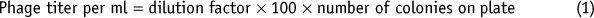Abstract
Directed evolution is a powerful tool to improve the characteristics of biomolecules. Here we present a protocol for the intracellular evolution of proteins with distinct differences and advantages in comparison with established techniques. These include the ability to select for a particular function from a library of protein variants inside cells, minimizing undesired coevolution and propagation of nonfunctional library members, as well as allowing positive and negative selection logics using basally active promoters. A typical evolution experiment comprises the following stages: (i) preparation of a combinatorial M13 phagemid (PM) library expressing variants of the gene of interest (GOI) and preparation of the Escherichia coli host cells; (ii) multiple rounds of an intracellular selection process toward a desired activity; and (iii) the characterization of the evolved target proteins. The system has been developed for the selection of new orthogonal transcription factors (TFs) but is capable of evolving any gene—or gene circuit function—that can be linked to conditional M13 phage replication. Here we demonstrate our approach using as an example the directed evolution of the bacteriophage λ cI TF against two synthetic bidirectional promoters. The evolved TF variants enable simultaneous activation and repression against their engineered promoters and do not cross-react with the wild-type promoter, thus ensuring orthogonality. This protocol requires no special equipment, allowing synthetic biologists and general users to evolve improved biomolecules within ∼7 weeks.
Similar content being viewed by others
Introduction
Directed evolution has emerged as a powerful tool to improve the characteristics of biomolecules1,2,3. The approach mimics natural selection to evolve biomolecules toward a desired activity4. One efficient and commonly used strategy to achieve this in a laboratory environment is to use filamentous bacteriophages such as M13 to link a mutable genotype to a selectable phenotype. In this way, a number of M13-phage-assisted methods, such as the widely used phage display technology5, have been developed and applied to improve a wide variety of proteins, including antibodies6,7,8, DNA-binding proteins9,10 and enzymes11,12. These systems are characterized by an extracellular (in vitro) or intracellular (in vivo) mode of operation. In vitro systems are generally easier to engineer in terms of selection stringency adjustments13, but possess certain limitations that can only be overcome by applying intracellular processes. For example, selection from combinatorial libraries in vivo ensures compatibility with the host cell machinery. This facilitates the optimization of synthetic proteins and gene circuits14,15,16, which ultimately have to function in a host cell context. In vivo methods promote selection for orthogonality17,18—a lack of cross-reactions—by intrinsically counterselecting against adverse effects inside the cell. To further broaden the applications of in vivo directed evolution, we recently developed an M13-phage-based method19 for the intracellular selection of proteins from combinatorial libraries with distinct differences and advantages to established techniques.
Overview of the protocol
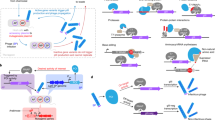
This protocol describes a general approach for the directed evolution of proteins from combinatorial libraries on PMs (Fig. 1). The selection process takes place inside E. coli cells by linking the target protein's activity to conditional phage production, thus allowing enrichment of functional library members. This is exemplified here by the directed evolution of orthogonal dual TFs based on bacteriophage λ cI variants19, selecting against synthetic promoters. However, the method can be readily adapted for other target biomolecules (“Applications of the method”). A typical evolution experiment consists of the following: (i) the preparation of a combinatorial M13 phage library (Steps 1–38) and E. coli host cells (Steps 39–47); (ii) the selection process toward a desired activity (Steps 48–61); and (iii) the characterization of the selected target proteins (Steps 62–68) (Fig. 2).
(1) Phagemid library generation: a combinatorial DNA library is generated from the target gene on a phagemid (PM) that also contains conditionally expressed M13 gene III and the M13 packaging signal. The DNA library members are then packaged into phage particles, which are the starting point for selection. (2) Selection cycles: TG1 cells containing a modified helper phage (HP) and an accessory plasmid (AP) are infected with the constructed phage library. The HP provides all that is required for phage propagation, except for two essential genes (gIII and gVI). The AP contains a conditional gene circuit that links the target protein's activity to conditional phage production (gene VI expression). Enrichment for a particular protein function occurs after several rounds of selection. (3) Gene isolation: cells are infected with selected phages, and the phagemid DNA is amplified and purified. (4) Protein characterization: the target protein's activity needs to be analyzed with a suitable reporter assay.
The system is based on E. coli cultures and three compatible plasmids (available from Addgene; see MATERIALS). Together, these conditionally produce phage (containing the evolving gene) in correlation to the activity of a library member. A selection experiment always begins with an E. coli culture that contains the first two plasmids: a modified helper phage plasmid (HP) and an accessory plasmid (AP) (Fig. 3a). The HP provides almost all that is needed for phage propagation, except for two essential genes (gIII and gVI). Furthermore, the weak M13 packaging signal (PS) is removed from the original M13KO7 HP to obtain the final M13KO7-ΔPS-ΔgIII-ΔgVI HP. The second plasmid, AP, contains a conditional gene circuit that links an inducible input (e.g., a promoter with a novel operator) to gVI expression. The evolving gene or gene circuit is placed on the third plasmid, termed a PM, which is packaged into an infectious phage particle only when all phage genes are expressed. The PM contains the second missing gene (gIII) and a combinatorially randomized GOI and is provided to the E. coli culture in the form of an infectious phage library (Fig. 3b). Crucially, our system moves Gene III onto the PM so that phage replication occurs only after initial infection, thus circumventing infection resistance20,21 and decreasing the chances of propagating nonfunctional library members because of multiple infections. A GOI with the desired characteristics upregulates gene VI expression on the AP, completing the phage life cycle. For example, a randomized TF library member that activates an artificial promoter upstream of gVI will increase its own phage production (Fig. 4a). In this way, a protein with novel desired properties can be selected after several rounds of reinfection.
(a) Plasmids needed to set up the phagemid-based selection system. The modified helper phage HP (M13KO7-ΔPS-ΔgIII-ΔgVI) contains the kanamycin-resistance gene (KanR) and all phage genes required for phage replication, except genes III and VI. The weak packaging signal (PS) is removed to prevent helper phage propagation. The accessory plasmid AP contains the chloramphenicol-resistance gene (CamR) and a conditional gene VI expression circuit, induced by an active library member on the phagemid (PM). The PM also provides the ampicillin-resistance gene (AmpR), the M13 packaging signal (PS; to allow DNA packaging in phage) and the constitutively expressed gene III. (b) The intracellular selection process. An active library member on the packaged PM induces gene VI expression to complete the phage life cycle, thus enriching this variant over time.
(a) An evolving transcription factor19 (e.g., λ cI) activates gene VI expression downstream of a specific promoter (e.g., λ PRM). In this protocol, as an example of this we selected new activators against engineered synthetic promoters. (b) An evolving RNA polymerase22 (gray) enables transcription and hence gene VI expression. (c) An evolving DNA-binding protein (red) derived from genome-editing systems (transcription-activator-like effector nucleases (TALENs))10 is linked to the ω subunit of bacterial RNA polymerase III (gray). Binding to a target DNA sequence (dark blue) upstream of a minimal lac promoter (black) induces transcription of gene VI. (d) The target protein (dark blue) is bound to the DNA upstream of the promoter PlacZ-opt (black) via a fused DNA-binding domain (orange), and the RNA polymerase ω subunit (RpoZ; yellow) is fused to the evolving protein (red). Target protein binding of the evolving protein24 enables the localization of RNA polymerase upstream of gene VI, initiating gene expression from the PlacZ-opt promoter. (e) The T7 polymerase (gray) is inhibited when bound to T7 lysozyme (dark blue), as it inhibits transcription initiation and the transition from initiation to elongation44. Proteolysis of the target cleavage site (red) by an evolving protease23 activates the T7 RNA polymerase, enabling gene VI expression downstream of the T7 promoter. Gene VI is annotated with an asterisk where originally conditional gene III was used instead of gene VI with PACE10,22,23,24. Gene III and gene VI are both minor coat proteins, each present in three to five copies per phage particle21.
