Abstract
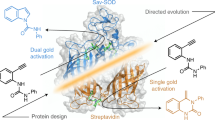
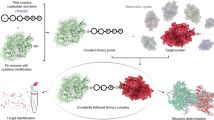
Artificial metalloenzymes (ArMs) based on the incorporation of a biotinylated metal cofactor within streptavidin (Sav) combine attractive features of both homogeneous and enzymatic catalysts. To speed up their optimization, we present a streamlined protocol for the design, expression, partial purification and screening of Sav libraries. Twenty-eight positions have been subjected to mutagenesis to yield 335 Sav isoforms, which can be expressed in 24-deep-well plates using autoinduction medium. The resulting cell-free extracts (CFEs) typically contain >1 mg of soluble Sav. Two straightforward alternatives are presented, which allow the screening of ArMs using CFEs containing Sav. To produce an artificial transfer hydrogenase, Sav is coupled to a biotinylated three-legged iridium pianostool complex Cp*Ir(Biot-p-L)Cl (the cofactor). To screen Sav variants for this application, you would determine the number of free binding sites, treat them with diamide, incubate them with the cofactor and then perform the reaction with your test compound (the example used in this protocol is 1-phenyl-3,4-dihydroisoquinoline). This process takes 20 d. If you want to perform metathesis reactions, Sav is coupled to a biotinylated second-generation Grubbs-Hoveyda catalyst. In this application, it is best to first immobilize Sav on Sepharose-iminobiotin beads and then perform washing steps. Elution from the beads is achieved in an acidic reaction buffer before incubation with the cofactor. Catalysis using your test compound (in this protocol, 2-(4-(N,N-diallylsulfamoyl)phenyl)-N,N,N-trimethylethan-1-aminium iodide) is performed using the formed metalloenzyme. Screening using this approach takes 19 d.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Yu, F. et al. Protein design: toward functional metalloenzymes. Chem. Rev. 114, 3495–3578 (2014).
Pàmies, O., Diéguez, M. & Bäckvall, J.-E. Artificial metalloenzymes in asymmetric catalysis: key developments and future directions. Adv. Synth. Catal. 357, 1567–1586 (2015).
Lewis, J.C. Artificial metalloenzymes and metallopeptide catalysts for organic synthesis. ACS Catal. 3, 2954–2975 (2013).
Ward, T.R. Artificial metalloenzymes based on the biotin–avidin technology: enantioselective catalysis and beyond. Acc. Chem. Res. 44, 47–57 (2010).
Wilson, M.E. & Whitesides, G.M. Conversion of a protein to a homogeneous asymmetric hydrogenation catalyst by site-specific modification with a diphosphinerhodium(I) moiety. J. Am. Chem. Soc. 100, 306–307 (1978).
Lin, C.-C., Lin, C.-W. & Chan, A.S.C. Catalytic hydrogenation of itaconic acid in a biotinylated pyrphos–rhodium(I) system in a protein cavity. Tetrahedron Asymmetr. 10, 1887–1893 (1999).
Reetz, M.T. Directed evolution of stereoselective hybrid catalysts. Top. Organomet. Chem. 25, 63–92 (2009).
Collot, J. et al. Artificial metalloenzymes for enantioselective catalysis based on biotin-avidin. J. Am. Chem. Soc. 125, 9030–9031 (2003).
Letondor, C., Humbert, N. & Ward, T.R. Artificial metalloenzymes based on biotin-avidin technology for the enantioselective reduction of ketones by transfer hydrogenation. Proc. Natl. Acad. Sci. USA 102, 4683–4687 (2005).
Pierron, J. et al. Artificial metalloenzymes for asymmetric allylic alkylation on the basis of the biotin–avidin technology. Angew. Chem. Int. Ed. 47, 701–705 (2008).
Hyster, T.K., Knörr, L., Ward, T.R. & Rovis, T. Biotinylated Rh(III) complexes in engineered streptavidin for accelerated asymmetric C-H activation. Science 338, 500–503 (2012).
Lo, C., Ringenberg, M.R., Gnandt, D., Wilson, Y. & Ward, T.R. Artificial metalloenzymes for olefin metathesis based on the biotin-(strept)avidin technology. Chem. Commun. 47, 12065–12067 (2011).
