Abstract
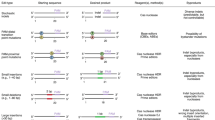
Genome editing harnesses programmable nucleases to cut and paste genetic information in a targeted manner in living cells and organisms. Here, I review the development of programmable nucleases, including zinc finger nucleases (ZFNs), TAL (transcription-activator-like) effector nucleases (TALENs) and CRISPR (cluster of regularly interspaced palindromic repeats)–Cas9 (CRISPR-associated protein 9) RNA-guided endonucleases (RGENs). I specifically highlight the key advances that set the foundation for the rapid and widespread implementation of CRISPR–Cas9 genome editing approaches that has revolutionized the field.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Kim, H. & Kim, J.S. A guide to genome engineering with programmable nucleases. Nat. Rev. Genet. 15, 321–334 (2014).
Cho, S.W., Kim, S., Kim, J.M. & Kim, J.S. Targeted genome engineering in human cells with the Cas9 RNA-guided endonuclease. Nat. Biotechnol. 31, 230–232 (2013).
Cong, L. et al. Multiplex genome engineering using CRISPR/Cas systems. Science 339, 819–823 (2013).
Mali, P. et al. RNA-guided human genome engineering via Cas9. Science 339, 823–826 (2013).
Jinek, M. et al. RNA-programmed genome editing in human cells. eLife 2, e00471 (2013).
Cho, S.W., Lee, J., Carroll, D., Kim, J.S. & Lee, J. Heritable gene knockout in Caenorhabditis elegans by direct injection of Cas9-sgRNA ribonucleoproteins. Genetics 195, 1177–1180 (2013).
Hwang, W.Y. et al. Efficient genome editing in zebrafish using a CRISPR-Cas system. Nat. Biotechnol. 31, 227–229 (2013).
Wang, H. et al. One-step generation of mice carrying mutations in multiple genes by CRISPR/Cas-mediated genome engineering. Cell 153, 910–918 (2013).
Sung, Y.H. et al. Highly efficient gene knockout in mice and zebrafish with RNA-guided endonucleases. Genome Res. 24, 125–131 (2014).
Shan, Q. et al. Targeted genome modification of crop plants using a CRISPR-Cas system. Nat. Biotechnol. 31, 686–688 (2013).
Woo, J.W. et al. DNA-free genome editing in plants with preassembled CRISPR-Cas9 ribonucleoproteins. Nat. Biotechnol. 33, 1162–1164 (2015).
Gilbert, L.A. et al. CRISPR-mediated modular RNA-guided regulation of transcription in eukaryotes. Cell 154, 442–451 (2013).
Qi, L.S. et al. Repurposing CRISPR as an RNA-guided platform for sequence-specific control of gene expression. Cell 152, 1173–1183 (2013).
Hilton, I.B. et al. Epigenome editing by a CRISPR-Cas9-based acetyltransferase activates genes from promoters and enhancers. Nat. Biotechnol. 33, 510–517 (2015).
Thakore, P.I. et al. Highly specific epigenome editing by CRISPR-Cas9 repressors for silencing of distal regulatory elements. Nat. Methods 12, 1143–1149 (2015).
Gaj, T., Gersbach, C.A. & Barbas, C.F. III. ZFN, TALEN, and CRISPR/Cas-based methods for genome engineering. Trends Biotechnol. 31, 397–405 (2013).
Capecchi, M.R. Gene targeting in mice: functional analysis of the mammalian genome for the twenty-first century. Nat. Rev. Genet. 6, 507–512 (2005).
Rudin, N. & Haber, J.E. Efficient repair of HO-induced chromosomal breaks in Saccharomyces cerevisiae by recombination between flanking homologous sequences. Mol. Cell. Biol. 8, 3918–3928 (1988).
Rouet, P., Smih, F. & Jasin, M. Introduction of double-strand breaks into the genome of mouse cells by expression of a rare-cutting endonuclease. Mol. Cell. Biol. 14, 8096–8106 (1994).
Epinat, J.C. et al. A novel engineered meganuclease induces homologous recombination in yeast and mammalian cells. Nucleic Acids Res. 31, 2952–2962 (2003).
Li, L., Wu, L.P. & Chandrasegaran, S. Functional domains in Fok I restriction endonuclease. Proc. Natl. Acad. Sci. USA 89, 4275–4279 (1992).
