Abstract
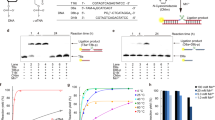
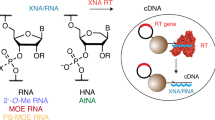
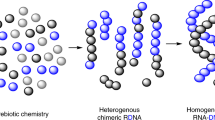
This protocol describes the directed evolution of artificial endonuclease and ligase enzymes composed of synthetic genetic polymers (XNAzymes), using 'cross-chemistry selective enrichment by exponential amplification' (X-SELEX). The protocol is analogous to (deoxy)ribozyme selections, but it enables the development of fully substituted catalysts. X-SELEX is initiated by the synthesis of diverse repertoires (here 1014 different sequences), using xeno nucleic acid (XNA) polymerases, on DNA templates primed with DNA, RNA or XNA oligonucleotides that double as substrates, allowing selection for XNA-catalyzed cleavage or ligation. XNAzymes are reverse-transcribed into cDNA using XNA-dependent DNA polymerases, and then PCR-amplified to generate templates for subsequent rounds or deep sequencing. We describe methods developed for four XNA chemistries, arabino nucleic acids (ANAs), 2′-fluoroarabino nucleic acids (FANAs), hexitol nucleic acids (HNAs) and cyclohexene nucleic acids (CeNAs), which require ∼1 week per round, and typically 10–20 rounds; in principle, these methods are scalable and applicable to a wide range of novel XNAzyme chemistries, substrates and reactions.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Pinheiro, V.B. & Holliger, P. The XNA world: progress towards replication and evolution of synthetic genetic polymers. Curr. Opin. Chem. Biol. 16, 245–252 (2012).
Pinheiro, V.B. et al. Synthetic genetic polymers capable of heredity and evolution. Science 336, 341–344 (2012).
Cozens, C., Pinheiro, V.B., Vaisman, A., Woodgate, R. & Holliger, P. A short adaptive path from DNA to RNA polymerases. Proc. Natl. Acad. Sci. USA 109, 8067–8072 (2012).
Taylor, A.I. et al. Catalysts from synthetic genetic polymers. Nature 518, 427–430 (2015).
Silverman, S.K. In vitro selection, characterization, and application of deoxyribozymes that cleave RNA. Nucleic Acids Res. 33, 6151–6163 (2005).
Keefe, A.D. & Cload, S.T. SELEX with modified nucleotides. Curr. Opin. Chem. Biol. 12, 448–456 (2008).
Vaught, J.D. et al. Expanding the chemistry of DNA for in vitro selection. J. Am. Chem. Soc. 132, 4141–4151 (2010).
Lin, Y., Qiu, Q., Gill, S.C. & Jayasena, S.D. Modified RNA sequence pools for in vitro selection. Nucleic Acids Res. 22, 5229–5234 (1994).
Hollenstein, M., Hipolito, C.J., Lam, C.H. & Perrin, D.M. Toward the combinatorial selection of chemically modified DNAzyme RNase A mimics active against all-RNA substrates. ACS Comb. Sci. 15, 174–182 (2013).
Renders, M., Miller, E., Hollenstein, M. & Perrin, D. A method for selecting modified DNAzymes without the use of modified DNA as a template in PCR. Chem. Commun. 51, 1360–1362 (2015).
Burnett, J.C. & Rossi, J.J. RNA-based therapeutics: current progress and future prospects. Chem. Biol. 19, 60–71 (2012).
Taylor, A.I., Arangundy-Franklin, S. & Holliger, P. Towards applications of synthetic genetic polymers in diagnosis and therapy. Curr. Opin. Chem. Biol. 22, 79–84 (2014).
Santoro, S.W. & Joyce, G.F. A general purpose RNA-cleaving DNA enzyme. Proc. Natl. Acad. Sci. USA 94, 4262–4266 (1997).
Wu, Y. et al. Inhibition of bcr-abl oncogene expression by novel deoxyribozymes (DNAzymes). Hum. Gene Ther. 10, 2847–2857 (1999).
Lan, N., Howrey, R.P., Lee, S.W., Smith, C.A. & Sullenger, B.A. Ribozyme-mediated repair of sickle β-globin mRNAs in erythrocyte precursors. Science 280, 1593–1596 (1998).
Breaker, R.R. & Joyce, G.F. The expanding view of RNA and DNA function. Chem. Biol. 21, 1059–1065 (2014).
Herdewijn, P. & Marliere, P. Toward safe genetically modified organisms through the chemical diversification of nucleic acids. Chem. Biodivers. 6, 791–808 (2009).