Applications of the method
The method has been used to evolve a set of dual activator–repressor switches for orthogonal logic gates, based on bacteriophage λ cI variants, and multi-input promoter architectures, and these switches have been successfully applied in downstream synthetic gene circuits19. In general, the method is capable of evolving any gene—or gene circuit function—on the PM that can be linked to pVI production. This is analogous to previous uses of phage-assisted continuous evolution (PACE)22 (Fig. 4). With PACE, a wide range of medically and biotechnologically relevant biomolecules, including polymerases22, proteases23 and genome-editing proteins10, as well as protein–protein interactions24, were linked to conditional M13 phage propagation. In principle, any application in which directed evolution approaches have been proposed (e.g., biosensors25 or hybrids with chemical evolution26) can be adapted to this method if the target protein's activity can be linked to conditional M13 phage production. However, certain applications (e.g., those involving membrane proteins) would be harder to adapt, which is why other methods such as liposome display27 have been developed.
Comparison with other methods
Several bacterial directed evolution methods have been developed based on phage replication22, display technologies5,27,28,29, genome engineering30, and conditional cell growth31,32. Linking a target protein's activity to cell growth is a widely used strategy and is particularly suitable when the evolving gene directly improves cellular fitness33,34. The use of bacteriophage offers a convenient way to uncouple the fitness function of a cell with target protein activity. This is achieved by linking a target gene's activity to phage replication using a conditional gene circuit. The main advantage of conditional phage production over display technologies is the compatibility of target genes or gene circuits with the host cell machinery, as these have to function in a host cell context. In contrast to PACE (which uses gIII as the sole conditional gene), our PM-based approach facilitates the selection of large combinatorial libraries and enables positive and negative selection logics using promoters with basal gene expression. Our system also minimizes the undesired coevolution of phage genes, as only the packaged PM is evolving and not the HP itself. In comparison with PACE, the protocol is performed in batch mode and therefore requires no special equipment for reactor assembly, instead relying on daily researcher intervention during selections. Moreover, the batch process facilitates the performance of multiple selections in parallel, enabling the scalability of each individual selection and easy handling. Continuous culture evolution systems can suffer from 'phage washout' (loss of phage) when conditional phage production rates are not compatible with the flow rates. By contrast, batch modes are not as sensitive to loss of phage. On the other hand, dozens of rounds of reinfections occur in a single day of PACE, whereas our system is currently limited to one round per overnight cycle. In addition, combinatorial libraries have to be designed and cloned because, unlike PACE, our system does not include a random mutagenesis plasmid35. This means that structural information or a partial understanding of how a set of amino acid changes will affect the target protein's activity is required to run our system.
Limitations of the phagemid-based system
The main limitation of the system is the combinatorial size of the library, which is linked to transformation efficiency (106–1010 variants)36. The selection process itself is not limited to a certain number of gene variants, but it has to be noted that the use of larger libraries comes with the cost of prolonged experiment times. Another limitation can be the linkage of the target protein's activity to conditional M13 phage replication, as this depends on the individual protein's characteristics. This is certainly more complicated for complex proteins such as membrane proteins than it is for cytosolic proteins. Furthermore, general limitations of bacterial expression over mammalian expression (e.g., protein solubility, disulfide bonds, post-translational modifications) need to be considered for individual target proteins. For instance, our system would need to be adapted to enable the selection of proteins that require disulfide bonds for proper folding in bacterial cells37.
Experimental design
Combinatorial library cloning on phagemid. Choosing which positions to randomize in the protein of interest is a critical step, as this affects the library size, the cloning strategy and ultimately the overall selection results. Small libraries with only one or two randomized positions can easily be obtained by round-the-world PCR, whereas bigger libraries require overlap extension PCR or end-to-end ligation36. Round-the-world PCR means, in this context, that single-base-pair mutations are inserted into the target region by amplification of the whole plasmid DNA with randomized primers so that no additional step for plasmid ligation is required. For round-the-world PCR, both randomized primers must contain the mutations and bind to the same DNA sequence on opposite strands of the plasmid. Primers are generally 30–60 nucleotides long (N) and contain mutations in the middle of the randomized primers, flanked with 15–20 bases of correct sequence on both sides. These primers should ideally have a minimum GC content of 40% (%GC), end with one or more C or G bases and are purified by PAGE. The annealing region should have a melting temperature (Tm) of ≥78 °C, as determined via the following formula: Tm = 81.5 + 0.41(%GC) – 675/N – %mismatch. In this protocol, we focus on an overlap PCR approach before Gibson assembly38, as this has been our method of choice for building λ cIopt libraries with a combinatorial space of >106 variants (Fig. 5a). These libraries are based on a λ cI optimized mutant (cIopt) with a strong activation region39. The protocol presented here is optimized for the construction of combinatorial libraries by Gibson assembly. It is our method of choice because it bypasses the need for restriction sites inside target genes, which makes it much easier to construct sequence-targeted libraries. However, the selection system itself is compatible with any other library generation method36,40 as long as our PM vector backbone is used. The design of randomized oligonucleotides for overlap PCR is similar to conventional Gibson primer design38. Briefly, PCR primers for insert amplification require a 15- to 25-bp overlap with each other, as well as a 15- to 25-bp overlap with the amplified PM vector backbone. Randomized positions should be avoided in the annealing regions, and primers should ideally have a Tm of 50–60 °C as determined via the following formula: Tm = 4(G + C) + 2(A + T) (where A, C, G and T are the numbers of each base in the primer). The temperature difference of the primer pairs should be matched and lie within a 5 °C range. The maximum insert size is limited by oligonucleotide synthesis (currently ∼120 bp; desalted oligonucleotides are sufficiently pure). To evolve a novel protein, the user should ideally start with a crystal structure of the target molecule (if available) and randomize positions known to affect the desired activity (e.g., change positions of the binding interface to alter the protein binding interaction). In other cases, biochemical information might also be sufficient to guide library construction.
(a) Combinatorial library construction on phagemids (PMs). Randomized oligonucleotides require a 15- to 25-bp overlap with each other, as well as a 15- to 25-bp overlap with the amplified vector backbone, and are fused by PCR. Randomized positions are marked with an 'x' and must be avoided within overlap regions. Primers 3 and 4 bind upstream and downstream of the randomized target region and are used for vector linearization. (b) Accessory plasmid (AP) design. The conditional gene circuit that links an inducible input to gene VI expression has to be adapted for individual needs. This is achieved by replacement of the λ PRM promoter with a different promoter or inducible input depending on the desired application. For example, an engineered promoter (e.g., PM,5G6G) is constructed by overlap extension PCR and inserted into the linearized fragment by Gibson assembly. Primers 3 and 4 bind upstream and downstream of λ PRM and are used to remove the PRM promoter. (c) Reporter plasmid (RP) design. The bidirectional λ PR/PRM is replaced by the same inducible input used on the AP (e.g., P/PM,5G6G). The fluorescent proteins mCherry and GFP on the RP are used to characterize the activity of the selected proteins on the PM. Note that the maximum insert size in overlap extension PCR is limited by oligonucleotide synthesis (currently ∼120 bp).
Accessory plasmid design. The conditional gene circuit that links an inducible input to gVI expression has to be adapted to individual needs. This is achieved by replacement of the λ PRM promoter (pJPC12-ΔPS-PRM-B0034-gVI) with a different promoter or inducible input depending on the desired application (Fig. 5b). The bidirectional promoter PR/PRM consists of three operator sites (O1, O2 and O3), where λ cI binding to O1–O2 leads to PRM activation41. Counterselection via repression is achieved by placement of a specific DNA sequence at operator position O3, which is located between the −35 and −10 regions. For example, the O3 site of the PRM promoter can be replaced with the consensus wild-type (WT) sequence, OCS. Thus, binding of a cIopt library member to O1–O2 of an engineered PM promoter activates gene VI expression (and so promotes selection), whereas simultaneous binding to WT O3 represses gene VI, enabling counterselection against unwanted WT activity. We chose positive and negative selections against the synthetic promoters PM,5G6G and PM,5T6T as examples for this protocol (Supplementary Fig. 1). The engineered promoters are designated according to the positions of the base substitutions in the consensus half-site of O1 and O2.
