980).
Mojica, F.J.M., Díez-Villaseñor, C., García-Martínez, J. & Soria, E. Intervening sequences of regularly spaced prokaryotic repeats derive from foreign genetic elements. J. Mol. Evol. 60, 174–182 (2005).
Ran, F.A. et al. Genome engineering using the CRISPR-Cas9 system. Nat. Protoc. 8, 2281–2308 (2013).
Jinek, M. et al. A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science 337, 816–821 (2012).
Gasiunas, G., Barrangou, R., Horvath, P. & Siksnys, V. Cas9-crRNA ribonucleoprotein complex mediates specific DNA cleavage for adaptive immunity in bacteria. Proc. Natl. Acad. Sci. USA 109, E2579–86 (2012).
Bolotin, A., Quinquis, B., Sorokin, A. & Ehrlich, S.D. Clustered regularly interspaced short palindrome repeats (CRISPRs) have spacers of extrachromosomal origin. Microbiology 151, 2551–2561 (2005).
Marraffini, L.A. & Sontheimer, E.J. CRISPR interference limits horizontal gene transfer in staphylococci by targeting DNA. Science 322, 1843–1845 (2008).
Hsu, P.D., Lander, E.S. & Zhang, F. Development and applications of CRISPR-Cas9 for genome engineering. Cell 157, 1262–1278 (2014).
Doudna, J.A. & Charpentier, E. Genome editing. The new frontier of genome engineering with CRISPR-Cas9. Science 346, 1258096–1258096 (2014).
Wang, T., Wei, J.J., Sabatini, D.M. & Lander, E.S. Genetic screens in human cells using the CRISPR-Cas9 system. Science 343, 80–84 (2014).
Shalem, O. et al. Genome-scale CRISPR-Cas9 knockout screening in human cells. Science 343, 84–87 (2014).
Sidik, S.M. et al. A genome-wide CRISPR screen in toxoplasma identifies essential apicomplexan genes. Cell 166, 1423–1435.e12 (2016).
Sidik, S.M., Hackett, C.G., Tran, F., Westwood, N.J. & Lourido, S. Efficient genome engineering of Toxoplasma gondii using CRISPR/Cas9. PLoS One 9, e100450 (2014).
Shen, B., Brown, K.M., Lee, T.D. & Sibley, L.D. Efficient gene disruption in diverse strains of Toxoplasma gondii using CRISPR/CAS9. MBio 5, e01114–14 (2014).
Wagner, J.C., Platt, R.J., Goldfless, S.J., Zhang, F. & Niles, J.C. Efficient CRISPR-Cas9-mediated genome editing in Plasmodium falciparum. Nat. Methods 11, 915–918 (2014).
Ghorbal, M. et al. Genome editing in the human malaria parasite Plasmodium falciparum using the CRISPR-Cas9 system. Nat. Biotechnol. 32, 819–821 (2014).
Vinayak, S. et al. Genetic modification of the diarrhoeal pathogen Cryptosporidium parvum. Nature 523, 477–480 (2015).
Weiss, L.M. & Kim, K. Toxoplasma gondii (Academic Press, 2014).
Donald, R.G. & Roos, D.S. Insertional mutagenesis and marker rescue in a protozoan parasite: cloning of the uracil phosphoribosyltransferase locus from Toxoplasma gondii. Proc. Natl. Acad. Sci. USA 92, 5749–5753 (1995).
Pfefferkorn, E.R. & Pfefferkorn, L.C. Toxoplasma gondii: characterization of a mutant resistant to 5-fluorodeoxyuridine. Exp. Parasitol. 42, 44–55 (1977).
Gilbert, L.A. et al. CRISPR-mediated modular RNA-guided regulation of transcription in eukaryotes. Cell 154, 442–451 (2013).
Gilbert, L.A. et al. Genome-scale CRISPR-mediated control of gene repression and activation. Cell 159, 647–661 (2014).
Bushell, E. et al. Functional profiling of a Plasmodium genome reveals an abundance of essential genes. Cell 170, 260–272.e8 (2017).
Chen, S. et al. Genome-wide CRISPR screen in a mouse model of tumor growth and metastasis. Cell 160, 1246–1260 (2015).
Hart, T. et al. High-resolution CRISPR screens reveal fitness genes and genotype-specific cancer liabilities. Cell 163, 1515–1526 (2015).
