Abstract
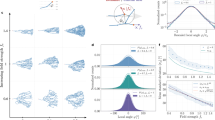
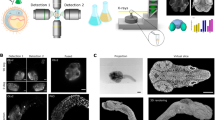
Developmental branching morphogenesis establishes organ architecture, and it is driven by iterative interactions between epithelial and mesenchymal progenitor cell populations. We describe an approach for analyzing this interaction and how it contributes to organ development. After initial in vivo cell labeling with the nucleoside analog 5-ethynyl-2′-deoxyuridine (EdU) and tissue-specific antibodies, optical projection tomography (OPT) and confocal microscopy are used to image the developing organ. These imaging data then inform a second analysis phase that quantifies (using Imaris and Tree Surveyor software), models and integrates these events at a cell and tissue level in 3D space and across developmental time. The protocol establishes a benchmark for assessing the impact of genetic change or fetal environment on organogenesis that does not rely on ex vivo organ culture or section-based reconstruction. By using this approach, examination of two developmental stages for an organ such as the kidney can be undertaken by a postdoctoral-level researcher in 6 weeks, with a full developmental analysis in mouse achievable in 5 months.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout











Similar content being viewed by others
References
Herriges, M. & Morrisey, E.E. Lung development: orchestrating the generation and regeneration of a complex organ. Development 141, 502–513 (2014).
Costantini, F. & Kopan, R. Patterning a complex organ: branching morphogenesis and nephron segmentation in kidney development. Dev. Cell 18, 698–712 (2010).
Huebner, R.J. & Ewald, A.J. Cellular foundations of mammary tubulogenesis. Semin. Cell Dev. Biol. 31, 24–31 (2014).
Patel, V.N. & Hoffman, M.P. Salivary gland development: a template for regeneration. Semin. Cell Dev. Biol. 25–26C, 52–60 (2014).
Grobstein, C. Inductive epithelio-mesenchymal interaction in cultured organ rudiments of the mouse. Science 118, 52–55 (1953).
Grobstein, C. Inductive interaction in the development of the mouse metanephros. J. Exp. Zool. 130, 319–339 (1955).
Short, K.M., Hodson, M.J. & Smyth, I.M. Tomographic quantification of branching morphogenesis and renal development. Kidney Int. 77, 1132–1139 (2010).
al-Awqati, Q. & Goldberg, M.R. Architectural patterns in branching morphogenesis in the kidney. Kidney Int. 54, 1832–1842 (1998).
Cebrian, C., Borodo, K., Charles, N. & Herzlinger, D.A. Morphometric index of the developing murine kidney. Dev. Dyn. 231, 601–608 (2004).
Sims-Lucas, S. et al. Three-dimensional imaging reveals ureteric and mesenchymal defects in Fgfr2-mutant kidneys. J. Am. Soc. Nephrol. 20, 2525–2533 (2009).
Hoy, W.E. et al. Nephron number, glomerular volume, renal disease and hypertension. Curr. Opin. Nephrol. Hypertens. 17, 258–265 (2008).
Luyckx, V.A. et al. Effect of fetal and child health on kidney development and long-term risk of hypertension and kidney disease. Lancet 382, 273–283 (2013).
Boyle, S. et al. Fate mapping using Cited1-CreERT2 mice demonstrates that the cap mesenchyme contains self-renewing progenitor cells and gives rise exclusively to nephronic epithelia. Dev. Biol. 313, 234–245 (2008).
Kobayashi, A. et al. Six2 defines and regulates a multipotent self-renewing nephron progenitor population throughout mammalian kidney development. Cell Stem Cell 3, 169–181 (2008).
El Agha, E. et al. Fgf10-positive cells represent a progenitor cell population during lung development and postnatally. Development 141, 296–306 (2014).
Cunha, G.R. et al. Mammary phenotypic expression induced in epidermal cells by embryonic mammary mesenchyme. Acta Anat. (Basel) 152, 195–204 (1995).
Short, K., Hodson, M. & Smyth, I. Spatial mapping and quantification of developmental branching morphogenesis. Development 140, 471–478 (2013).