Reporter plasmid design. This protocol describes the downstream functional characterization of evolved TFs by fluorescence analysis. It has to be noted that a suitable reporter assay needs to be adapted to the target protein's properties according to the user's needs. For this, the bidirectional PR/PRM promoter on the reporter plasmid (RP) (pJPC12-ΔPS-mCherry-PR/PRM-GFP) has to be replaced by the same inducible input used on the AP for selection (Fig. 5c). The insertion of the bidirectional promoters P/PM,5G6G and P/PM,5T6T into the RP is depicted as an example for this protocol (Supplementary Fig. 2). For other target proteins, it might be sufficient to use one of the two reporters to analyze the activity of the selected proteins.
Control selections. Enrichment assays can be performed to test the efficiency of the selection process. Mix plasmids containing λ cIopt (Addgene plasmid ID 80852) and one of the orthogonal cI variants (e.g., cI5G6G,P; Addgene plasmid ID 80861) in different ratios (e.g., 10−3 and 10−6). Then transform these into TOP10 cells with the modified HP M13KO7-ΔPS-ΔgeneIII-ΔgeneVI and the AP pJPC12-ΔPS-PRM-B0034-geneVI. This will allow the production of a phage stock packaged with cIopt and cI5G6G,P (Steps 23–38). Use the obtained phage population and run a batch selection using the AP pJPC12-ΔPS-PRM-B0034-geneVI (Steps 41–61). One can monitor the enrichment of λ cIopt by infecting TG1 cells (containing the plasmid pJPC12-ΔPS-mCherry-PR/PRM-GFP; Addgene plasmid ID 80859) with the phage titer obtained after each round of selection. Streak out infected cells on agar plates supplemented with chloramphenicol and ampicillin and grow them overnight at 37 °C. The next day, analyze the plates under the UV light of a gel documentation system. The ratio of green to red colonies should increase over time because the nonactive TF cI5G6G,P results in red colonies, whereas the enriched active cIopt leads to green colonies because of GFP activation and mCherry repression. As an alternative control selection, one can replace the TF cI5G6G,P with a reporter (e.g., a red fluorescent protein (RFP)) on the PM and then monitor the selection process by infecting TG1 cells and counting the ratio of red to white colonies after each round of selection19.
Materials
REAGENTS
Cloning and plasmid construction
-
Plasmids: M13KO7-ΔPS-ΔgeneIII-ΔgeneVI (Addgene plasmid ID 80840), pLITMUS-rpoN-cIopt-J23106-geneIII (Addgene plasmid ID 80852), pJPC12-ΔPS-PRM-B0034-geneVI (Addgene plasmid ID 80858); optional: pJPC12-ΔPS-mCherry-PR/PRM-GFP (Addgene plasmid ID 80859), pLITMUS-rpoN-cI5G6G,P-J23106-geneIII (Addgene plasmid ID 80861) (Supplementary Fig. 3, Supplementary Table 1). Sequences of all plasmids are listed in Supplementary Data Sets 1–5.
-
Oligonucleotides (Sigma). Primers used for cloning are listed in Supplementary Table 2.
-
KOD Hot Start DNA Polymerase (Merck Millipore, cat. no. 71086). PCR reaction components are listed in the Equipment Setup.
-
Diethyl pyrocarbonate (DEPC)-treated and sterile-filtered water (Sigma, cat. no. 95284)
-
Gibson Assembly Master Mix (New England BioLabs, cat. no. E2611)
-
DpnI endonuclease (New England BioLabs, cat. no. R0176)
-
Super optimal broth with catabolite repression (SOC) medium (Sigma, cat. no. 15544034)
-
DNA gel loading dye, 6× (Thermo Scientific, cat. no. R0611)
-
1-kb Plus DNA Ladder (Thermo Scientific, cat. no. 10787026)
-
SYBR Safe DNA gel stain (Life Technologies, cat. no. S33102)
-
Tris-borate-ethylenediaminetetraacetic acid (TBE) buffer (10×; Sigma, cat. no. T4415)
-
Agarose for gel electrophoresis (Sigma, cat. no. A9539)
-
QIAquick gel extraction kit (Qiagen, cat. no. 28704)
-
QIAquick PCR purification kit (Qiagen, cat. no. 28104)
-
MinElute PCR purification kit (Qiagen, cat. no. 28004)
-
QIAprep Spin miniprep kit (Qiagen, cat. no. 27104)
-
HiSpeed Plasmid Maxi kit (Qiagen, cat. no. 12663)
Strains, buffers and media
-
One Shot chemically competent TOP10 E. coli (Fisher Scientific, cat. no. C404010)
-
Mix & Go competent cells, strain TG1 (Zymo Research, cat. no. T3017)
-
5-Alpha electrocompetent E. coli, optional (New England BioLabs, cat. no. C2989K)
-
Lysogeny broth (LB) with agar (Sigma, cat. no. L2897)
-
Ampicillin (Sigma, cat. no. A0166), chloramphenicol (Sigma, cat. no. C0378), kanamycin (Sigma, cat. no. K4000), and carbenicillin disodium salt (Sigma, cat. no. C1389)
-
2× tryptone yeast extract (2× TY): NaCl (Sigma, cat. no. S9888), yeast extract (Sigma, cat. no. Y1625), tryptone (Sigma, cat. no. T7293)
-
Glycerol (Sigma, cat. no. G5516)
-
Ethanol (≥99.8% vol/vol) for molecular biology (Merck Millipore, cat. no. 1085430250)
-
M9 minimal salts (5×; Sigma, cat. no. M6030)
-
M9 plates: bacteriological agar (Sigma, cat. no. A5306), MgSO4 (Sigma, cat. no. M7506), D-(+)-glucose (Sigma, cat. no. G8270), CaCl2 (Sigma, cat. no. C1016), thiamine-HCl (Sigma, cat. no. T1270)
EQUIPMENT
-
PCR tubes (VWR, cat. no. 732-0545)
-
Microcentrifuge tubes (1.5 ml; Thermo Scientific, cat. no. 05-408-129)
-
Conical centrifuge tube, polypropylene (15 ml; BD Falcon, cat. no. 352097)
-
Conical centrifuge tubes, polypropylene (50 ml; Corning, cat. no. 430829)
-
Schott culture flasks (250 ml; Sigma, cat. no. Z620033)
-
Nunc CryoTubes (Thermo Scientific, cat. no. 366656)
-
Serological pipettes (5, 10 and 25 ml; Fisher Scientific, cat. nos. 13-678-11D, 13-678-11E and 13-678-11)
-
Sterile filters (0.22-μm pore size, Millex-GV, cat. no. SLGV033RS)
-
L-shaped cell spreaders (Fisher Scientific, cat. no. 14-665-231)
-
Cell culture centrifuge Avanti J-26XP (Beckman Coulter, cat. no. 393124)
-
Microcentrifuge (Eppendorf, 5415D)
-
Dri-block heater (Techne, DB100/2)
-
Eppendorf Thermomixer Compact (Sigma, cat. no. T1317)
-
Balance Sartorius Excellence (Sartorius)
-
NanoDrop Lite spectrophotometer (Thermo Scientific)
-
Biophotometer (Eppendorf)
-
Biophotometer cuvettes (Sigma, cat. no. Z605050)
-
Horizontal gel electrophoresis systems (Bio-Rad)
-
Gel documentation system (InGenius 3, Syngene)
-
Gene Pulser Cuvette, 0.1-cm electrode (Bio-Rad, cat. no. 165-2089)
-
Gene Pulser Xcell microbial system (Bio-Rad, cat. no. 1652662)
-
PCR thermocycler (Bio-Rad S1000, cat. no. 1852196)
-
Petri dishes, 57 cm2 (Sigma, cat. no. P7741)
-
Mini incubator (Labnet International, I5110A)
-
Nunc Square BioAssay dishes, 24.1 cm × 24.1 cm (Thermo Scientific, cat. no. 10570502)
-
SI500 shaking incubator (Stuart)
-
Cell culture microplate, 96-well (optional) (Greiner Bio-One, cat. no. 655090)
-
Infinite M200 plate reader (optional) (Tecan)
-
Research pipettes: 10 μl, 100 μl and 1,000 μl (Sigma, cat. no. Z683884)
-
Tips: 10 μl, 200 μl and 1,000 μl (Starlab, cat. nos. S1111-3700-C, S1113-1700-C and S1111-6701-C)
REAGENT SETUP
Antibiotic stocks
-
Prepare 100 mg ml−1 ampicillin in H2O (sterile filtered), 100 mg ml−1 kanamycin in H2O (sterile filtered) and 100 mg ml−1 chloramphenicol in ethanol. Prepare aliquots in sterile 1.5-ml tubes and store them at −20 °C for up to 6 months. The final concentrations, if not stated otherwise, are 100 μg ml−1 ampicillin (1:1,000), 50 μg ml−1 kanamycin (1:2,000) and 25 μg ml−1 chloramphenicol (1:4,000).