Pettitt, S., Krastev, D.B., Song, F., Ashworth, A. & Lord, C.J. Abstract 2743: finding determinants of PARP inhibitor sensitivity using genome-wide and focused CRISPR screens. Cancer Res. 76, 2743–2743 (2016).
Steinhart, Z. et al. Genome-wide CRISPR screens reveal a Wnt-FZD5 signaling circuit as a druggable vulnerability of RNF43-mutant pancreatic tumors. Nat. Med. 23, 60–68 (2017).
Parnas, O. et al. A genome-wide CRISPR screen in primary immune cells to dissect regulatory networks. Cell 162, 675–686 (2015).
Marceau, C.D. et al. Genetic dissection of Flaviviridae host factors through genome-scale CRISPR screens. Nature 535, 159–163 (2016).
Peters, J.M. et al. A comprehensive, CRISPR-based functional analysis of essential genes in bacteria. Cell 165, 1493–1506 (2016).
Tsai, S.Q. et al. GUIDE-seq enables genome-wide profiling of off-target cleavage by CRISPR-Cas nucleases. Nat. Biotechnol. 33, 187–197 (2015).
Doench, J.G. et al. Optimized sgRNA design to maximize activity and minimize off-target effects of CRISPR-Cas9. Nat. Biotechnol. 34, 184–191 (2016).
Kleinstiver, B.P. et al. High-fidelity CRISPR-Cas9 nucleases with no detectable genome-wide off-target effects. Nature 529, 490–495 (2016).
Slaymaker, I.M. et al. Rationally engineered Cas9 nucleases with improved specificity. Science 351, 84–88 (2016).
Wang, T. et al. Identification and characterization of essential genes in the human genome. Science 350, 1096–1101 (2015).
Tzelepis, K. et al. A CRISPR dropout screen identifies genetic vulnerabilities and therapeutic targets in acute myeloid leukemia. Cell Rep. 17, 1193–1205 (2016).
Gibson, D.G. et al. Enzymatic assembly of DNA molecules up to several hundred kilobases. Nat. Methods 6, 343–345 (2009).
Winter, J. et al. CRISPRAnalyzeR: interactive analysis, annotation and documentation of pooled CRISPR screens. Preprint at bioRxiv, https://www.biorxiv.org/content/early/2017/02/20/109967 (2017).
Drewry, L.L. & Sibley, L.D. Toxoplasma actin is required for efficient host cell invasion. MBio 6, e00557 (2015).
Burg, J.L., Perelman, D., Kasper, L.H., Ware, P.L. & Boothroyd, J.C. Molecular analysis of the gene encoding the major surface antigen of Toxoplasma gondii. J. Immunol. 141, 3584–3591 (1988).
Acknowledgements
We thank J.P.J. Saeij and T. Wang for helpful advice during the development of the protocol, G. Bell and P. Thiru for assistance with bioinformatics, and E. Shortt for helpful comments during the preparation of the manuscript. Antibodies against SAG1 and ACT1 were kindly provided by L.D. Sibley (Washington University). This work was supported by the NIH Director's Early Independence Award (1DP5OD017892) and an NIH Exploratory R21 grant (1R21AI123746) to S.L.
Author information
Authors and Affiliations
Contributions
S.M.S., D.H., and S.L. designed and performed the experiments. S.M.S. wrote the manuscript, which was edited and revised in collaboration with D.H. and S.L.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Methods
Detailed protocols outlining the procedures for cloning an sgRNA pool into the pU6-DHFR plasmid, library transformation and preparation, and confirmation of Cas9 expression. (PDF 270 kb)
Rights and permissions
About this article
Cite this article
Sidik, S., Huet, D. & Lourido, S. CRISPR-Cas9-based genome-wide screening of Toxoplasma gondii. Nat Protoc 13, 307–323 (2018). https://doi.org/10.1038/nprot.2017.131
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2017.131
This article is cited by
-
A splitCas9 phenotypic screen in Toxoplasma gondii identifies proteins involved in host cell egress and invasion
Nature Microbiology (2022)
-
Efficient CRISPR/Cas9 genome editing in a salmonid fish cell line using a lentivirus delivery system
BMC Biotechnology (2020)
-
Genetic screens reveal a central role for heme metabolism in artemisinin susceptibility
Nature Communications (2020)
-
Genome-wide screens identify Toxoplasma gondii determinants of parasite fitness in IFNγ-activated murine macrophages
Nature Communications (2020)
-
One gene to rule them all in a chronic brain infection
Nature (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.