Schmidt, M. Xenobiology: a new form of life as the ultimate biosafety tool. Bioessays 32, 322–331 (2010).
Lagoja, I.M., Marchand, A., Van Aerschot, A. & Herdewijn, P. Synthesis of 1,5-anhydrohexitol building blocks for oligonucleotide synthesis. Curr. Protoc. Nucleic Acid Chem. 14, 1.9.1–1.9.22 (2003).
Zlatev, I., Manoharan, M., Vasseur, J.-J. & Morvan, F. Solid-phase chemical synthesis of 5-triphosphate DNA, RNA, and chemically modified oligonucleotides. Curr. Protoc. Nucleic Acid Chem. 50, 1.28.1–1.28.16 (2012).
Deleavey, G.F. et al. Synergistic effects between analogs of DNA and RNA improve the potency of siRNA-mediated gene silencing. Nucleic Acids Res. 38, 4547–4557 (2010).
Chu, B.C., Wahl, G.M. & Orgel, L.E. Derivatization of unprotected polynucleotides. Nucleic Acids Res. 11, 6513–6529 (1983).
Breaker, R.R. & Joyce, G.F. A DNA enzyme that cleaves RNA. Chem. Biol. 1, 223–229 (1994).
Breaker, R.R. & Joyce, G.F. A DNA enzyme with Mg2+-dependent RNA phosphoesterase activity. Chem. Biol. 2, 655–660 (1995).
Elshire, R.J., Glaubitz, J.C., Sun, Q. & Poland, J.A. A robust, simple genotyping-by-sequencing (GBS) approach for high-diversity species. PLoS ONE 6, e19379 (2011).
Brody, J.R. & Kern, S.E. Sodium boric acid: a tris-free, cooler conductive medium for DNA electrophoresis. Biotechniques 36, 214–216 (2004).
Tolle, F., Wilke, J., Wengel, J. & Mayer, G. By-product formation in repetitive PCR amplification of DNA libraries during SELEX. PLoS ONE 9, e114693 (2014).
Shao, K. et al. Emulsion PCR: a high efficient way of PCR amplification of random DNA libraries in aptamer selection. PLoS ONE 6, e24910 (2011).
Matochko, W.L., Cory Li, S., Tang, S.K.Y. & Derda, R. Prospective identification of parasitic sequences in phage display screens. Nucleic Acids Res. 42, 1784–1798 (2014).
Goecks, J., Nekrutenko, A., Taylor, J. & Galaxy Team Galaxy: a comprehensive approach for supporting accessible, reproducible, and transparent computational research in the life sciences. Genome Biol. 11, R86 (2010).
Blankenberg, D. et al. Galaxy: a web-based genome analysis tool for experimentalists. Curr. Protoc. Mol. Biol. 89, 19.10.1 19.10. 21 (2010).
Giardine, B. et al. Galaxy: a platform for interactive large-scale genome analysis. Genome Res. 15, 1451–1455 (2005).
Alam, K.K., Chang, J.L. & Burke, D.H. FASTAptamer: a bioinformatic toolkit for high-throughput sequence analysis of combinatorial selections. Mol. Ther. Nucleic Acids 4, e230 (2015).
Elle, I.C. et al. Selection of LNA-containing DNA aptamers against recombinant human CD73. Mol. Biosyst. 11, 1260–1270 (2015).
Acknowledgements
The authors thank J. Sutherland for helpful discussions. This work was supported by the UK Medical Research Council (MRC) programme grant U105178804 and by grants from the European Science Foundation (ESF) and the UK Biotechnology and Biological Sciences Research Council (BBSRC; 09-EuroSYNBIO-OP-013).
Author information
Authors and Affiliations
Contributions
A.I.T. and P.H. conceived and designed the protocols, and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Taylor, A., Holliger, P. Directed evolution of artificial enzymes (XNAzymes) from diverse repertoires of synthetic genetic polymers. Nat Protoc 10, 1625–1642 (2015). https://doi.org/10.1038/nprot.2015.104
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2015.104
This article is cited by
-
A two-residue nascent-strand steric gate controls synthesis of 2′-O-methyl- and 2′-O-(2-methoxyethyl)-RNA
Nature Chemistry (2023)
-
XNAzymes targeting the SARS-CoV-2 genome inhibit viral infection
Nature Communications (2022)
-
A modular XNAzyme cleaves long, structured RNAs under physiological conditions and enables allele-specific gene silencing
Nature Chemistry (2022)
-
Recent advances of functional nucleic acid-based sensors for point-of-care detection of SARS-CoV-2
Microchimica Acta (2022)
-
Organ-on-a-Chip: The Future of Therapeutic Aptamer Research?
BioChip Journal (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.