Abstract
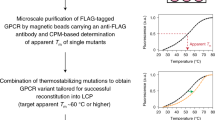
The thermostability of an integral membrane protein (MP) in detergent solution is a key parameter that dictates the likelihood of obtaining well-diffracting crystals that are suitable for structure determination. However, many mammalian MPs are too unstable for crystallization. We developed a thermostabilization strategy based on systematic mutagenesis coupled to a radioligand-binding thermostability assay that can be applied to receptors, ion channels and transporters. It takes ∼6–12 months to thermostabilize a G-protein-coupled receptor (GPCR) containing 300 amino acid (aa) residues. The resulting thermostabilized MPs are more easily crystallized and result in high-quality structures. This methodology has facilitated structure-based drug design applied to GPCRs because it is possible to determine multiple structures of the thermostabilized receptors bound to low-affinity ligands. Protocols and advice are given on how to develop thermostability assays for MPs and how to combine mutations to make an optimally stable mutant suitable for structural studies. The steps in the procedure include the generation of ∼300 site-directed mutants by Ala/Leu scanning mutagenesis, the expression of each mutant in mammalian cells by transient transfection and the identification of thermostable mutants using a thermostability assay that is based on binding of an 125I-labeled radioligand to the unpurified, detergent-solubilized MP. Individual thermostabilizing point mutations are then combined to make an optimally stable MP that is suitable for structural biology and other biophysical studies.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Bill, R.M. et al. Overcoming barriers to membrane protein structure determination. Nat. Biotechnol. 29, 335–340 (2011).
Drew, D., Lerch, M., Kunji, E., Slotboom, D.J. & de Gier, J.W. Optimization of membrane protein overexpression and purification using GFP fusions. Nat. Methods 3, 303–313 (2006).
Drew, D. et al. A scalable, GFP-based pipeline for membrane protein overexpression screening and purification. Protein Sci. 14, 2011–2017 (2005).
Hattori, M., Hibbs, R.E. & Gouaux, E. A fluorescence-detection size-exclusion chromatography-based thermostability assay for membrane protein precrystallization screening. Structure 20, 1293–1299 (2012).
Kawate, T. & Gouaux, E. Fluorescence-detection size-exclusion chromatography for precrystallization screening of integral membrane proteins. Structure 14, 673–681 (2006).
Mancia, F. & Love, J. High throughput platforms for structural genomics of integral membrane proteins. Curr. Opin. Struct. Biol. 21, 517–522 (2011).
Drew, D. et al. GFP-based optimization scheme for the overexpression and purification of eukaryotic membrane proteins in Saccharomyces cerevisiae. Nat. Protoc. 3, 784–798 (2008).
Newstead, S., Kim, H., von Heijne, G., Iwata, S. & Drew, D. High-throughput fluorescent-based optimization of eukaryotic membrane protein overexpression and purification in Saccharomyces cerevisiae. Proc. Natl. Acad. Sci. USA 104, 13936–13941 (2007).
Sonoda, Y. et al. Benchmarking membrane protein detergent stability for improving throughput of high-resolution X-ray structures. Structure 19, 17–25 (2011).
Zhang, X., Stevens, R.C. & Xu, F. The importance of ligands for G protein-coupled receptor stability. Trends Biochem. Sci. 40, 79–87 (2015).
Tate, C.G. A crystal clear solution for determining G-protein-coupled receptor structures. Trends Biochem. Sci. 37, 343–352 (2012).
Magnani, F., Shibata, Y., Serrano-Vega, M.J. & Tate, C.G. Co-evolving stability and conformational homogeneity of the human adenosine A2a receptor. Proc. Natl. Acad. Sci. USA 105, 10744–10749 (2008).
Serrano-Vega, M.J., Magnani, F., Shibata, Y. & Tate, C.G. Conformational thermostabilization of the β1-adrenergic receptor in a detergent-resistant form. Proc. Natl. Acad. Sci. USA 105, 877–882 (2008).
Shibata, Y. et al. Thermostabilization of the neurotensin receptor NTS1. J. Mol. Biol. 390, 262–277 (2009).
Sato, T. et al. Understanding the activation of the β1-adrenergic receptor through pharmacological analysis of 7-methylcyanopindolol and structure determination of the receptor-ligand complex. Mol. Pharm. 88, 1024–1034 (2015).
Warne, T. et al. The structural basis for agonist and partial agonist action on a β(1)-adrenergic receptor. Nature 469, 241–244 (2011).
