Abstract
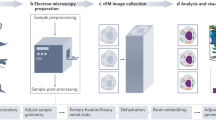
Our groups have recently developed related approaches for sample preparation for super-resolution imaging within endogenous cellular environments using correlative light and electron microscopy (CLEM). Four distinct techniques for preparing and acquiring super-resolution CLEM data sets for aldehyde-fixed specimens are provided, including Tokuyasu cryosectioning, whole-cell mount, cell unroofing and platinum replication, and resin embedding and sectioning. The choice of the best protocol for a given application depends on a number of criteria that are discussed in detail. Tokuyasu cryosectioning is relatively rapid but is limited to small, delicate specimens. Whole-cell mount has the simplest sample preparation but is restricted to surface structures. Cell unroofing and platinum replication creates high-contrast, 3D images of the cytoplasmic surface of the plasma membrane but is more challenging than whole-cell mount. Resin embedding permits serial sectioning of large samples but is limited to osmium-resistant probes, and is technically difficult. Expected results from these protocols include super-resolution localization (∼10–50 nm) of fluorescent targets within the context of electron microscopy ultrastructure, which can help address cell biological questions. These protocols can be completed in 2–7 d, are compatible with a number of super-resolution imaging protocols, and are broadly applicable across biology.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Shimomura, O., Johnson, F.H. & Saiga, Y. Extraction, purification and properties of aequorin, a bioluminescent protein from the luminous hydromedusan, Aequorea. J. Cell. Physiol. 59, 223–239 (1962).
Tsien, R.Y. The green fluorescent protein. Annu. Rev. Biochem. 67, 509–544 (1998).
Shaner, N.C., Patterson, G.H. & Davidson, M.W. Advances in fluorescent protein technology. J. Cell Sci. 120, 4247–4260 (2007).
Lippincott-Schwartz, J. & Manley, S. Putting super-resolution fluorescence microscopy to work. Nat. Methods 6, 21–23 (2009).
Betzig, E. et al. Imaging intracellular fluorescent proteins at nanometer resolution. Science 313, 1642–1645 (2006).
Hess, S.T., Girirajan, T.P.K. & Mason, M.D. Ultra-high resolution imaging by fluorescence photoactivation localization microscopy. Biophys. J. 91, 4258–4272 (2006).
Rust, M.J., Bates, M. & Zhuang, X. Sub-diffraction-limit imaging by stochastic optical reconstruction microscopy (STORM). Nat. Methods 3, 793–795 (2006).
Westphal, V. & Hell, S.W. Nanoscale resolution in the focal plane of an optical microscope. Phys. Rev. Lett. 94, 143903 (2005).
Gustafsson, M.G. Surpassing the lateral resolution limit by a factor of two using structured illumination microscopy. J. Microsc. 198, 82–87 (2000).
Li, D. et al. Extended-resolution structured illumination imaging of endocytic and cytoskeletal dynamics. Science 349, aab3500 (2015).
Heilemann, M., van de Linde, S., Mukherjee, A., Mukherjee, A. & Sauer, M. Super-resolution imaging with small organic fluorophores. Angew. Chem. Int. Ed. Engl. 48, 6903–6908 (2009).
Kim, D. et al. Correlative stochastic optical reconstruction microscopy and electron microscopy. PLoS One 10, e0124581–20 (2015).
Watanabe, S. et al. Protein localization in electron micrographs using fluorescence nanoscopy. Nat. Methods 8, 80–84 (2011).
Nanguneri, S., Flottmann, B., Horstmann, H., Heilemann, M. & Kuner, T. Three-dimensional, tomographic super-resolution fluorescence imaging of serially sectioned thick samples. PLoS One 7, e38098 (2012).
Suleiman, H. et al. Nanoscale protein architecture of the kidney glomerular basement membrane. eLife 2, e01149 (2013).
Perkovic, M. et al. Correlative light- and electron microscopy with chemical tags. J. Struct. Biol. 186, 205–213 (2014).