Short, K.M. & Smyth, I.M. Analysis of native kidney structures in three dimensions. Methods Mol. Biol. 886, 95–107 (2012).
Lefevre, J. et al. Modelling cell turnover in a complex tissue during development. J. Theor. Biol. 338, 66–79 (2013).
Packard, A. et al. Luminal mitosis drives epithelial cell dispersal within the branching ureteric bud. Dev. Cell 27, 319–330 (2013).
Rumballe, B.A. et al. Nephron formation adopts a novel spatial topology at cessation of nephrogenesis. Dev. Biol. 360, 110–122 (2011).
Combes, A.N. et al. Three-dimensional visualization of testis cord morphogenesis, a novel tubulogenic mechanism in development. Dev. Dyn. 238, 1033–1041 (2009).
Combes, A.N. et al. Endothelial cell migration directs testis cord formation. Dev. Biol. 326, 112–120 (2009).
Davies, J.A. et al. Access and use of the GUDMAP database of genitourinary development. Methods Mol. Biol. 886, 185–201 (2012).
Georgas, K.M., Chiu, H.S., Lesieur, E., Rumballe, B.A. & Little, M.H. Expression of metanephric nephron-patterning genes in differentiating mesonephric tubules. Dev. Dyn. 240, 1600–1612 (2011).
Harding, S.D. et al. The GUDMAP database—an online resource for genitourinary research. Development 138, 2845–2853 (2011).
Little, M.H. et al. A high-resolution anatomical ontology of the developing murine genitourinary tract. Gene Expr. Patterns 7, 680–699 (2007).
McMahon, A.P. et al. GUDMAP: the genitourinary developmental molecular anatomy project. J. Am. Soc. Nephrol. 19, 667–671 (2008).
Short, K.M. et al. Global quantification of tissue dynamics in the developing mouse kidney. Dev. Cell 29, 188–202 (2014).
Sharpe, J. et al. Optical projection tomography as a tool for 3D microscopy and gene expression studies. Science 296, 541–545 (2002).
Schuchardt, A., D'Agati, V., Pachnis, V. & Costantini, F. Renal agenesis and hypodysplasia in ret-k– mutant mice result from defects in ureteric bud development. Development 122, 1919–1929 (1996).
Enomoto, H. et al. RET signaling is essential for migration, axonal growth and axon guidance of developing sympathetic neurons. Development 128, 3963–3974 (2001).
Schuchardt, A., D'Agati, V., Larsson-Blomberg, L., Costantini, F. & Pachnis, V. Defects in the kidney and enteric nervous system of mice lacking the tyrosine kinase receptor Ret. Nature 367, 380–383 (1994).
Metzger, R.J., Klein, O.D., Martin, G.R. & Krasnow, M.A. The branching programme of mouse lung development. Nature 453, 745–750 (2008).
Hockendorf, B., Thumberger, T. & Wittbrodt, J. Quantitative analysis of embryogenesis: a perspective for light sheet microscopy. Dev. Cell 23, 1111–1120 (2012).
Francois, M. et al. Segmental territories along the cardinal veins generate lymph sacs via a ballooning mechanism during embryonic lymphangiogenesis in mice. Dev. Biol. 364, 89–98 (2012).
Livet, J. et al. Transgenic strategies for combinatorial expression of fluorescent proteins in the nervous system. Nature 450, 56–62 (2007).
Chung, K. & Deisseroth, K. CLARITY for mapping the nervous system. Nat. Methods 10, 508–513 (2013).
Becker, K., Jahrling, N., Saghafi, S., Weiler, R. & Dodt, H.U. Chemical clearing and dehydration of GFP expressing mouse brains. PLoS ONE 7, e33916 (2012).
Kuwajima, T. et al. ClearT: a detergent- and solvent-free clearing method for neuronal and non-neuronal tissue. Development 140, 1364–1368 (2013).
Hornblad, A., Cheddad, A. & Ahlgren, U. An improved protocol for optical projection tomography imaging reveals lobular heterogeneities in pancreatic islet and beta cell mass distribution. Islets 3, 204–208 (2011).
Theiler, K. The House Mouse: Atlas of Embryonic Development 2nd edn. (Springer-Verlag, 1989).