Christopher, J.A. et al. Biophysical fragment screening of the β1-adrenergic receptor: identification of high affinity arylpiperazine leads using structure-based drug design. J. Med. Chem. 56, 3446–3455 (2013).
Miller-Gallacher, J.L. et al. The 2.1 Å resolution structure of cyanopindolol-bound β1-adrenoceptor identifies an intramembrane Na+ ion that stabilises the ligand-free receptor. PLoS ONE 9, e92727 (2014).
Moukhametzianov, R. et al. Two distinct conformations of helix 6 observed in antagonist-bound structures of a β1-adrenergic receptor. Proc. Natl. Acad. Sci. USA 108, 8228–8232 (2011).
Warne, T. et al. Structure of a β1-adrenergic G-protein-coupled receptor. Nature 454, 486–491 (2008).
Warne, T., Edwards, P.C., Leslie, A.G. & Tate, C.G. Crystal structures of a stabilized β1-adrenoceptor bound to the biased agonists bucindolol and carvedilol. Structure 20, 841–849 (2012).
Lebon, G., Edwards, P.C., Leslie, A.G. & Tate, C.G. Molecular determinants of CGS21680 binding to the human adenosine A2A receptor. Mol. Pharmacol. 87, 907–915 (2015).
Lebon, G. et al. Agonist-bound adenosine A2A receptor structures reveal common features of GPCR activation. Nature 474, 521–525 (2011).
Congreve, M. et al. Discovery of 1,2,4-triazine derivatives as adenosine A(2A) antagonists using structure based drug design. J. Med. Chem. 55, 1898–1903 (2012).
Dore, A.S. et al. Structure of the adenosine A(2A) receptor in complex with ZM241385 and the xanthines XAC and caffeine. Structure 19, 1283–1293 (2011).
White, J.F. et al. Structure of the agonist-bound neurotensin receptor. Nature 490, 508–513 (2012).
Egloff, P. et al. Structure of signaling-competent neurotensin receptor 1 obtained by directed evolution in Escherichia coli. Proc. Natl. Acad. Sci. USA 111, E655–E662 (2014).
Krumm, B.E., White, J.F., Shah, P. & Grisshammer, R. Structural prerequisites for G-protein activation by the neurotensin receptor. Nat. Commun. 6, 7895 (2015).
Leslie, A.G., Warne, T. & Tate, C.G. Ligand occupancy in crystal structure of β1-adrenergic G protein-coupled receptor. Nat. Struct. Mol. Biol. 22, 941–942 (2015).
Robertson, N. et al. The properties of thermostabilised G protein-coupled receptors (StaRs) and their use in drug discovery. Neuropharmacology 60, 36–44 (2011).
Hollenstein, K. et al. Structure of class B GPCR corticotropin-releasing factor receptor 1. Nature 499, 438–443 (2013).
Dore, A.S. et al. Structure of class C GPCR metabotropic glutamate receptor 5 transmembrane domain. Nature 511, 557–562 (2014).
Tan, Q. et al. Structure of the CCR5 chemokine receptor-HIV entry inhibitor maraviroc complex. Science 341, 1387–1390 (2013).
Srivastava, A. et al. High-resolution structure of the human GPR40 receptor bound to allosteric agonist TAK-875. Nature 513, 124–127 (2014).
Penmatsa, A., Wang, K.H. & Gouaux, E. X-ray structure of dopamine transporter elucidates antidepressant mechanism. Nature 503, 85–90 (2013).
Chen, L., Durr, K.L. & Gouaux, E. X-ray structures of AMPA receptor-cone snail toxin complexes illuminate activation mechanism. Science 345, 1021–1026 (2014).
Tucker, J. & Grisshammer, R. Purification of a rat neurotensin receptor expressed in Escherichia coli. Biochem. J. 317, 891–899 (1996).
Warne, T., Chirnside, J. & Schertler, G.F. Expression and purification of truncated, non-glycosylated turkey beta-adrenergic receptors for crystallization. Biochim. Biophys. Acta 1610, 133–140 (2003).
Weiss, H.M. & Grisshammer, R. Purification and characterization of the human adenosine A(2a) receptor functionally expressed in Escherichia coli. Eur. J. Biochem. 269, 82–92 (2002).
Lebon, G., Bennett, K., Jazayeri, A. & Tate, C.G. Thermostabilisation of an agonist-bound conformation of the human adenosine A(2A) receptor. J. Mol. Biol. 409, 298–310 (2011).
Miller, J.L. & Tate, C.G. Engineering an ultra-thermostable β(1)-adrenoceptor. J. Mol. Biol. 413, 628–638 (2011).