Kim, Y.G. & Chandrasegaran, S. Chimeric restriction endonuclease. Proc. Natl. Acad. Sci. USA 91, 883–887 (1994).
Kim, Y.G., Cha, J. & Chandrasegaran, S. Hybrid restriction enzymes: zinc finger fusions to Fok I cleavage domain. Proc. Natl. Acad. Sci. USA 93, 1156–1160 (1996).
Pavletich, N.P. & Pabo, C.O. Zinc finger-DNA recognition: crystal structure of a Zif268-DNA complex at 2.1 A. Science 252, 809–817 (1991).
Desjarlais, J.R. & Berg, J.M. Redesigning the DNA-binding specificity of a zinc finger protein: a data base-guided approach. Proteins 12, 101–104 (1992).
Rebar, E.J. & Pabo, C.O. Zinc finger phage: affinity selection of fingers with new DNA-binding specificities. Science 263, 671–673 (1994).
Bibikova, M., Golic, M., Golic, K.G. & Carroll, D. Targeted chromosomal cleavage and mutagenesis in Drosophila using zinc-finger nucleases. Genetics 161, 1169–1175 (2002).
Bibikova, M., Beumer, K., Trautman, J.K. & Carroll, D. Enhancing gene targeting with designed zinc finger nucleases. Science 300, 764 (2003).
Morton, J., Davis, M.W., Jorgensen, E.M. & Carroll, D. Induction and repair of zinc-finger nuclease-targeted double-strand breaks in Caenorhabditis elegans somatic cells. Proc. Natl. Acad. Sci. USA 103, 16370–16375 (2006).
Doyon, Y. et al. Heritable targeted gene disruption in zebrafish using designed zinc-finger nucleases. Nat. Biotechnol. 26, 702–708 (2008).
Meng, X., Noyes, M.B., Zhu, L.J., Lawson, N.D. & Wolfe, S.A. Targeted gene inactivation in zebrafish using engineered zinc-finger nucleases. Nat. Biotechnol. 26, 695–701 (2008).
Lloyd, A., Plaisier, C.L., Carroll, D. & Drews, G.N. Targeted mutagenesis using zinc-finger nucleases in Arabidopsis. Proc. Natl. Acad. Sci. USA 102, 2232–2237 (2005).
Townsend, J.A. et al. High-frequency modification of plant genes using engineered zinc-finger nucleases. Nature 459, 442–445 (2009).
Kim, H.J., Lee, H.J., Kim, H., Cho, S.W. & Kim, J.S. Targeted genome editing in human cells with zinc finger nucleases constructed via modular assembly. Genome Res. 19, 1279–1288 (2009).
Porteus, M.H. & Baltimore, D. Chimeric nucleases stimulate gene targeting in human cells. Science 300, 763 (2003).
Urnov, F.D. et al. Highly efficient endogenous human gene correction using designed zinc-finger nucleases. Nature 435, 646–651 (2005).
Maeder, M.L. et al. Rapid “open-source” engineering of customized zinc-finger nucleases for highly efficient gene modification. Mol. Cell 31, 294–301 (2008).
Ramirez, C.L. et al. Unexpected failure rates for modular assembly of engineered zinc fingers. Nat. Methods 5, 374–375 (2008).
Kim, J.S., Lee, H.J. & Carroll, D. Genome editing with modularly assembled zinc-finger nucleases. Nat. Methods 7, 91 (2010).
Kim, S., Lee, M.J., Kim, H., Kang, M. & Kim, J.S. Preassembled zinc-finger arrays for rapid construction of ZFNs. Nat. Methods 8, 7 (2011).
Miller, J.C. et al. An improved zinc-finger nuclease architecture for highly specific genome editing. Nat. Biotechnol. 25, 778–785 (2007).
Bhakta, M.S. et al. Highly active zinc-finger nucleases by extended modular assembly. Genome Res. 23, 530–538 (2013).
Sander, J.D. et al. Selection-free zinc-finger-nuclease engineering by context-dependent assembly (CoDA). Nat. Methods 8, 67–69 (2011).
Szczepek, M. et al. Structure-based redesign of the dimerization interface reduces the toxicity of zinc-finger nucleases. Nat. Biotechnol. 25, 786–793 (2007).
Gupta, A. et al. An optimized two-finger archive for ZFN-mediated gene targeting. Nat. Methods 9, 588–590 (2012).
Moscou, M.J. & Bogdanove, A.J. A simple cipher governs DNA recognition by TAL effectors. Science 326, 1501 (2009).