Glycerol
-
Prepare a sterile 10% (vol/vol) glycerol solution in H2O for making electrocompetent cells. To obtain glycerol stocks, prepare a sterile 50% (vol/vol) glycerol solution in H2O and add glycerol to the cell culture (final concentration, 20% (vol/vol)) before freezing at −80 °C. Store glycerol stock at 4 °C for up to 3 months.
Tris-borate-ethylenediaminetetraacetic acid electrophoresis buffer
-
Dilute TBE buffer in dH2O to a 1× working solution and store it at room temperature (15–25 °C) for up to 6 months.
Culture medium
-
Autoclave 2× tryptone yeast extract (TY) medium (5 g L−1 NaCl, 10 g L−1 yeast extract, 16 g L−1 tryptone) and add antibiotics where appropriate before use. Store the medium at 4 °C for up to several months.
LB plates
-
Add 35 g of LB powder in 1 L of water and autoclave. Add antibiotics where appropriate, pour into Petri dishes and allow to solidify. Store the plates at 4 °C for up to several weeks. Note that antibiotics degrade over time, which might affect the concentration when stored for prolonged times.
M9 minimal medium plates
-
Autoclave 7 g of bacteriological agar in 500 ml of 1× M9 medium. Add 1 ml of 1 M MgSO4 (autoclaved), 5 ml of 20% (wt/vol) D-(+)-glucose (sterile filtered), 50 μl of 1 M CaCl2 (autoclaved) and 500 μl of 1 M thiamine-HCl (sterile filtered) to M9 agar just before use. Add antibiotics where appropriate. Store the plates at 4 °C for up to several months.
EQUIPMENT SETUP
PCR thermocycler
-
The PCR reaction components are listed below.
Table 5 Critical
For targets >2 kb, final Mg2+ concentrations are adjusted to 2 mM.
The following conditions are used for all PCR reactions:
Table 6 See Supplementary Table 3 for PCR conditions specific for individual reactions.
Infinite M200 plate reader
-
The following setting are used: temperature, 37 °C; duration, 10 h; shaking, 281 r.p.m.; absorbance, 600 ± 9 nm; mCherry fluorescence, excitation 585 ± 9 nm, emission 625 ± 20 nm, gain value 70; GFP fluorescence, excitation 485 ± 9 nm, emission 520 ± 20 nm, gain value 40.
Procedure
Phagemid construction by Gibson assembly
Timing 2 weeks
-
1
Design and order generic forward and reverse primers (e.g., pLITMUS-F and pLITMUS-R; Supplementary Table 2) for the amplification of the PM vector backbone (pLITMUS-rpoN-cIopt-J23106-geneIII) upstream of cIopt and downstream of the terminator BBa_B0015. Note that the terminator BBa_B0015 occurs twice in the parental plasmid. The 'medium strength' rpoN promoter is used to express the evolving gene to achieve a balance between functional expression and any potential metabolic load. The levels of the expressed target gene may need to be adjusted to the function in other cases.
-
2
Design and order user-specific primers (e.g., cI-F and cI-R; Supplementary Table 2) for the GOI plus a terminator of choice (e.g., BBa_B0015) with a 15- to 25-bp overlap with the PM vector backbone.
-
3
Amplify the GOI and vector backbone by PCR (Equipment Setup) and purify the samples using the QIAquick PCR purification kit.
Critical Step
If the PCR reactions contain unwanted by-products, gel extraction should be performed throughout the protocol with the QIAquick gel extraction kit. Use a DNA polymerase with proofreading activity (e.g., KOD DNA polymerase) for all PCR reactions throughout the protocol.
-
4
Remove the parental plasmid by adding 1 μl of DpnI per 50 μl of PCR reaction product and incubating for 1–2 h at 37 °C and 400 r.p.m. (Thermomixer Compact).
-
5
Fuse the two fragments by Gibson assembly38 according to the manufacturer's instructions. Note that Gibson reactions can be downscaled to 5 μl per reaction.
-
6
Dilute the assembled products fourfold with H2O, add 2 μl of the diluted product to 50 μl of chemically competent Top10 cells and transform the cells according to the manufacturer's instructions.
-
7
Incubate the cells for 1 h at 37 °C at 220 r.p.m. (SI500 incubator) and spread them onto LB plates supplemented with 100 μg ml−1 ampicillin.
-
8
Allow the cells to grow overnight at 37 °C.
-
9
The next day, pick single colonies and grow them in 5 ml of 2× TY supplemented with 100 μg ml−1 ampicillin overnight at 37 °C and 220 r.p.m. (SI500 incubator).
-
10
Extract the PM DNA (QIAprep Spin miniprep kit) according to the manufacturer's instructions and confirm the nucleotide sequences by DNA sequencing using the primers pLITMUS-F and pLITMUS-R (Table 1).
Table 1 Oligonucleotides used for sequencing. Pause point
The extracted PM DNA can be stored at –20 °C for several years.
Combinatorial library cloning on phagemids
Timing 2 weeks
-
11
Design and order user-specific forward and reverse primers for the amplification of the PM vector backbone (e.g., pLITMUS-Lib-F and pLITMUS-Lib-R; Supplementary Table 2) and the insertion of the randomized target sequence (e.g., Library 1-F, Library 1-R; Supplementary Table 2). PCR primers for insert amplification require a 15- to 25-bp overlap with each other, as well as a 15- to 25-bp overlap with the amplified vector backbone.
Critical Step
Avoid randomized library positions within the primer overlap regions.
-
12
Amplify the PCR fragments (Equipment Setup) and purify the samples using the QIAquick PCR purification kit or the MinElute PCR purification kit (for samples <100 bp).
Critical Step
The PCR product concentration affects the efficiency of the assembly reaction. Optimized cloning efficiency requires at least 20 ng μl−1 of the PM vector backbone.
-
13
Add 1 μl of DpnI per 50-μl PCR reaction product and incubate for 1–2 h at 37 °C and 400 r.p.m. (Thermomixer Compact).
-
14
Fuse the DNA fragments by Gibson assembly38. Upscale Gibson reactions (e.g., 4 × 20 μl) to increase the total plasmid concentration.
-
15
Pool the Gibson reactions, purify the assembled plasmid using the QIAquick PCR purification kit and elute in 30 μl of H2O.
Critical Step
Note that purification is important to decrease the salt concentration and to decrease the Gibson reaction components, as these are toxic to the cells at high concentrations.
-
16
Measure the plasmid concentration with a spectrophotometer (NanoDrop Lite).
Critical Step
DNA concentrations should be >10 ng μl−1 for high transformation efficiency.
Pause point
The assembled plasmid can be stored at –20 °C for several years.