Serrano-Vega, M.J. & Tate, C.G. Transferability of thermostabilizing mutations between beta-adrenergic receptors. Mol. Membr. Biol. 26, 385–396 (2009).
Shibata, Y. et al. Optimising the combination of thermostabilising mutations in the neurotensin receptor for structure determination. Biochim. Biophys. Acta 1828, 1293–1301 (2013).
Abdul-Hussein, S., Andrell, J. & Tate, C.G. Thermostabilisation of the serotonin transporter in a cocaine-bound conformation. J. Mol. Biol. 425, 2198–2207 (2013).
AbdulHussein, S. Conformational thermostabilisation of a mammalian serotonin transporter. PhD thesis, University of Cambridge, UK, (2011).
Andrell, J. & Tate, C.G. Overexpression of membrane proteins in mammalian cells for structural studies. Mol. Membr. Biol. 30, 52–63 (2013).
Grisshammer, R. & Tate, C.G. Overexpression of integral membrane proteins for structural studies. Q. Rev. Biophys. 28, 315–422 (1995).
Tate, C.G. Overexpression of mammalian integral membrane proteins for structural studies. FEBS Lett. 504, 94–98 (2001).
Thomas, J. & Tate, C.G. Quality control in eukaryotic membrane protein overproduction. J. Mol. Biol. 426, 4139–4154 (2014).
Tate, C.G. et al. Comparison of seven different heterologous protein expression systems for the production of the serotonin transporter. Biochim. Biophys. Acta 1610, 141–153 (2003).
Tate, C.G. & Blakely, R.D. The effect of N-linked glycosylation on activity of the Na(+)- and Cl(-)-dependent serotonin transporter expressed using recombinant baculovirus in insect cells. J. Biol. Chem. 269, 26303–26310 (1994).
Tate, C.G., Whiteley, E. & Betenbaugh, M.J. Molecular chaperones stimulate the functional expression of the cocaine-sensitive serotonin transporter. J. Biol. Chem. 274, 17551–17558 (1999).
Tate, C.G. Practical considerations of membrane protein instability during purification and crystallisation. Methods Mol. Biol. 601, 187–203 (2010).
Helenius, A. & Simons, K. Solubilization of membranes by detergents. Biochim. Biophys. Acta 415, 29–79 (1975).
le Maire, M., Champeil, P. & Moller, J.V. Interaction of membrane proteins and lipids with solubilizing detergents. Biochim. Biophys. Acta 1508, 86–111 (2000).
Seddon, A.M., Curnow, P. & Booth, P.J. Membrane proteins, lipids and detergents: not just a soap opera. Biochim. Biophys. Acta 1666, 105–117 (2004).
Toyoshima, C., Nakasako, M., Nomura, H. & Ogawa, H. Crystal structure of the calcium pump of sarcoplasmic reticulum at 2.6 A resolution. Nature 405, 647–655 (2000).
Chae, P.S. et al. A new class of amphiphiles bearing rigid hydrophobic groups for solubilization and stabilization of membrane proteins. Chemistry 18, 9485–9490 (2012).
Butler, P.J., Ubarretxena-Belandia, I., Warne, T. & Tate, C.G. The Escherichia coli multidrug transporter EmrE is a dimer in the detergent-solubilised state. J. Mol. Biol. 340, 797–808 (2004).
Warne, T., Serrano-Vega, M.J., Tate, C.G. & Schertler, G.F. Development and crystallization of a minimal thermostabilised G protein-coupled receptor. Protein Expr. Purif. 65, 204–213 (2009).
Zhukov, A. et al. Biophysical mapping of the adenosine A2A receptor. J. Med. Chem. 54, 4312–4323 (2011).
Rosenbaum, D.M. et al. GPCR engineering yields high-resolution structural insights into beta2-adrenergic receptor function. Science 318, 1266–1273 (2007).
Steyaert, J. & Kobilka, B.K. Nanobody stabilization of G protein-coupled receptor conformational states. Curr. Opin. Struct. Biol. 21, 567–572 (2011).
Caffrey, M. Crystallizing membrane proteins for structure determination: use of lipidic mesophases. Annu. Rev. Biophys. 38, 29–51 (2009).
Zhou, Y. & Bowie, J.U. Building a thermostable membrane protein. J. Biol. Chem. 275, 6975–6979 (2000).
Li, D. et al. Crystal structure of the integral membrane diacylglycerol kinase. Nature 497, 521–524 (2013).