Boch, J. et al. Breaking the code of DNA binding specificity of TAL-type III effectors. Science 326, 1509–1512 (2009).
Miller, J.C. et al. A TALE nuclease architecture for efficient genome editing. Nat. Biotechnol. 29, 143–148 (2011).
Cermak, T. et al. Efficient design and assembly of custom TALEN and other TAL effector-based constructs for DNA targeting. Nucleic Acids Res. 39, e82 (2011).
Briggs, A.W. et al. Iterative capped assembly: rapid and scalable synthesis of repeat-module DNA such as TAL effectors from individual monomers. Nucleic Acids Res. 40, e117 (2012).
Reyon, D. et al. FLASH assembly of TALENs for high-throughput genome editing. Nat. Biotechnol. 30, 460–465 (2012).
Kim, Y. et al. A library of TAL effector nucleases spanning the human genome. Nat. Biotechnol. 31, 251–258 (2013).
Jinek, M. et al. A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science 337, 816–821 (2012).
Cradick, T.J., Fine, E.J., Antico, C.J. & Bao, G. CRISPR/Cas9 systems targeting b-globin and CCR5 genes have substantial off-target activity. Nucleic Acids Res. 41, 9584–9592 (2013).
Fu, Y. et al. High-frequency off-target mutagenesis induced by CRISPR-Cas nucleases in human cells. Nat. Biotechnol. 31, 822–826 (2013).
Pattanayak, V. et al. High-throughput profiling of off-target DNA cleavage reveals RNA-programmed Cas9 nuclease specificity. Nat. Biotechnol. 31, 839–843 (2013).
Cho, S.W. et al. Analysis of off-target effects of CRISPR/Cas-derived RNA-guided endonucleases and nickases. Genome Res. 24, 132–141 (2014).
Lin, Y. et al. CRISPR/Cas9 systems have off-target activity with insertions or deletions between target DNA and guide RNA sequences. Nucleic Acids Res. 42, 7473–7485 (2014).
Koo, T., Lee, J. & Kim, J.S. Measuring and reducing off-target activities of programmable nucleases including CRISPR-Cas9. Mol. Cells 38, 475–481 (2015).
Kim, D. et al. Digenome-seq: genome-wide profiling of CRISPR-Cas9 off-target effects in human cells. Nat. Methods 12, 237–243, 1, 243 (2015).
Tsai, S.Q. et al. GUIDE-seq enables genome-wide profiling of off-target cleavage by CRISPR-Cas nucleases. Nat. Biotechnol. 33, 187–197 (2015).
Kleinstiver, B.P. et al. High-fidelity CRISPR-Cas9 nucleases with no detectable genome-wide off-target effects. Nature 529, 490–495 (2016).
Slaymaker, I.M. et al. Rationally engineered Cas9 nucleases with improved specificity. Science 351, 84–88 (2016).
Fu, Y., Sander, J.D., Reyon, D., Cascio, V.M. & Joung, J.K. Improving CRISPR-Cas nuclease specificity using truncated guide RNAs. Nat. Biotechnol. 32, 279–284 (2014).
Tsai, S.Q. et al. Dimeric CRISPR RNA-guided FokI nucleases for highly specific genome editing. Nat. Biotechnol. 32, 569–576 (2014).
Guilinger, J.P., Thompson, D.B. & Liu, D.R. Fusion of catalytically inactive Cas9 to FokI nuclease improves the specificity of genome modification. Nat. Biotechnol. 32, 577–582 (2014).
Bolukbasi, M.F. et al. DNA-binding-domain fusions enhance the targeting range and precision of Cas9. Nat. Methods 12, 1150–1156 (2015).
Mali, P. et al. CAS9 transcriptional activators for target specificity screening and paired nickases for cooperative genome engineering. Nat. Biotechnol. 31, 833–838 (2013).
Ramirez, C.L. et al. Engineered zinc finger nickases induce homology-directed repair with reduced mutagenic effects. Nucleic Acids Res. 40, 5560–5568 (2012).
Ran, F.A. et al. Double nicking by RNA-guided CRISPR Cas9 for enhanced genome editing specificity. Cell 154, 1380–1389 (2013).
Kim, E. et al. Precision genome engineering with programmable DNA-nicking enzymes. Genome Res. 22, 1327–1333 (2012).