-
17
Transform 1–2 μl of DNA into 50 μl of electrocompetent cells (DH5-α or TG1) and add 950 μl of SOC medium.
Critical Step
Use electroporation as the method of choice for transformation, as it allows much larger library sizes.
-
18
Incubate for 1 h at 37 °C at 220 r.p.m. (SI500 incubator).
-
19
Plate the transformation reaction on Nunc Square BioAssay dishes (24.1 × 24.1 cm) supplemented with 100 μg ml−1 ampicillin and incubate overnight at 37 °C.
-
20
The next day, harvest the cells with a cell spreader.
Critical Step
Only use plates with more than 105 clones. Estimate the transformation efficiency by plating serial dilutions (10−2 and 10−4 in 2× TY) on additional Petri dishes (57 cm2) supplemented with 100 μg ml−1 ampicillin and counting the colonies the following day. Ideally, to cover the whole library space, at least a threefold excess of colonies relative to the theoretical library size is desired.
-
21
Purify the combinatorial DNA library using the HiSpeed Plasmid Maxi kit and elute in 0.5 ml of TE buffer. Measure the plasmid concentration with a spectrophotometer (NanoDrop Lite). The obtained plasmid concentration should ideally be >50 ng ml−1.
-
22
Pick individual colonies (10–100 clones of a library depending on the library size and quality control desired) from the Petri dishes, which were used to estimate the transformation efficiency (Step 20), and culture each in 5 ml of 2× TY supplemented with 100 μg ml−1 ampicillin overnight at 37 °C at 220 r.p.m. (SI500 incubator). The next day, extract PM DNA (QIAprep Spin miniprep kit) and sequence the GOI using the primers pLITMUS-F and/or pLITMUS-R to confirm library diversity (Table 1).
Pause point
The extracted PM DNA can be stored at –20 °C for several years.
Production of M13 phage from a combinatorial phagemid library
Timing 1 week
-
23
Transform 50 μl of chemically competent Top10 cells with equal mole amounts of HP (M13KO7-ΔPS-ΔgeneIII-ΔgeneVI) and AP (pJPC12-ΔPS-PRM-B0034-geneVI) (10–20 fmol per plasmid; typically 1–2 μl in total). Note that the PRM promoter can be replaced by an alternative promoter (e.g., T7) to obtain higher phage titers in the absence of the activator λ cI.
-
24
Add 250 μl of SOC medium to the samples and incubate for 1 h at 37 °C at 220 r.p.m. (SI500 incubator).
-
25
Spread the cells on LB plates supplemented with 25 μg ml−1 chloramphenicol and 50 μg ml−1 kanamycin. Allow them to grow overnight at 37 °C.
-
26
The next day, pick a single colony and grow it in 2× TY supplemented with 12.5 μg ml−1 chloramphenicol and 25 μg ml−1 kanamycin at 37 °C at 250 r.p.m. (SI500 incubator) until the OD600 reaches 0.4–0.6 (mid-exponential phase) and make cells electrocompetent as described by Gonzales et al.42.
Pause point
Competent cells are stored at –80 °C. Stored electrocompetent cells can be used for the construction of any phage library.
-
27
Transfer 50 μl of the electrocompetent cells to a prechilled 1.5-ml tube on ice and add 1–2 μl of the cloned combinatorial PM library.
-
28
Electroporate cells, and immediately add 950 μl of SOC medium and incubate for 1 h at 37 °C at 220 r.p.m. (SI500 incubator).
-
29
Estimate the actual phage library by colony-counting serial dilutions (10−2 and 10−4 in 2× TY) on LB plates supplemented with ampicillin (Step 20).
Critical Step
Make sure not to lose any library members through low transformation efficiencies.
-
30
Add 3 ml of 2× TY supplemented with 12.5 μg ml−1 chloramphenicol, 25 μg ml−1 kanamycin and 50 μg ml−1 ampicillin to the transformation reaction and grow for 18–20 h at 30 °C at 250 r.p.m. (SI500 incubator). Note that the volume can be adjusted depending on the desired volume of the phage titer.
-
31
The next day, centrifuge the sample for 5 min at 5,000g and 4 °C.
-
32
Filter-sterilize the phage supernatant (0.22-μm pore size).
Pause point
The phage library can be kept at 4 °C for short-term storage (weeks) or at −20 °C for long-term storage (years).
Phage titer analysis
Timing 3 d
-
33
Streak out TG1 cells from a glycerol stock (∼1–5 μl) on an M9 minimal medium plate and incubate overnight at 37 °C. Note that TG1 plates can be used for a maximum of 2 weeks when stored at 4 °C.
Critical Step
Use M9 minimal medium plates to select F-pilus-positive TG1 cells.
-
34
The next day, pick one to four single isolated colonies from the M9 minimal medium plate and inoculate in 10 ml of 2× TY medium in a 50-ml conical centrifuge tube.
-
35
Incubate at 37 °C and 250 r.p.m. (SI500 incubator) until the OD600 reaches 0.4–0.6 (mid-exponential phase). It typically takes 4–6 h for the culture to reach the desired OD600.
Critical Step
Do not let the cells grow into stationary phase, as TG1 cells tend to lose the F episome, and this lowers the overall infection rate.
-
36
In the meantime, prepare serial dilutions (10−2, 10−4, 10−6, 10−8 in 2× TY) of the phage library in sterile 1.5-ml tubes. Phage stocks are diluted before infection to ensure that each cell is infected by only one phage particle (the number of colonies on plates equals the number of phage particles).
-
37
Add 100 μl of the phage dilutions to 900 μl of TG1 cells in a sterile microcentrifuge tube. Mix gently and incubate the samples for 1 h at 37 °C without shaking. Plate 100 μl of cell suspension on prewarmed LB plates supplemented with 100 μg ml−1 ampicillin and incubate overnight at 37 °C.
-
38
The next day, count the number of colonies and calculate the phage titer (equation (1)). Ideally, use the plates containing 20–400 colonies. Note that the 100-fold dilution (Step 37) has to be taken into consideration.
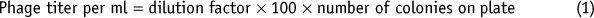
Critical Step
The phage titer should lie between 108 and 1,013 colony-forming units (c.f.u.) per milliliter.
Construction of accessory and reporter plasmids
Timing 2 weeks
-
39
Order user-specific forward and reverse primers to replace the PRM promoter on the AP plasmid (pJPC12-ΔPS-PRM-B0034-geneVI) with a different promoter or inducible input. For vector amplification, use primers B0034-gVI-F and gVI-R, which bind upstream and downstream of PRM (Supplementary Table 2). For insert amplification, make sure to add a 15- to 25-bp overlap for assembly. Optional: clone the same inducible input into pJPC12-ΔPS-mCherry-PR/PRM-GFP to obtain a reporter for the functional characterization of selected proteins. Use primers GFP-F and mCherry-R for vector amplification (Supplementary Table 2).
-
40
Clone the AP and RPs as described in Steps 3–10. Use sequencing primers pJPC12-F and/or pJPC12-R for gene VI constructs and pJPC12-F2 for reporters (Table 1).
Preparation of host cells for directed evolution
Timing 3 d
-
41
Transform 50 μl of competent TG1 cells with equal mole amounts of HP M13KO7-ΔPS-ΔgeneIII-ΔgeneVI and the cloned AP (10–20 fmol per plasmid; typically 1–2 μl in total).
Critical Step
Always use an E. coli strain that contains the F-factor needed for M13 phage infection.
-
42
Add 250 μl of SOC medium to the sample and incubate for 1 h at 37 °C at 220 r.p.m. (SI500 incubator).
-
43
Spread the cells on LB plates supplemented with 25 μg ml−1 chloramphenicol and 50 μg ml−1 kanamycin and allow the cells to grow overnight at 37 °C.
-
44
The next day, pick a single colony and allow the cells to grow in 2 ml of 2× TY supplemented with kanamycin and chloramphenicol for 4–6 h at 37 °C at 250 r.p.m. (SI500 incubator) until the cells reach the late-exponential phase.
