Abstract
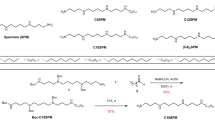
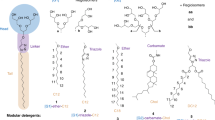
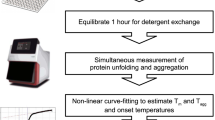
Despite the great importance of membrane proteins, structural and functional studies of these proteins present major challenges. A significant hurdle is the extraction of the functional protein from its natural lipid membrane. Traditionally achieved with detergents, purification procedures can be costly and time consuming. A critical flaw with detergent approaches is the removal of the protein from the native lipid environment required to maintain functionally stable protein. This protocol describes the preparation of styrene maleic acid (SMA) co-polymer to extract membrane proteins from prokaryotic and eukaryotic expression systems. Successful isolation of membrane proteins into SMA lipid particles (SMALPs) allows the proteins to remain with native lipid, surrounded by SMA. We detail procedures for obtaining 25 g of SMA (4 d); explain the preparation of protein-containing SMALPs using membranes isolated from Escherichia coli (2 d) and control protein-free SMALPS using E. coli polar lipid extract (1–2 h); investigate SMALP protein purity by SDS–PAGE analysis and estimate protein concentration (4 h); and detail biophysical methods such as circular dichroism (CD) spectroscopy and sedimentation velocity analytical ultracentrifugation (svAUC) to undertake initial structural studies to characterize SMALPs (∼2 d). Together, these methods provide a practical tool kit for those wanting to use SMALPs to study membrane proteins.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Filmore, D. It's a GPCR world. Mod. Drug Discov. 7, 24–28 (2004).
Arinaminpathy, Y., Khurana, E., Engelman, D.M. & Gerstein, M.B. Computational analysis of membrane proteins: the largest class of drug targets. Drug Discov. Today 14, 1130–1135 (2009).
Seddon, A.M., Curnow, P. & Booth, P.J. Membrane proteins, lipids and detergents: not just a soap opera. Biochim. Biophys. Acta 1666, 105–117 (2004).
le Maire, M., Champeil, P. & Moller, J.V. Interaction of membrane proteins and lipids with solubilizing detergents. Biochim. Biophys. Acta 1508, 86–111 (2000).
Gohon, Y. & Popot, J.-L. Membrane protein–surfactant complexes. Curr. Opin. Colloid Interface Sci. 8, 15–22 (2003).
Lin, S.-H. & Guidotti, G. Purification of membrane proteins. Methods Enzymol. 463, 619–629 (2009).
Damianoglou, A. et al. The synergistic action of melittin and phospholipase A2 with lipid membranes: development of linear dichroism for membrane-insertion kinetics. Protein Pept. Lett. 17, 1351–1362 (2010).
Charalambous, K., Miller, D., Curnow, P. & Booth, P.J. Lipid bilayer composition influences small multidrug transporters. BMC Biochem. 9, 31 (2008).
Wiener, M.C. & White, S.H. Structure of a fluid dioleoylphosphatidylcholine bilayer determined by joint refinement of X-ray and neutron diffraction data. III. Complete structure. Biophys. J. 61, 434–447 (1992).
Simons, K. & Sampaio, J.L. Membrane organization and lipid rafts. Cold Spring Harb. Perspect. Biol. 3, a004697 (2011).
Kellosalo, J., Kajander, T., Honkanen, R. & Goldman, A. Crystallization and preliminary X-ray analysis of membrane-bound pyrophosphatases. Mol. Membr. Biol. 30, 64–74 (2013).
Breyton, C., Tribet, C., Olive, J., Dubacq, J.P. & Popot, J.L. Dimer to monomer conversion of the cytochrome b6 f complex. Causes and consequences. J. Biol. Chem. 272, 21892–21900 (1997).
Esmann, M. Solubilized (Na+ + K+)-ATPase from shark rectal gland and ox kidney--an inactivation study. Biochim. Biophys. Acta 857, 38–47 (1986).
Popot, J.-L. Amphipols, nanodiscs, and fluorinated surfactants: three nonconventional approaches to studying membrane proteins in aqueous solutions. Annu. Rev. Biochem. 79, 737–775 (2010).
Zoonens, M. & Popot, J.-L. Amphipols for each season. J. Membr. Biol. 247, 759–796 (2014).
Debnath, D.K., Basaiawmoit, R.V., Nielsen, K.L. & Otzen, D.E. The role of membrane properties in Mistic folding and dimerisation. Protein Eng. Des. Sel. 24, 89–97 (2011).