Löschberger, A., Franke, C., Krohne, G., van de Linde, S. & Sauer, M. Correlative super-resolution fluorescence and electron microscopy of the nuclear pore complex with molecular resolution. J. Cell Sci. 127, 4351–4355 (2014).
Kopek, B.G., Shtengel, G., Xu, C.S., Clayton, D.A. & Hess, H.F. Correlative 3D superresolution fluorescence and electron microscopy reveal the relationship of mitochondrial nucleoids to membranes. Proc. Natl. Acad. Sci. USA 109, 6136–6141 (2012).
Kopek, B.G., Shtengel, G., Grimm, J.B., Clayton, D.A. & Hess, H.F. Correlative photoactivated localization and scanning electron microscopy. PLoS One 8, e77209 (2013).
Paez-Segala, M.G. et al. Fixation-resistant photoactivatable fluorescent proteins for CLEM. Nat. Methods 12, 215–218 (2015).
Van Engelenburg, S.B. et al. Distribution of ESCRT machinery at HIV assembly sites reveals virus scaffolding of ESCRT subunits. Science 343, 653–656 (2014).
Sochacki, K.A., Shtengel, G., Van Engelenburg, S.B., Hess, H.F. & Taraska, J.W. Correlative super-resolution fluorescence and metal-replica transmission electron microscopy. Nat. Methods 11, 305–308 (2014).
Gould, T.J., Verkhusha, V.V. & Hess, S.T. Imaging biological structures with fluorescence photoactivation localization microscopy. Nat. Protoc. 4, 291–308 (2009).
Shtengel, G. et al. Imaging cellular ultrastructure by PALM, iPALM, and correlative iPALM-EM. Methods Cell Biol. 123, 273–294 (2014).
van de Linde, S. et al. Direct stochastic optical reconstruction microscopy with standard fluorescent probes. Nat. Protoc. 6, 991–1009 (2011).
Koga, D., Kusumi, S., Shodo, R., Dan, Y. & Ushiki, T. High-resolution imaging by scanning electron microscopy of semithin sections in correlation with light microscopy. Microscopy 64, 387–394 (2015).
Collman, F. et al. Mapping synapses by conjugate light-electron array tomography. J. Neurosci. 35, 5792–5807 (2015).
Micheva, K.D. & Smith, S.J. Array tomography: a new tool for imaging the molecular architecture and ultrastructure of neural circuits. Neuron 55, 25–36 (2007).
Egner, A. et al. Fluorescence nanoscopy in whole cells by asynchronous localization of photoswitching emitters. Biophys. J. 93, 3285–3290 (2007).
Dertinger, T., Colyer, R., Iyer, G., Weiss, S. & Enderlein, J. Fast, background-free, 3D super-resolution optical fluctuation imaging (SOFI). Proc. Natl. Acad. Sci. USA 106, 22287–22292 (2009).
Rego, E.H. et al. Nonlinear structured-illumination microscopy with a photoswitchable protein reveals cellular structures at 50-nm resolution. Proc. Natl. Acad. Sci. USA 109, E135–E143 (2012).
Bornfleth, H., Satzler, K., Eils, R. & Cremer, C. High-precision distance measurements and volume-conserving segmentation of objects near and below the resolution limit in three dimensional confocal fluorescence microscopy. J. Microsc. 189, 118–136 (1998).
Van Oijen, A.M., Köhler, J., Schmidt, J., Müller, M. & Brakenhoff, G.J. 3-Dimensional super-resolution by spectrally selective imaging. Chem. Phys. Lett. 292, 183–187 (1998).
Lemmer, P. et al. SPDM: light microscopy with single-molecule resolution at the nanoscale. Appl. Phys. B 93, 1–12 (2008).
Hofmann, M., Eggeling, C., Jakobs, S. & Hell, S.W. Breaking the diffraction barrier in fluorescence microscopy at low light intensities by using reversibly photoswitchable proteins. Proc. Natl. Acad. Sci. USA 102, 17565–17569 (2005).