-
45
(Optional) Make glycerol stock (Reagent Setup).
Pause point
The glycerol stocks can be stored at −80 °C for up to several years.
-
46
Make serial dilutions of the cell suspension (e.g., 10−6 or 10−8 in M9 medium) and spread the diluted cells on an M9 minimal medium plate supplemented with 25 μg ml−1 chloramphenicol and 50 μg ml−1 kanamycin. These conditions promote phage infectability by maintaining F′ pili.
-
47
Incubate the plates for 30–48 h at 37 °C. Note that bacteria grow much slower on minimal media than on rich media. This plate is used as a source of fresh colonies for selection experiments and can be used for up to 2 weeks when stored at 4 °C.
Phage-assisted batch selection
Timing 2 weeks
-
48
Inoculate 10–20 ml of 2× TY containing 12.5 μg ml−1 chloramphenicol and 12.5 μg ml−1 kanamycin with one to four colonies from the prepared M9 plate (Step 47) in a 50-ml tube (Fig. 6).
Figure 6: Phage-assisted batch selection. A starter culture from TG1 cells containing the modified M13 helper phage (HP; M13KO7-ΔPS-ΔgIII-ΔgVI) and an accessory plasmid (AP) is prepared and cells are grown for 6–8 h at 37 °C until the OD600 reaches 0.4–0.6. Starter cells are infected with the constructed phagemid library to start the first round of selection. Conditional phage production is carried out in a shaking incubator for 16–18 h at 30 °C, and the resulting phage particles are separated from the cells by centrifugation. The obtained phage stock is used to start a new round of selection via infection of a fresh starter culture (round 2). After several rounds of reinfection and selection, a TG1 preculture is infected with the obtained phage stock, and infected TG1 cells are selected on LB plates supplemented with ampicillin. Single colonies are picked, and cells are grown overnight at 37 °C in a shaking incubator. The next day, phagemid DNA is purified and the gene of interest is sequenced.
-
49
Grow the starter culture for 6–8 h at 37 °C and 250 r.p.m. (SI500 incubator) until the OD600 reaches 0.4–0.6.
-
50
Infect 10 ml of the starter culture with the combinatorial phage library at a multiplicity of infection (MOI) of 0.5–5. An excess of cell culture can be chilled on ice and then stored at 4 °C for up to 1 week. This culture may be used for the next rounds of selection. Note that the selection volume can be easily up- or downscaled according to the user's need.
-
51
Incubate the infected cells at 37 °C without stirring for 5 min.
-
52
Incubate the sample for 18–20 h at 30 °C and 250 r.p.m. (SI500 incubator).
-
53
The next day, centrifuge the culture for 5 min at 5,000g and 4 °C, and transfer 1 ml of the supernatant into a sterile microcentrifuge tube. This sample is used to start a new round of selection.
Pause point
Phage supernatants for each round of selection can be stored at 4 °C for short-term storage (several weeks) or at −20 °C for the long term (several years), to continue selection at a later time.
-
54
Infect the starter culture (Step 49) at a ratio of 10−3–10−1 (e.g., 10–1,000 μl of phage supernatant in 10 ml of culture) for the next round of selection.
-
55
Run the selection cycle (Steps 51–54) for several rounds until the target proteins are enriched. This usually takes four to eight rounds depending on the target protein's activity and thus the conditional gene VI expression.
Critical Step
The phage titer should ideally stay between 106 and 1012 c.f.u. ml−1 after each round of the selection (Step 56). Very high infection rates (MOI >10) lead to multiple infections and thus propagation of nonfunctional library members ('cheaters'), whereas very low rates (MOI <0.1) decrease the performance of the system.
-
56
(Optional) During the selection process, monitor the phage titer for each round by phage titer analysis (Steps 33–38).
-
57
(Optional) Monitor the selection process by infecting reporter cells (TG1 with a suitable RP; e.g., pJPC12-ΔPS-mCherry-P/PM,5G6G-GFP) with the obtained phage titer for each round (analogous to Steps 33–38). Streak out infected cells on LB plates supplemented with 25 μg ml−1 chloramphenicol and 100 μg ml−1 ampicillin and allow them to grow overnight at 37 °C. The next day, analyze the plates under the UV light of a gel documentation system. Nonactive library members result in red colonies, whereas active library members lead to green colonies because of GFP activation and mCherry repression. Store the plates at 4 °C overnight for improved mCherry signals.
-
58
After selection, filter-sterilize the phage supernatant (0.22-μm pore size) and serially dilute the sample with 2× TY medium before infecting TG1 cells with an OD600 of 0.4–0.6. Incubate the infected cells for 1 h at 37 °C before plating (Step 37).
-
59
Select infected cells on 100 μg ml−1 ampicillin plates overnight at 37 °C.
-
60
The next day, pick at least three colonies per selection and grow each colony in 5 ml of 2× TY supplemented with ampicillin overnight at 37 °C and 250 r.p.m. (SI500 incubator).
-
61
The next day, extract PM DNA (QIAprep Spin miniprep kit) and sequence the GOI using pLITMUS-F and/or pLITMUS-R primers (Table 1).
Pause point
The extracted PM DNA can be stored at –20 °C for several years.
Characterization of evolved proteins (optional)
Timing 3 d
-
62
Transform 50 μl of competent TG1 cells with equal moles of a selected PM and a suitable RP (e.g., pJPC12-ΔPS-mCherry-P/PM,5G6G-GFP) (10–20 fmol per plasmid, typically 1–2 μl in total). Transform the RP into TG1 cells and use as a control. (Optional) Delete the expression cassette rpoN-cIopt-B0015 from the PM (e.g., pLITMUS-ΔcIopt-F, pLITMUS-ΔcIopt-R) and transform the obtained plasmid (pLITMUS-J23106-geneIII) together with the reporter to compensate for growth effects between control and selected PMs.
-
63
Spread the cells on LB plates supplemented with 25 μg ml−1 chloramphenicol and 100 μg ml−1 ampicillin and allow the cells to grow overnight at 37 °C.
-
64
The next day, pick single colonies and allow them to grow in 1 ml of 2× TY supplemented with 5 μg ml−1 chloramphenicol and 5 μg ml−1 carbenicillin for 4–6 h at 37 °C at 250 r.p.m. (SI500 incubator). Analyze at least three replicates per transformation.
-
65
Measure the OD600 of each replicate (150 μl) with the Tecan Infinite M200 plate reader.
-
66
Dilute cultures in 2× TY supplemented with 5 μg ml−1 chloramphenicol and 5 μg ml−1 carbenicillin to a final OD600 of 0.01 (150 μl) in a 96-well microplate.
-
67
Measure the absorbance at 600 nm (green fluorescence, excitation 485 nm, emission 520 nm; red fluorescence, excitation 585 nm, emission 625 nm) every 10 min with the Infinite M200 plate reader (at 37 °C, shaking between readings) until the cells reach stationary phase.
-
68
For data analysis, use fluorescence readings in the mid-exponential phase (OD600 of 0.2) and correct absorbance and fluorescence against readings of a TG1 culture. Normalize the fluorescence for the number of cells by dividing by the absorbance.
Troubleshooting
Troubleshooting advice can be found in Table 2.
