Abstract
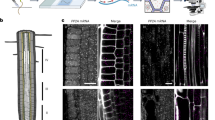
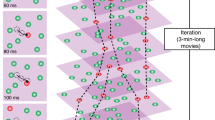
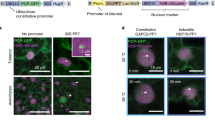
Measuring the mobility and interactions of proteins is key to understanding cellular signaling mechanisms; however, quantitative analysis of protein dynamics in living plant cells remains a major challenge. Here we describe an automated, single-molecule protocol based on total internal reflection fluorescence microscopy (TIRFM) imaging that allows protein tracking and subunit counting in living plant cells. This protocol uses TIRFM to image transgenic plant tissues expressing fluorescently tagged proteins that are localized to the plasma membrane. Next, a tracking algorithm quantifies dynamic changes in fluorescent protein motion types, temporary particle displacement and protein photobleaching steps. This protocol allows researchers to study the kinetic characteristics of heterogeneously distributed proteins. The approach has potential applications for studies of protein dynamics and subunit stoichiometry for a wide variety of plasma membrane and intracellular proteins in living plant cells and other biological specimens visualized by TIRFM or other fluorescence imaging techniques. The whole protocol can be completed in 5–6 h.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Jaqaman, K. et al. Robust single-particle tracking in live-cell time-lapse sequences. Nat. Methods 5, 695–702 (2008).
Parthasarathy, R. Rapid, accurate particle tracking by calculation of radial symmetry centers. Nat. Methods 9, 724–726 (2012).
Fan, L. et al. Dynamic analysis of Arabidopsis AP2 sigma subunit reveals a key role in clathrin-mediated endocytosis and plant development. Development 140, 3826–3837 (2013).
Li, X. et al. Single-molecule analysis of PIP2;1 dynamics and partitioning reveals multiple modes of Arabidopsis plasma membrane aquaporin regulation. Plant Cell 23, 3780–3897 (2011).
Li, R. et al. A membrane microdomain-associated protein, Arabidopsis Flot1, is involved in a clathrin-independent endocytic pathway and is required for seedling development. Plant Cell 24, 2105–2122 (2012).
Wang, Q. et al. Single-particle analysis reveals shutoff control of the Arabidopsisammonium transporter AMT1;3 by clustering and internalization. Proc. Natl. Acad. Sci. USA 110, 13204–13209 (2013).
Hao, H. et al. Clathrin and membrane microdomains cooperatively regulate RbohD dynamics and activity in Arabidopsis. Plant Cell 26, 1729–1745 (2014).
Meijering, E., Smal, I. & Danuser, G. Tracking in molecular bioimaging. Signal Process. Mag. IEEE 23, 46–53 (2006).
Serge, A., Bertaux, N., Rigneault, H. & Marguet, D. Dynamic multiple-target tracing to probe spatiotemporal cartography of cell membranes. Nat. Methods 5, 687–694 (2008).
Kalaidzidis, Y. Intracellular objects tracking. Eur. J. Cell Biol. 86, 569–578 (2007).
Chenouard, N. et al. Objective comparison of particle tracking methods. Nat. Methods 11, 281–289 (2014).
Chenouard, N., Bloch, I. & Olivo-Marin, J.C. Multiple hypothesis tracking in microscopy images. IEEE International Symposium on Biomedical Imaging: From Nano to Macro, ISBI'09, 1346–1349 (2009).
Jonker, R. & Volgenant, A. A shortest augmenting path algorithm for dense and sparse linear assignment problems. Computing 38, 325–340 (1987).
Vizcay-Barrena, G., Webb, S.E., Martin-Fernandez, M.L. & Wilson, Z.A. Subcellular and single-molecule imaging of plant fluorescent proteins using total internal reflection fluorescence microscopy (TIRFM). J. Exp. Bot. 62, 5419–5428 (2011).
Martiniere, A. et al. Cell wall constrains lateral diffusion of plant plasma-membrane proteins. Proc. Natl. Acad. Sci. USA 109, 12805–12810 (2012).
Konopka, C.A. & Bednarek, S.Y. Variable-angle epifluorescence microscopy: a new way to look at protein dynamics in the plant cell cortex. Plant J. 53, 186–196 (2008).
Konopka, C.A., Backues, S.K. & Bednarek, S.Y. Dynamics of Arabidopsis dynamin-related protein 1C and a clathrin light chain at the plasma membrane. Plant Cell 20, 1363–1380 (2008).
Zacharias, D.A., Violin, J.D., Newton, A.C. & Tsien, R.Y. Partitioning of lipid-modified monomeric GFPs into membrane microdomains of live cells. Science 296, 913–916 (2002).
Uttenweiler, D., Weber, C., Jahne, B., Fink, R.H. & Scharr, H. Spatiotemporal anisotropic diffusion filtering to improve signal-to-noise ratios and object restoration in fluorescence microscopic image sequences. J. Biomed. Opt. 8, 40–47 (2003).
Gohring, J., Fulcher, N., Schilcher, K., Barta, A. & Jacak, J. Suitable transfection methods for single particle tracing in plant suspension cells. Plant Methods 10, 15 (2014).
Ulbrich, M.H. & Isacoff, E.Y. Subunit counting in membrane-bound proteins. Nat. Methods 4, 319–321 (2007).
McGuire, H., Aurousseau, M.R., Bowie, D. & Blunck, R. Automating single subunit counting of membrane proteins in mammalian cells. J. Biol. Chem. 287, 35912–35921 (2012).
Ge, Y., Matov, A. & Danuser, G. Reliable tracking of large-scale dense antiparallel particle motion for fluorescence live cell imaging. In IEEE Computer Society Conference on Computer Vision and Pattern Recognition - Workshops, 2005. CVPR Workshops. 138 (2005).
Acknowledgements
We thank K. Jaqaman and R. Parthasarathy for providing the original algorithm source code. This work is supported by the Program of Introducing Talents of Discipline to Universities (111 project, B13007), the Major Science Foundation of the Ministry of Education of China (no. 313008), the National Basic Research Program of China (973 Program 2011CB809103) and the National Nature Science Foundation of China Project (grant nos. 31270412 and 31270224).
Author information
Authors and Affiliations
Contributions
X.W. and X.L. combined the MATLAB scripts and wrote the paper. X.D. and D.-T.L. performed the computational analysis. C.M. edited and revised the paper. J.L. designed the experiment and revised the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Software 1
MATLAB source code. (ZIP 18676 kb)
Rights and permissions
About this article
Cite this article
Wang, X., Li, X., Deng, X. et al. Single-molecule fluorescence imaging to quantify membrane protein dynamics and oligomerization in living plant cells. Nat Protoc 10, 2054–2063 (2015). https://doi.org/10.1038/nprot.2015.132
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2015.132
This article is cited by
-
VA-TIRFM-based SM kymograph analysis for dwell time and colocalization of plasma membrane protein in plant cells
Plant Methods (2023)
-
Multiple functions of the vacuole in plant growth and fruit quality
Molecular Horticulture (2021)
-
Determination of G-protein–coupled receptor oligomerization by molecular brightness analyses in single cells
Nature Protocols (2021)
-
Single-particle tracking photoactivated localization microscopy of membrane proteins in living plant tissues
Nature Protocols (2021)
-
Mass photometry enables label-free tracking and mass measurement of single proteins on lipid bilayers
Nature Methods (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.