Fölling, J. et al. Fluorescence nanoscopy by ground-state depletion and single-molecule return. Nat. Methods 5, 943–945 (2008).
Egner, A., Jakobs, S. & Hell, S.W. Fast 100-nm resolution three-dimensional microscope reveals structural plasticity of mitochondria in live yeast. Proc. Natl. Acad. Sci. USA 99, 3370–3375 (2002).
Egner, A., Verrier, S., Goroshkov, A., Söling, H.-D. & Hell, S.W. 4Pi-microscopy of the Golgi apparatus in live mammalian cells. J. Struct. Biol. 147, 70–76 (2004).
Schrader, M., Bahlmann, K., Giese, G. & Hell, S.W. 4Pi-confocal imaging in fixed biological specimens. Biophys. J. 75, 1659–1668 (1998).
Perinetti, G. et al. Correlation of 4Pi and electron microscopy to study transport through single golgi stacks in living cells with super resolution. Traffic 10, 379–391 (2009).
Schübbe, S., Cavelius, C., Schumann, C., Koch, M. & Kraegeloh, A. STED microscopy to monitor agglomeration of silica particles inside A549 cells. Adv. Eng. Mater. 12, 417–422 (2010).
Chang, Y.-W. et al. Correlated cryogenic photoactivated localization microscopy and cryo-electron tomography. Nat. Methods 11, 737–739 (2014).
Liu, B. et al. Three-dimensional super-resolution protein localization correlated with vitrified cellular context. Sci. Rep. 5, 13017 (2015).
Kaufmann, R. et al. Super-resolution microscopy using standard fluorescent proteins in intact cells under cryo-conditions. Nano Lett. 14, 4171–4175 (2014).
Johnson, E. et al. Correlative in-resin super-resolution and electron microscopy using standard fluorescent proteins. Sci. Rep. 5, 9583 (2015).
Ding, B., Turgeon, R. & Parthasarathy, M.V. Substructure of freeze-substituted plasmodesmata. Protoplasma 169, 28–41 (1992).
Los, G.V. et al. HaloTag: a novel protein labeling technology for cell imaging and protein analysis. ACS Chem. Biol. 3, 373–382 (2008).
Keppler, A. et al. A general method for the covalent labeling of fusion proteins with small molecules in vivo. Nat. Biotechnol. 21, 86–89 (2003).
Sharonov, A. & Hochstrasser, R.M. Wide-field subdiffraction imaging by accumulated binding of diffusing probes. Proc. Natl. Acad. Sci. USA 103, 18911–18916 (2006).
Endesfelder, U. Advances in correlative single-molecule localization microscopy and electron microscopy. NanoBioImaging 1, 1–9 (2015).
de Boer, P., Hoogenboom, J.P. & Giepmans, B.N.G. Correlated light and electron microscopy: ultrastructure lights up!. Nat. Methods 12, 503–513 (2015).
Li, J., Wang, Y., Chiu, S.-L. & Cline, H.T. Membrane targeted horseradish peroxidase as a marker for correlative fluorescence and electron microscopy studies. Front. Neural Circuits 4, 6 (2010).
Connolly, C.N., Futter, C.E., Gibson, A., Hopkins, C.R. & Cutler, D.F. Transport into and out of the Golgi complex studied by transfecting cells with cDNAs encoding horseradish peroxidase. J. Cell Biol. 127, 641–652 (1994).
Lam, S.S. et al. Directed evolution of APEX2 for electron microscopy and proximity labeling. Nat. Methods 12, 51–54 (2014).
Martell, J.D. et al. Engineered ascorbate peroxidase as a genetically encoded reporter for electron microscopy. Nat. Biotechnol. 30, 1143–1148 (2012).
Shu, X. et al. A genetically encoded tag for correlated light and electron microscopy of intact cells, tissues, and organisms. PLoS Biol. 9, e1001041 (2011).