Timing
Phagemid construction by Gibson assembly
Steps 1 and 2, design of primers and oligo synthesis by supplier: 1 week
Steps 3–10, cloning of phagemid: 1 week
Combinatorial library cloning on phagemids
Step 11, design of primers and oligo synthesis by supplier: 1 week
Steps 12–22, cloning of combinatorial library: 1 week
Production of M13 phage from a combinatorial phagemid library
Steps 23–32, transfer from plasmid library to phage library: 1 week
Phage titer analysis
Steps 33–38, analysis of phage concentration: 3 d
Construction of accessory and reporter plasmids (can be done in parallel with phage library cloning)
Step 39, design of primers and oligo synthesis by supplier: 1 week
Step 40, cloning of AP and reporter plasmid: 1 week
Preparation of host cells for directed evolution (can be done in parallel after successful AP cloning)
Steps 41–47, transformation and plating of cells: 3 d
Phage-assisted batch selection
Steps 48–57, batch selections: 1 week
Steps 58–61, extraction and sequencing of selected genes: 1 week
Characterization of evolved proteins (optional)
Steps 62–68, functional characterization by reporter assay: 3 d
Anticipated results
The first section of this protocol describes the construction of combinatorial libraries used for subsequent directed evolution experiments. As examples, we describe the construction of two cIopt libraries, each of which contains five randomized positions: library 1 (45S, 46G, 47V, 48G, 55N) and library 2 (45S, 46G, 48G, 49A, 55N). Quality-control sequencing of 10–100 clones of a library may be performed to confirm diversity, depending on the library size and quality control desired. For example, ten individual clones of a constructed library should ideally result in ten different variants (Table 3).
The second section illustrates the directed evolution of proteins based on conditional M13 phage propagation. Libraries 1 and 2 are selected against engineered promoters for six to eight rounds, leading to enrichment of TFs with binding activations against their novel promoters (Table 4). We frequently obtain amino acid substitutions that occur spontaneously at certain positions not covered by the combinatorial space of the library. These mutations can originate either from mutations during library cloning or from the spontaneous error rate of M13 phage replication, which is ∼0.0046 mutations per genome per replication43. Such mutations can provide function19 and contribute to directed evolution.
The last section of the protocol describes the characterization of selected TFs. The reporter assay is designed in such a way that TF binding to the bidirectional promoter results in GFP activation and mCherry repression. For baseline comparison, GFP and mCherry expression are measured for each promoter in the absence of a TF. The evolved TF variants enable simultaneous activation and repression against their engineered bidirectional promoters. For the selected cI variant (cI5G6G,P) against the bidirectional promoter P/PM,5G6G, GFP production is upregulated tenfold, and 94% of mCherry is repressed (Fig. 7a,b). The evolved cI variant (cI5T6T,P) against the bidirectional promoter P/PM,5T6T results in a ninefold activation and 98% mCherry repression (Fig. 7c,d). This protocol further shows a method to analyze cross-reactivities for DNA-binding proteins. WT cI and cIopt activate GFP sixfold and ninefold and simultaneously repress 90% and 82% of mCherry production on the WT PR/PRM promoter, whereas this effect is not observed for any of the engineered promoter variants (Fig. 7e,f). The selected TFs also do not cross-react with each other, thus ensuring orthogonality.
The activity of the selected TFs needs to be verified by a reporter assay. (a,b) Basal promoter strength of the bidirectional promoter P/PM,5G6G and its dual activation and repression by the selected TF cI5G6G,P, as determined on the basis of GFP (a) or mCherry (b) fluorescence. (c,d) Basal promoter strength of the bidirectional promoter P/PM,5T6T and its dual activation and repression by the selected TF cI5T6T,P, as determined on the basis of GFP (c) or mCherry (d) fluorescence. (e,f) Basal promoter strength of PR/PRM and its activation/repression by WT λ cI and cIopt, as determined on the basis of GFP (e) or mCherry (f) fluorescence. Cross-reactivity of TF variants was ruled out by reporter analysis. Basal mCherry expression varies between promoters because of base-pair substitutions next to the −35 and −10 regions, and y-axes are adjusted accordingly. In all plots, GFP and mCherry expression was normalized to OD600. Four biological replicates were measured for each sample; data points represent individual replicates. Data are shown as mean and s.d. a.u., arbitrary units.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
References
Packer, M.S. & Liu, D.R. Methods for the directed evolution of proteins. Nat. Rev. Genet. 16, 379–394 (2015).
Jäckel, C., Kast, P. & Hilvert, D. Protein design by directed evolution. Annu. Rev. Biophys. 37, 153–173 (2008).
Cobb, R.E., Sun, N. & Zhao, H. Directed evolution as a powerful synthetic biology tool. Methods 60, 81–90 (2013).
Lutz, S. Beyond directed evolution—semi-rational protein engineering and design. Curr. Opin. Biotechnol. 21, 734–743 (2010).
Smith, G.P. Filamentous fusion phage: novel expression vectors that display cloned antigens on the virion surface. Science 228, 1315 (1985).
Lee, C.M.Y., Iorno, N., Sierro, F. & Christ, D. Selection of human antibody fragments by phage display. Nat. Protoc. 2, 3001–3008 (2007).
McCafferty, J., Griffiths, A.D., Winter, G. & Chiswell, D.J. Phage antibodies: filamentous phage displaying antibody variable domains. Nature 348, 552–554 (1990).
Hoogenboom, H.R. in Antibody Phage Display: Methods and Protocols (eds. O'Brien, P.M. & Aitken, R.) 1–37 (Humana Press, 2002).
Isalan, M., Klug, A. & Choo, Y. A rapid, generally applicable method to engineer zinc fingers illustrated by targeting the HIV-1 promoter. Nat. Biotechnol. 19, 656–660 (2001).
Hubbard, B.P. et al. Continuous directed evolution of DNA-binding proteins to improve TALEN specificity. Nat. Methods 12, 939–942 (2015).
Fernandez-Gacio, A., Uguen, M. & Fastrez, J. Phage display as a tool for the directed evolution of enzymes. Trends Biotechnol. 21, 408–414 (2003).
Demartis, S. et al. A strategy for the isolation of catalytic activities from repertoires of enzymes displayed on phage1. J. Mol. Biol. 286, 617–633 (1999).
Badran, A.H. & Liu, D.R. In vivo continuous directed evolution. Curr. Opin. Chem. Biol. 24, 1–10 (2015).
Hasty, J., Dolnik, M., Rottschäfer, V. & Collins, J.J. Synthetic gene network for entraining and amplifying cellular oscillations. Phys. Rev. Lett. 88, 148101 (2002).
Hasty, J., Isaacs, F., Dolnik, M., McMillen, D. & Collins, J. Designer gene networks: towards fundamental cellular control. Chaos 11, 207–220 (2001).
Guet, C.C., Elowitz, M.B., Hsing, W. & Leibler, S. Combinatorial synthesis of genetic networks. Science 296, 1466–1470 (2002).
Stanton, B.C. et al. Genomic mining of prokaryotic repressors for orthogonal logic gates. Nat. Chem. Biol. 10, 99–105 (2014).
Rhodius, V.A. et al. Design of orthogonal genetic switches based on a crosstalk map of σs, anti-σs, and promoters. Mol. Syst. Biol. 9, 702–702 (2013).
Brödel, A.K., Jaramillo, A. & Isalan, M. Engineering orthogonal dual transcription factors for multi-input synthetic promoters. Nat. Commun. 7, 13858 (2016).
Boeke, J., Model, P. & Zinder, N. Effects of bacteriophage f1 gene III protein on the host cell membrane. Mol. Gen. Genet. 186, 185–192 (1982).
Rakonjac, J. & Model, P. Roles of pIII in filamentous phage assembly. J. Mol. Biol. 282, 25–41 (1998).
Esvelt, K.M., Carlson, J.C. & Liu, D.R. A system for the continuous directed evolution of biomolecules. Nature 472, 499–503 (2011).
Dickinson, B.C., Packer, M.S., Badran, A.H. & Liu, D.R. A system for the continuous directed evolution of proteases rapidly reveals drug-resistance mutations. Nat. Commun. 5, 5352 (2014).
Badran, A.H. et al. Continuous evolution of Bacillus thuringiensis toxins overcomes insect resistance. Nature 533, 58–63 (2016).
Galvão, T.C. & de Lorenzo, V. Transcriptional regulators à la carte: engineering new effector specificities in bacterial regulatory proteins. Curr. Opin. Biotechnol. 17, 34–42 (2006).
Gutierrez, J.M.P., Hinkley, T., Taylor, J.W., Yanev, K. & Cronin, L. Evolution of oil droplets in a chemorobotic platform. Nat. Commun. 5, 5571 (2014).