Denisov, I.G., Grinkova, Y.V., Lazarides, A.A. & Sligar, S.G. Directed self-assembly of monodisperse phospholipid bilayer nanodiscs with controlled size. J. Am. Chem. Soc. 126, 3477–3487 (2004).
Luthra, A., Gregory, M., Grinkova, Y.V., Denisov, I.G. & Sligar, S.G. Nanodiscs in the studies of membrane-bound cytochrome P450 enzymes. Methods Mol. Biol. 987, 115–127 (2013).
Alami, M., Dalal, K., Lelj-Garolla, B., Sligar, S.G. & Duong, F. Nanodiscs unravel the interaction between the SecYEG channel and its cytosolic partner SecA. EMBO J. 26, 1995–2004 (2007).
Leitz, A.J., Bayburt, T.H., Barnakov, A.N., Springer, B.A. & Sligar, S.G. Functional reconstitution of Beta2-adrenergic receptors utilizing self-assembling nanodisc technology. Biotechniques 40 601–2, 604, 606, passim (2006).
Knowles, T.J. et al. Membrane proteins solubilized intact in lipid containing nanoparticles bounded by styrene maleic acid copolymer. J. Am. Chem. Soc. 131, 7484–7485 (2009).
Jamshad, M. et al. G-protein coupled receptor solubilization and purification for biophysical analysis and functional studies, in the total absence of detergent. Biosci. Rep. 35 http://dx.doi.org/10.1042/BSR20140171 (2015).
Tonge, S.R. & Tighe, B.J. Responsive hydrophobically associating polymers: a review of structure and properties. Adv. Drug Deliv. Rev. 53, 109–122 (2001).
Tonge, S., Stephen, T., Vincent, R. & Tighe, B.J. Dynamic surface activity of biological fluids and ophthalmic solutions. Cornea 19, S133 (2000).
Jamshad, M. et al. Structural analysis of a nanoparticle containing a lipid bilayer used for detergent-free extraction of membrane proteins. Nano Res. 8, 774–789 (2014).
Scheidelaar, S. et al. Molecular model for the solubilization of membranes into nanodisks by styrene maleic acid copolymers. Biophys. J. 108, 279–290 (2015).
Postis, V. et al. The use of SMALPs as a novel membrane protein scaffold for structure study by negative stain electron microscopy. Biochim. Biophys. Acta 1848, 496–501 (2015).
Gulati, S. et al. Detergent-free purification of ABC (ATP-binding-cassette) transporters. Biochem. J 461, 269–278 (2014).
Dörr, J.M. et al. Detergent-free isolation, characterization, and functional reconstitution of a tetrameric K+ channel: the power of native nanodiscs. Proc. Natl. Acad. Sci. USA 111, 18607–18612 (2014).
Paulin, S. et al. Surfactant-free purification of membrane protein complexes from bacteria: application to the staphylococcal penicillin-binding protein complex PBP2/PBP2a. Nanotechnology 25, 285101 (2014).
Orwick-Rydmark, M. et al. Detergent-free incorporation of a seven-transmembrane receptor protein into nanosized bilayer Lipodisq particles for functional and biophysical studies. Nano Lett. 12, 4687–4692 (2012).
Long, A.R. et al. A detergent-free strategy for the reconstitution of active enzyme complexes from native biological membranes into nanoscale discs. BMC Biotechnol. http://dx.doi.org/10.1186/1472-6750-13-41 (2013).
Gomori, G., Colowick, S.P. & Kaplan, N.O. in Methods in Enzymology 138–148 (Academic Press, 1955).
Lin, Y. Over-expression and Biophysical Characterisation of Membrane Proteins Solubilised in a Styrene Maleic Acid Polymer http://etheses.bham.ac.uk/1738/1/Lin_11_PhD.pdf (2011).
Schuck, P. Size-distribution analysis of macromolecules by sedimentation velocity ultracentrifugation and Lamm equation modeling. Biophys. J. 78, 1606–1619 (2000).
Fotiadis, D., Harder, D. & Fotiadis, D. Preparation of detergent-solubilized membranes from Escherichia coli. Protoc. Exchange http://dx.doi.org/10.1038/protex.2012.033 (2012).
Panaretou, B. & Piper, P. in Yeast Protocols (ed. Evans, I.H.) 117–121 (Humana Press).
Jamshad, M. et al. G-protein coupled receptor solubilization and purification for biophysical analysis and functional studies, in the total absence of detergent. Biosci. Rep. 35, 1–10 (2015).