Pordea, A., Mathis, D. & Ward, T.R. Incorporation of biotinylated manganese-salen complexes into streptavidin: new artificial metalloenzymes for enantioselective sulfoxidation. J. Organomet. Chem. 694, 930–936 (2009).
Pordea, A. et al. Artificial metalloenzyme for enantioselective sulfoxidation based on vanadyl-loaded streptavidin. J. Am. Chem. Soc. 130, 8085–8088 (2008).
Köhler, V. et al. OsO4·streptavidin: a tunable hybrid catalyst for the enantioselective cis-dihydroxylation of olefins. Angew. Chem. Int. Ed. 50, 10863–10866 (2011).
Thomas, C.M., Letondor, C., Humbert, N. & Ward, T.R. Aqueous oxidation of alcohols catalyzed by artificial metalloenzymes based on the biotin–avidin technology. J. Organomet. Chem. 690, 4488–4491 (2005).
Sano, T. & Cantor, C.R. Expression of a cloned streptavidin gene in Escherichia coli. Proc. Natl. Acad. Sci. USA 87, 142–146 (1990).
Chilkoti, A., Tan, P.H. & Stayton, P.S. Site-directed mutagenesis studies of the high-affinity streptavidin-biotin complex: contributions of tryptophan residues 79, 108, and 120. Proc. Natl. Acad. Sci. USA 92, 1754–1758 (1995).
Sletten, E.M. & Bertozzi, C.R. Bioorthogonal chemistry: fishing for selectivity in a sea of functionality. Angew. Chem. Int. Ed. 48, 6974–6998 (2009).
Köhler, V. et al. Synthetic cascades are enabled by combining biocatalysts with artificial metalloenzymes. Nat. Chem. 5, 93–99 (2013).
Laitinen, O.H., Hytönen, V.P., Nordlund, H.R. & Kulomaa, M.S. Genetically engineered avidins and streptavidins. Cell. Mol. Life Sci. 63, 2992–3017 (2006).
Schmidt, T.G.M. & Skerra, A. The Strep-tag system for one-step purification and high-affinity detection or capturing of proteins. Nat. Protoc. 2, 1528–1535 (2007).
Wilchek, M. & Bayer, E.A. Introduction to avidin-biotin technology. Methods Enyzmol. 184, 5–13 (1990).
Reetz, M.T., Soni, P., Fernandez, L., Gumulya, Y. & Carballeira, J.D. Increasing the stability of an enzyme toward hostile organic solvents by directed evolution based on iterative saturation mutagenesis using the B-FIT method. Chem. Commun. 46, 8657–8658 (2010).
Dürrenberger, M. et al. Artificial transfer hydrogenases for the enantioselective reduction of cyclic imines. Angew. Chem. Int. Ed. 50, 3026–3029 (2011).
Wilson, Y.M., Dürrenberger, M., Nogueira, E.S. & Ward, T.R. Neutralizing the detrimental effect of glutathione on precious metal catalysts. J. Am. Chem. Soc. 136, 8928–8932 (2014).
Creus, M. et al. X-ray structure and designed evolution of an artificial transfer hydrogenase. Angew. Chem. Int. Ed. 47, 1400–1404 (2008).
Reetz, M.T., Carballeira, J.D. & Vogel, A. Iterative saturation mutagenesis on the basis of B factors as a strategy for increasing protein thermostability. Angew. Chem. Int. Ed. 45, 7745–7751 (2006).
Papworth, C., Bauer, J.C., Braman, J. & Wright, D.A. Site-directed mutagenesis in one day with >80% efficiency. Strategies 9, 3–4 (1996).
Zimbron, J.M. et al. Chemo-genetic optimization of DNA recognition by metallodrugs using a presenter-protein strategy. Chemistry 16, 12883–12889 (2010).
Zimbron, J.M. et al. A dual anchoring strategy for the localization and activation of artificial metalloenzymes based on the biotin-streptavidin technology. J. Am. Chem. Soc. 135, 5384–5388 (2013).
Zheng, L., Baumann, U. & Reymond, J.L. An efficient one-step site-directed and site-saturation mutagenesis protocol. Nucleic Acids Res. 32, e115 (2004).