Gaj, T., Guo, J., Kato, Y., Sirk, S.J. & Barbas, C.F. III. Targeted gene knockout by direct delivery of zinc-finger nuclease proteins. Nat. Methods 9, 805–807 (2012).
Kim, S., Kim, D., Cho, S.W., Kim, J. & Kim, J.S. Highly efficient RNA-guided genome editing in human cells via delivery of purified Cas9 ribonucleoproteins. Genome Res. 24, 1012–1019 (2014).
Zuris, J.A. et al. Cationic lipid-mediated delivery of proteins enables efficient protein-based genome editing in vitro and in vivo. Nat. Biotechnol. 33, 73–80 (2015).
Ramakrishna, S. et al. Gene disruption by cell-penetrating peptide-mediated delivery of Cas9 protein and guide RNA. Genome Res. 24, 1020–1027 (2014).
Lee, H.J., Kim, E. & Kim, J.S. Targeted chromosomal deletions in human cells using zinc finger nucleases. Genome Res. 20, 81–89 (2010).
Lee, H.J., Kweon, J., Kim, E., Kim, S. & Kim, J.S. Targeted chromosomal duplications and inversions in the human genome using zinc finger nucleases. Genome Res. 22, 539–548 (2012).
Brunet, E. et al. Chromosomal translocations induced at specified loci in human stem cells. Proc. Natl. Acad. Sci. USA 106, 10620–10625 (2009).
Park, C.Y. et al. Targeted inversion and reversion of the blood coagulation factor 8 gene in human iPS cells using TALENs. Proc. Natl. Acad. Sci. USA 111, 9253–9258 (2014).
Park, C.Y. et al. Functional correction of large factor VIII gene chromosomal inversions in hemophilia A patient-derived iPSCs using CRISPR-Cas9. Cell Stem Cell 17, 213–220 (2015).
Young, C.S. et al. A single CRISPR-Cas9 deletion strategy that targets the majority of DMD patients restores dystrophin function in hiPSC-derived muscle cells. Cell Stem Cell 18, 533–540 (2016).
Beerli, R.R., Dreier, B. & Barbas, C.F. III. Positive and negative regulation of endogenous genes by designed transcription factors. Proc. Natl. Acad. Sci. USA 97, 1495–1500 (2000).
Park, K.S., Jang, Y.S., Lee, H. & Kim, J.S. Phenotypic alteration and target gene identification using combinatorial libraries of zinc finger proteins in prokaryotic cells. J. Bacteriol. 187, 5496–5499 (2005).
Blancafort, P., Magnenat, L. & Barbas, C.F. III. Scanning the human genome with combinatorial transcription factor libraries. Nat. Biotechnol. 21, 269–274 (2003).
Park, K.S. et al. Phenotypic alteration of eukaryotic cells using randomized libraries of artificial transcription factors. Nat. Biotechnol. 21, 1208–1214 (2003).
Komor, A.C., Kim, Y.B., Packer, M.S., Zuris, J.A. & Liu, D.R. Programmable editing of a target base in genomic DNA without double-stranded DNA cleavage. Nature 533, 420–424 (2016).
Zetsche, B. et al. Cpf1 is a single RNA-guided endonuclease of a class 2 CRISPR-Cas system. Cell 163, 759–771 (2015).
Kim, D. et al. Genome-wide analysis reveals specificities of Cpf1 endonucleases in human cells. Nat. Biotechnol. http://dx.doi.org/10.1038/nbt.3609 (2016).
Acknowledgements
This work was supported by a grant from the Institute for Basic Science in South Korea (IBS-R021-D1). I would like to thank S.H. Ye for providing Figure 2.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
J.-S.K. is a co-founder and shareholder of ToolGen, Inc., a biotechnology company focused on genome editing.
Rights and permissions
About this article
Cite this article
Kim, JS. Genome editing comes of age. Nat Protoc 11, 1573–1578 (2016). https://doi.org/10.1038/nprot.2016.104
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2016.104
This article is cited by
-
Epigenome editing: targeted manipulation of epigenetic modifications in plants
Genes & Genomics (2022)
-
Live-cell imaging of circadian clock protein dynamics in CRISPR-generated knock-in cells
Nature Communications (2021)
-
Genetic engineering in organoids
Journal of Molecular Medicine (2021)
-
Gene editing of the multi-copy H2A.B gene and its importance for fertility
Genome Biology (2019)
-
Unexpected genomic rearrangements at targeted loci associated with CRISPR/Cas9-mediated knock-in
Scientific Reports (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.