Viswanathan, S. et al. High-performance probes for light and electron microscopy. Nat. Methods 12, 568–576 (2015).
Mercogliano, C.P. & DeRosier, D.J. Concatenated metallothionein as a clonable gold label for electron microscopy. J. Struct. Biol. 160, 70–82 (2007).
Wang, Q., Mercogliano, C.P. & Löwe, J. A ferritin-based label for cellular electron cryotomography. Structure 19, 147–154 (2011).
Collins, J.S. & Goldsmith, T.H. Spectral properties of fluorescence induced by glutaraldehyde fixation. J. Histochem. Cytochem. 29, 411–414 (1981).
Clancy, B. & Cauller, L.J. Reduction of background autofluorescence in brain sections following immersion in sodium borohydride. J. Neurosci. Methods 83, 97–102 (1998).
McKinney, S.A., Murphy, C.S., Hazelwood, K.L., Davidson, M.W. & Looger, L.L. A bright and photostable photoconvertible fluorescent protein. Nat. Methods 6, 131–133 (2009).
Chudakov, D.M., Lukyanov, S. & Lukyanov, K.A. Tracking intracellular protein movements using photoswitchable fluorescent proteins PS-CFP2 and Dendra2. Nat. Protoc. 2, 2024–2032 (2007).
Zhang, M. et al. Rational design of true monomeric and bright photoactivatable fluorescent proteins. Nat. Methods 9, 727–729 (2012).
Wang, S., Moffitt, J.R., Dempsey, G.T., Xie, X.S. & Zhuang, X. Characterization and development of photoactivatable fluorescent proteins for single-molecule-based superresolution imaging. Proc. Natl. Acad. Sci. USA 111, 8452–8457 (2014).
Wysocki, L.M. et al. Facile and general synthesis of photoactivatable xanthene dyes. Angew. Chem. Int. Ed. Engl. 50, 11206–11209 (2011).
Slot, J.W., Slot, J.W., Geuze, H.J. & Geuze, H.J. Cryosectioning and immunolabeling. Nat. Protoc. 2, 2480–2491 (2007).
Webster, P. & Webster, A. in Methods in Molecular Biology 369, 257–289 (Humana Press, 2007).
Shtengel, G. et al. Interferometric fluorescent super-resolution microscopy resolves 3D cellular ultrastructure. Proc. Natl. Acad. Sci. USA 106, 3125–3130 (2009).
Knott, G., Marchman, H., Wall, D. & Lich, B. Serial section scanning electron microscopy of adult brain tissue using focused ion beam milling. J. Neurosci. 28, 2959–2964 (2008).
Tokuyasu, K.T. Use of poly(vinylpyrrolidone) and poly(vinyl alcohol) for cryoultramicrotomy. Histochem. J. 21, 163–171 (1989).
Larson, D.R., Johnson, M.C., Webb, W.W. & Vogt, V.M. Visualization of retrovirus budding with correlated light and electron microscopy. Proc. Natl. Acad. Sci. USA 102, 15453–15458 (2005).
Ryan, U.S. & Hart, M.A. Electron microscopy of endothelial cells in culture: II. Scanning electron microscopy and OTOTO impregnation method. J. Tissue Cult. Methods 10, 35–36 (1986).
Svitkina, T. Methods in Molecular Biology 586, 187–206 (Humana Press, 2009).
Collins, A., Warrington, A., Taylor, K.A. & Svitkina, T. Structural organization of the actin cytoskeleton at sites of clathrin-mediated endocytosis. Curr. Biol. 21, 1167–1175 (2011).
Heuser, J. Quick-freeze, deep-etch preparation of samples for 3-D electron microscopy. Trends Biochem. Sci. 6, 64–68 (1980).
Heuser, J. The production of 'cell cortices' for light and electron microscopy. Traffic 1, 545–552 (2000).
Caplan, J. et al. Correlative Protein Localization in Yeast Carl Zeiss Microscopy (2013).