Studier, F.W. Protein production by auto-induction in high-density shaking cultures. Protein Expr. Purif. 41, 207–234 (2005).
Robles, V.M. et al. Structural, kinetic, and docking studies of artificial imine reductases based on biotin–streptavidin technology: an induced lock-and-key hypothesis. J. Am. Chem. Soc. 136, 15676–15683 (2014).
Kada, G., Kaiser, K., Falk, H. & Gruber, H.J. Rapid estimation of avidin and streptavidin by fluorescence quenching or fluorescence polarization. Biochim. Biophys. Acta 1427, 44–48 (1999).
Waner, M.J. & Mascotti, D.P. A simple spectrophotometric streptavidin-biotin binding assay utilizing biotin-4-fluorescein. J. Biochem. Biophys. Methods 70, 873–877 (2008).
Wilson, Y.M., Dürrenberger, M. & Ward, T.R. Organometallic chemistry in protein scaffolds. in Protein Engineering Handbook Vol. 3 (eds. Lütz, S. & Bornscheuer, U.T.) 215–238 (Wiley-VCH, Weinheim, 2012).
Bennett, B.D. et al. Absolute metabolite concentrations and implied enzyme active site occupancy in Escherichia coli. Nat. Chem. Biol. 5, 593–599 (2009).
Kajetanowicz, A., Chatterjee, A., Reuter, R. & Ward, T. Biotinylated metathesis catalysts: synthesis and performance in ring closing metathesis. Catal. Lett. 144, 373–379 (2014).
Lantos, I., Bhattacharjee, D. & Eggleston, D.S. Synthesis of 1-phenyl-5H-2-benzazepines by ring expansion of 1-phenyl-1,2-dihydroisoquinolines. J. Org. Chem. 51, 4147–4150 (1986).
Bertani, G. Studies on lysogenesis. I. The mode of phage liberation by lysogenic Escherichia coli. J. Bacteriol. 62, 293–300 (1951).
Hanahan, D. Studies on transformation of Escherichia coli with plasmids. J. Mol. Biol. 166, 557–580 (1983).
Hanahan, D., Jessee, J. & Bloom, F.R. Techniques for transformation of E. coli. in DNA Cloning (Oxford University Press, 1995).
Muto, T. et al. Preparation of 1,4-diazepane-3,5-dione derivatives as chymase inhibitors and pharmaceutical use thereof. Japanese patent no. WO 2010053182A1 (2010).
Becker, G.O. et al. Organikum 19th edn. (Barth Verlagsgesellschaft, 1993).
Angov, E. Codon usage: nature's roadmap to expression and folding of proteins. Biotechnol. J. 6, 650–659 (2011).
Acknowledgements
T.R.W. thanks the Swiss National Science Foundation (grants 200020_162348 and the NCCR (National Centres of Competence in Research) Molecular Systems Engineering) and the Seventh Framework Programme Project METACODE (KBBE (Knowledge-Based BioEconomy), 'Code-engineered new-to nature microbial cell factories for novel and safety enhanced bioproduction') and the US National Institutes of Health (NIH; grant GM050781) for generous support. M.H. thanks the SNI (Swiss Nanoscience Institute) for a Ph.D. scholarship. The authors are happy to provide the library free of charge upon request to academic institutions.
Author information
Authors and Affiliations
Contributions
T.R.W. conceived and established the concept of ArMs using the Sav-biotin technology and planned the experiments. H.M. planned experiments, designed the mutant library and established the expression protocol. M.H. and R.R. designed and performed the screening catalysis experiments. T.R.W., H.M., M.H. and R.R. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–7, Supplementary Methods, Supplementary Table 1 and Supplementary Data (PDF 1571 kb)
Rights and permissions
About this article
Cite this article
Mallin, H., Hestericová, M., Reuter, R. et al. Library design and screening protocol for artificial metalloenzymes based on the biotin-streptavidin technology. Nat Protoc 11, 835–852 (2016). https://doi.org/10.1038/nprot.2016.019
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2016.019
This article is cited by
-
The importance of catalytic promiscuity for enzyme design and evolution
Nature Reviews Chemistry (2019)
-
Recent advances in the engineering and application of streptavidin-like molecules
Applied Microbiology and Biotechnology (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.