Scott, R.E. Plasma membrane vesiculation: a new technique for isolation of plasma membranes. Science 194, 743–745 (1976).
Del Piccolo, N., Placone, J., He, L., Agudelo, S.C. & Hristova, K. Production of plasma membrane vesicles with chloride salts and their utility as a cell membrane mimetic for biophysical characterization of membrane protein interactions. Anal. Chem. 84, 8650–8655 (2012).
Cohen, S., Ushiro, H., Stoscheck, C. & Chinkers, M. A native 170,000 epidermal growth factor receptor-kinase complex from shed plasma membrane vesicles. J. Biol. Chem. 257, 1523–1531 (1982).
Gulati, S. et al. Detergent free purification of ABC transporters. Biochem. J 44, 1–24 (2014).
Nordén, B., Rodger, A. & Dafforn, T. Linear Dichroism and Circular Dichroism: A Textbook on Polarized-light Spectroscopy (Royal Society of Chemistry, 2010).
Burgess, S.A., Walker, M.L., Sakakibara, H., Oiwa, K. & Knight, P.J. The structure of dynein-c by negative stain electron microscopy. J. Struct. Biol. 146, 205–216 (2004).
Booth, D.S., Avila-Sakar, A. & Cheng, Y. Visualizing proteins and macromolecular complexes by negative stain EM: from grid preparation to image acquisition. J. Vis. Exp. 58, 3227 (2011).
Grassucci, R.A., Taylor, D.J. & Frank, J. Preparation of macromolecular complexes for cryo-electron microscopy. Nat. Protoc. 2, 3239–3246 (2007).
Grassucci, R.A., Taylor, D. & Frank, J. Visualization of macromolecular complexes using cryo-electron microscopy with FEI Tecnai transmission electron microscopes. Nat. Protoc. 3, 330–339 (2008).
Scheres, S.H.W. RELION: implementation of a Bayesian approach to cryo-EM structure determination. J. Struct. Biol. 180, 519–530 (2012).
Tang, G. et al. EMAN2: an extensible image processing suite for electron microscopy. J. Struct. Biol. 157, 38–46 (2007).
Rico, A.I., Krupka, M. & Vicente, M. In the beginning, Escherichia coli assembled the proto-ring: an initial phase of division. J. Biol. Chem. 288, 20830–20836 (2013).
Hale, C.A., Rhee, A.C. & De Boer, P.A.J. ZipA-induced bundling of FtsZ polymers mediated by an interaction between C-terminal domains. J. Bacteriol. 182, 5153–5166 (2000).
Acknowledgements
This work was supported by the Biotechnology and Biosciences Research Council (BBSRC; grants BB/J017310/1, BB/H016309/1, BB/I005579/1, BB/I020349/1, BB/K004441/1 and BB/M018261/1 to T.R.D.) and the Wellcome Trust (ref. 091322/7/10/Z to V.L.G.P.). S.P.M. is supported by an MRC Career Development Award (G100567). We dedicate this work to the late Professor Stephen A. Baldwin.
Author information
Authors and Affiliations
Contributions
S.C.L., V.L.G.P., T.J.K., R.A.P., M.J., A.G., S.P.M., M.O. and T.R.D. designed the research. T.J.K., T.R.D. and M.O. devised the SMALP method. S.C.L., V.L.G.P., Y.L., M.J., R.A.P., T.J.K. and P.S. optimized the detailed protocol. S.C.L., V.L.G.P., T.J.K., A.G., R.A.P., S.P.M. and T.R.D. wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Lee, S., Knowles, T., Postis, V. et al. A method for detergent-free isolation of membrane proteins in their local lipid environment. Nat Protoc 11, 1149–1162 (2016). https://doi.org/10.1038/nprot.2016.070
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2016.070
This article is cited by
-
A nanobody-based strategy for rapid and scalable purification of human protein complexes
Nature Protocols (2024)
-
Direct purification and immobilization of his-tagged enzymes using unmodified nickel ferrite NiFe2O4 magnetic nanoparticles
Scientific Reports (2023)
-
UbiB proteins regulate cellular CoQ distribution in Saccharomyces cerevisiae
Nature Communications (2021)
-
Lipidomic and in-gel analysis of maleic acid co-polymer nanodiscs reveals differences in composition of solubilized membranes
Communications Biology (2021)
-
Cycloalkane-modified amphiphilic polymers provide direct extraction of membrane proteins for CryoEM analysis
Communications Biology (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.