Fujii, S. et al. Liposome display for in vitro selection and evolution of membrane proteins. Nat. Protoc. 9, 1578–1591 (2014).
Wilson, D.S., Keefe, A.D. & Szostak, J.W. The use of mRNA display to select high-affinity protein-binding peptides. Proc. Natl. Acad. Sci. USA 98, 3750–3755 (2001).
Seelig, B. mRNA display for the selection and evolution of enzymes from in vitro–translated protein libraries. Nat. Protoc. 6, 540–552 (2011).
Wang, H.H. et al. Programming cells by multiplex genome engineering and accelerated evolution. Nature 460, 894–898 (2009).
Kleinstiver, B.P. et al. Engineered CRISPR-Cas9 nucleases with altered PAM specificities. Nature 523, 481–485 (2015).
Digianantonio, K.M. & Hecht, M.H. A protein constructed de novo enables cell growth by altering gene regulation. Proc. Natl. Acad. Sci. USA 113, 2400–2405 (2016).
Fisher, M.A., McKinley, K.L., Bradley, L.H., Viola, S.R. & Hecht, M.H. De novo designed proteins from a library of artificial sequences function in Escherichia coli and enable cell growth. PLoS One 6, e15364 (2011).
Kim, Y.-S., Jung, H.-C. & Pan, J.-G. Bacterial cell surface display of an enzyme library for selective screening of improved cellulase variants. Appl. Environ. Microbiol. 66, 788–793 (2000).
Badran, A.H. & Liu, D.R. Development of potent in vivo mutagenesis plasmids with broad mutational spectra. Nat. Commun. 6, 8425 (2015).
Isalan, M. Construction of semi-randomized gene libraries with weighted oligonucleotide synthesis and PCR. Nat. Protoc. 1, 468–475 (2006).
de Marco, A. Strategies for successful recombinant expression of disulfide bond-dependent proteins in Escherichia coli. Microb. Cell Fact. 8, 26 (2009).
Gibson, D.G. et al. Enzymatic assembly of DNA molecules up to several hundred kilobases. Nat. Methods 6, 343–345 (2009).
Bushman, F.D., Shang, C. & Ptashne, M. A single glutamic acid residue plays a key role in the transcriptional activation function of lambda repressor. Cell 58, 1163–1171 (1989).
Weiss, G.A., Watanabe, C.K., Zhong, A., Goddard, A. & Sidhu, S.S. Rapid mapping of protein functional epitopes by combinatorial alanine scanning. Proc. Natl. Acad. Sci. USA 97, 8950–8954 (2000).
Hochschild, A. & Lewis, M. The bacteriophage λ CI protein finds an asymmetric solution. Curr. Opin. Struct. Biol. 19, 79–86 (2009).
Gonzales, M.F., Brooks, T., Pukatzki, S.U. & Provenzano, D. Rapid protocol for preparation of electrocompetent Escherichia coli and Vibrio cholerae. J. Vis. Exp. 80, 50684 (2013).
Drake, J.W., Charlesworth, B., Charlesworth, D. & Crow, J.F. Rates of spontaneous mutation. Genetics 148, 1667–1686 (1998).
Stano, N.M. & Patel, S.S. T7 lysozyme represses T7 RNA polymerase transcription by destabilizing the open complex during initiation. J. Biol. Chem. 279, 16136–16143 (2004).
Albright, R.A. & Matthews, B.W. How Cro and λ-repressor distinguish between operators: the structural basis underlying a genetic switch. Proc. Natl. Acad. Sci. USA 95, 3431–3436 (1998).
Beamer, L.J. & Pabo, C.O. Refined 1.8 Å crystal structure of the λ repressor-operator complex. J. Mol. Biol. 227, 177–196 (1992).
Stayrook, S., Jaru-Ampornpan, P., Ni, J., Hochschild, A. & Lewis, M. Crystal structure of the λ repressor and a model for pairwise cooperative operator binding. Nature 452, 1022–1025 (2008).
Acknowledgements
This research was supported by European Commission grant FP7-ICT-2013-10 (No. 610730, EVOPROG). A.J. was funded by FP7-KBBE (Nos. 613745, PROMYS), H2020 Marie Sklodowska-Curie (No. 642738, MetaRNA) and EPSRC-BBSRC (No. BB/M017982/1, WISB center). M.I. is funded by New Investigator award no. WT102944 from the Wellcome Trust UK. The authors thank T. Bartels and M. Storch for their critical reading of the manuscript.
Author information
Authors and Affiliations
Contributions
A.K.B., A.J. and M.I. developed the protocol. A.K.B. performed the experiments. A.K.B. and M.I. wrote the manuscript. M.I. and A.J. supervised the project and contributed reagents, materials and analysis tools.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Sequences of engineered promoters used for phage-assisted selections.
The natural operator O3 of PR/PRM is replaced by the natural operator OR1 from cI434 in pJPC12-ΔPS-PRM-B0034-geneVI (Addgene plasmid ID: 80858) in order to bypass autorepression at high cI concentration (mismatches are shown in red). Synthetic promoters are derived from the consensus sequence CS (TATCACCGCCGGTGATA) of bacteriophage λ operators. Mutated base pairs in O1 and O2 of synthetic promoters are bold and the engineered promoters are named after the position of the base substitution in the consensus half-site. WT cI binding to O3 was restored for counterselections by inserting the CS at O3. For selections, the promoter variants are used as unidirectional promoters with M13 gene VI downstream PM variants on the accessory plasmid. Operators are highlighted as follows: O1 blue, O2 green, O3 grey.
Supplementary Figure 2 Sequences of synthetic promoters used for functional characterization of selected TF variants.
The natural operator O3 of PR/PRM is replaced by an obliterated O3 site in order to bypass autorepression at high cI concentration. Synthetic promoters are derived from the consensus sequence CS (TATCACCGCCGGTGATA) of bacteriophage λ operators. Mutated base pairs in O1 and O2 of synthetic promoters are bold and the engineered promoters are named after the position of the base substitution in the consensus half-site. For functional characterization of selected TFs, the bidirectional promoter variants are used upstream of GFP and mCherry on the reporter plasmid. Operators are highlighted as follows: O1 blue, O2 green, O3 grey.
Supplementary Figure 3 Plasmid maps needed to set up the directed evolution system.
(a) The modified helper phage HP (M13KO7-ΔPS-ΔgeneIII-ΔgeneVI) contains all phage genes required for phage replication except the genes III and VI. The weak packaging signal (PS) is removed to bypass helper phage propagation. (b) The phagemid pLITMUS-rpoN-cIopt-J23106-gIII provides the gene of interest cIopt, the M13 packaging signal (PS) as well as constitutively expressed gene III. (c) The accessory plasmid pJPC12-ΔPS-PRM-gVI contains a conditional gene VI expression circuit, induced by an active library member on the phagemid. (d) The reporter plasmid pJPC12-ΔPS-mCherry-PR/PRM-GFP is used to characterize the selected library members. For additional information on the pJPC12 vector see reference. DNA sequences are shown in Supplementary Data 1–5.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–3, Supplementary Tables 1–3 and Supplementary Dataset 1–5. (PDF 1974 kb)
Rights and permissions
About this article
Cite this article
Brödel, A., Jaramillo, A. & Isalan, M. Intracellular directed evolution of proteins from combinatorial libraries based on conditional phage replication. Nat Protoc 12, 1830–1843 (2017). https://doi.org/10.1038/nprot.2017.084
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2017.084
This article is cited by
-
A systematic approach to inserting split inteins for Boolean logic gate engineering and basal activity reduction
Nature Communications (2021)
-
Phage-assisted continuous and non-continuous evolution
Nature Protocols (2020)
-
Trp-ing upon new repressors
Nature Chemical Biology (2018)
-
Carbohydrate active enzyme domains from extreme thermophiles: components of a modular toolbox for lignocellulose degradation
Extremophiles (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.