Kobs, S.F. & Behrman, E.J. Reactions of osmium tetroxide with imidazoles. Inorg. Chim. Acta 138, 113–120 (1987).
Nielson, A.J. & Griffith, W.P. Tissue fixation by osmium tetroxide. A possible role for proteins. J. Histochem. Cytochem. 27, 997–999 (1979).
Pertsinidis, A., Zhang, Y. & Chu, S. Subnanometre single-molecule localization, registration and distance measurements. Nature 466, 647–651 (2010).
Nieuwenhuizen, R.P.J. et al. Measuring image resolution in optical nanoscopy. Nat. Methods 10, 557–562 (2013).
Mollenhauer, H.H. Artifacts caused by dehydration and epoxy embedding in transmission electron microscopy. Microsc. Res. Tech. 26, 496–512 (1993).
Acknowledgements
We thank R. Fetter and W.-P. Li of the HHMI-Janelia Research Campus for assistance with electron microscopy. We thank M. Daniels and the National Heart, Lung, and Blood Institute (NHLBI) EM core facility for help with platinum replica TEM. U.S. Army Medical Research Institute of Infectious Disease (USAMRIID) disclaimer: Opinions, interpretations, conclusions and recommendations are those of the authors and are not necessarily endorsed by the U.S. Army. M.G.P.-S., G.S., C.S.X., L.L.L. and H.F.H. were supported by the Howard Hughes Medical Institute. J.W.T. was supported by the Intramural Research Program of the NHLBI, US National Institutes of Health. This research was supported in part by funds from the Boettcher Foundation's Webb-Waring Biomedical Research Awards program and The National Institute of Allergy and Infectious Disease, NIH_1R21AI127192-01 to S.B.vE.
Author information
Authors and Affiliations
Contributions
Tokuyasu protocol: B.G.K., G.S., C.S.X. and H.F.H. Whole-mount protocol: S.B.v.E., G.S. and H.F.H. Unroofing protocol: K.A.S., G.S., S.B.v.E., H.F.H. and J.W.T. Resin-embedding protocol: M.G.P.-S., M.G.S., G.S., H.F.H., Y.W. and L.L.L. iPALM/FIB-SEM protocol: C.S.X., G.S. and H.F.H. All authors contributed to writing the paper. Correspondence should be addressed to B.G.K. (kopek@hope.edu) for the Tokuyasu cryosectioning protocol, S.B.v.E. (schuyler.vanengelenburg@du.edu) for the whole-mount protocol, J.W.T. (justin.taraska@nih.gov) for the platinum replica/TEM unroofing protocol, L.L.L. (loogerl@janelia.hhmi.org) for the fluorescent protein engineering/resin embedding protocol and H.F.H. (hessh@janelia.hhmi.org) for iPALM and FIB-SEM.
Corresponding authors
Ethics declarations
Competing interests
Harald Hess is part owner of a patent and license on PALM.
Supplementary information
Supplementary Note
Supplementary Note. (PDF 128 kb)
Rights and permissions
About this article
Cite this article
Kopek, B., Paez-Segala, M., Shtengel, G. et al. Diverse protocols for correlative super-resolution fluorescence imaging and electron microscopy of chemically fixed samples. Nat Protoc 12, 916–946 (2017). https://doi.org/10.1038/nprot.2017.017
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2017.017
This article is cited by
-
Focal adhesions contain three specialized actin nanoscale layers
Nature Communications (2024)
-
Label-free nanoscale mapping of intracellular organelle chemistry
Communications Biology (2023)
-
Reconstruction of ovine axonal cytoarchitecture enables more accurate models of brain biomechanics
Communications Biology (2022)
-
Landmark-based retrieval of inflamed skin vessels enabled by 3D correlative intravital light and volume electron microscopy
Histochemistry and Cell Biology (2022)
-
Chip-based multimodal super-resolution microscopy for histological investigations of cryopreserved tissue sections
Light: Science & Applications (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.