Grisshammer, R., Duckworth, R. & Henderson, R. Expression of a rat neurotensin receptor in Escherichia coli. Biochem. J. 295, 571–576 (1993).
Sarkar, C.A. et al. Directed evolution of a G protein-coupled receptor for expression, stability, and binding selectivity. Proc. Natl. Acad. Sci. USA 105, 14808–14813 (2008).
Scott, D.J. & Pluckthun, A. Direct molecular evolution of detergent-stable G protein-coupled receptors using polymer encapsulated cells. J. Mol. Biol. 425, 662–677 (2013).
Isogai, S. et al. Backbone NMR reveals allosteric signal transduction networks in the β1-adrenergic receptor. Nature 530, 237–241 (2016).
Zhao, J., Benlekbir, S. & Rubinstein, J.L. Electron cryomicroscopy observation of rotational states in a eukaryotic V-ATPase. Nature 521, 241–245 (2015).
Allegretti, M. et al. Horizontal membrane-intrinsic α-helices in the stator a-subunit of an F-type ATP synthase. Nature 521, 237–240 (2015).
Congreve, M., Langmead, C.J., Mason, J.S. & Marshall, F.H. Progress in structure based drug design for G protein-coupled receptors. J. Med. Chem. 54, 4283–4311 (2011).
Jazayeri, A., Dias, J.M. & Marshall, F.H. From G protein-coupled receptor structure resolution to rational drug design. J. Biol. Chem. 290, 19489–19495 (2015).
Bennett, K.A. et al. Pharmacology and structure of isolated conformations of the adenosine A(2)A receptor define ligand efficacy. Mol. Pharmacol. 83, 949–958 (2013).
Zheng, L., Baumann, U. & Reymond, J.L. An efficient one-step site-directed and site-saturation mutagenesis protocol. Nucleic Acids Res. 32, e115 (2004).
Palczewski, K. et al. Crystal structure of rhodopsin: a G protein-coupled receptor. Science 289, 739–745 (2000).
Tate, C.G. & Schertler, G.F. Engineering G protein-coupled receptors to facilitate their structure determination. Curr. Opin. Struct. Biol. 19, 386–395 (2009).
Alexandrov, A.I., Mileni, M., Chien, E.Y., Hanson, M.A. & Stevens, R.C. Microscale fluorescent thermal stability assay for membrane proteins. Structure 16, 351–359 (2008).
Dodevski, I. & Pluckthun, A. Evolution of three human GPCRs for higher expression and stability. J. Mol. Biol. 408, 599–615 (2011).
Schlinkmann, K.M. et al. Maximizing detergent stability and functional expression of a GPCR by exhaustive recombination and evolution. J. Mol. Biol. 422, 414–428 (2012).
Schaffner, W. & Weissmann, C. A rapid, sensitive, and specific method for the determination of protein in dilute solution. Anal. Biochem. 56, 502–514 (1973).
Acknowledgements
Funding for the thermostabilization of membrane proteins in the laboratory of C.G.T. was from the Medical Research Council (MRC U105197215), Medical Research Council Technology Development Gap Fund, Pfizer, Heptares Therapeutics and an ERC Advanced Grant (EMPSI 339995). We thank R. Henderson, F. Marshall, A. Jazayeri and M. Weir for constructive comments on the manuscript.
Author information
Authors and Affiliations
Contributions
All authors contributed to the development of techniques described in this paper. C.G.T. wrote the manuscript and coordinated contributions from all the other authors.
Corresponding author
Ethics declarations
Competing interests
C.G.T. is a consultant for Heptares Therapeutics, and this work was funded partly by Pfizer and Heptares Therapeutics.
Rights and permissions
About this article
Cite this article
Magnani, F., Serrano-Vega, M., Shibata, Y. et al. A mutagenesis and screening strategy to generate optimally thermostabilized membrane proteins for structural studies. Nat Protoc 11, 1554–1571 (2016). https://doi.org/10.1038/nprot.2016.088
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2016.088
This article is cited by
-
Stabilization and structure determination of integral membrane proteins by termini restraining
Nature Protocols (2022)
-
Universal platform for the generation of thermostabilized GPCRs that crystallize in LCP
Nature Protocols (2022)
-
The rapid “teabag” method for high-end purification of membrane proteins
Scientific Reports (2020)
-
Antibodies and venom peptides: new modalities for ion channels
Nature Reviews Drug Discovery (2019)
-
An engineered thermal-shift screen reveals specific lipid preferences of eukaryotic and prokaryotic membrane proteins
Nature Communications (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.