Wanek, N., Muneoka, K., Holler-Dinsmore, G., Burton, R. & Bryant, S.V. A staging system for mouse limb development. J. Exp. Zool. 249, 41–49 (1989).
Nagy, A., Gertsenstein, M., Vintersten, K. & Behringer, R. Alcian blue staining of the mouse fetal cartilaginous skeleton. Cold Spring Harbor Protoc. 2009 10.1101/pdb.prot5169 (2009).
McDonald, J.H. Handbook of Biological Statistics 161–168 (Sparky House Publishing, 2009).
Welch, B.L. The generalization of 'Student's' problem when several different population variances are involved. Biometrika 34, 28–35 (1947).
Buck, S.B. et al. Detection of S-phase cell cycle progression using 5-ethynyl-2′-deoxyuridine incorporation with click chemistry, an alternative to using 5-bromo-2′-deoxyuridine antibodies. BioTechniques 44, 927–929 (2008).
Salic, A. & Mitchison, T.J. A chemical method for fast and sensitive detection of DNA synthesis in vivo. Proc. Natl. Acad. Sci. USA 105, 2415–2420 (2008).
Zeng, C. et al. Evaluation of 5-ethynyl-2′-deoxyuridine staining as a sensitive and reliable method for studying cell proliferation in the adult nervous system. Brain Res. 1319, 21–32 (2010).
Farah, M.H. Cumulative labeling of embryonic mouse neural retina with bromodeoxyuridine supplied by an osmotic minipump. J. Neurosci. Methods 134, 169–178 (2004).
Alanentalo, T. et al. High-resolution three-dimensional imaging of islet-infiltrate interactions based on optical projection tomography assessments of the intact adult mouse pancreas. J. Biomed. Optics 13, 054070 (2008).
Walls, J.R., Coultas, L., Rossant, J. & Henkelman, R.M. Three-dimensional analysis of vascular development in the mouse embryo. PLoS ONE 3, e2853 (2008).
Acknowledgements
This work was supported by the National Health and Medical Research Council of Australia (APP1002748, APP1022239 and APP1063989), the Australian Research Council (DP10310086) and the Human Frontiers in Science Program (RGP0039/2011). Microscopy was performed at the Australian Cancer Research Foundation (ARCF)/Institute for Molecular Bioscience Cancer Biology Imaging Facility, which was established with the support of the ARCF. I.M.S. holds a Future Fellowship from the Australian Research Council. M.H.L. is a Senior Principal Research Fellow of the Australian National Health and Medical Research Council.
Author information
Authors and Affiliations
Contributions
A.N.C. and J.L. developed the methodologies used in analyzing confocal data, cell cycle length and population modeling. K.M.S. developed the Tree Surveyor protocol and tools to examine tree morphometrics. I.M.S. undertook the correlative microscopy experiments. N.A.H. oversaw and contributed to the development of approaches to mathematical analyses. I.M.S. and M.H.L. were responsible for the initial conception of the project, directed the collaboration and were involved in the design of experiments, ongoing planning, and analysis and interpretation of the data. K.M.S., A.N.C., J.L. and I.M.S. wrote the manuscript with substantive input from M.H.L. and N.A.H.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Whole Organ Confocal Z-stack.
Confocal imaging data from a whole 14.5 dpc kidney demonstrate that the quality of imaging can be maintained throughout the sample. Panels show every 10th optical slice throughout the Z-stack (depth from the fist slice is indicated in each panel). The whole file is composed of 101 Z-slices that encompass a total thickness of 352 microns. Scale bar = 100 μm.
Supplementary information
Supplementary Figure 1 and Supplementary Manual
(PDF 213 kb)
Introduction to the Tree Surveyor application.
The movie explains the interface, project setup, loading data, and an explanation of the algorithms used and the sliders and buttons which control them. (MOV 6995 kb)
A guide to running a Tree Surveyor process.
Discussed are the processing algorithms, the assessment of the first stage of skeletonization, editing of the tree and placement of root. (MOV 6082 kb)
Completion of processing and export of data from Tree Surveyor.
Visualisation modes, explanation of segment/branch naming conventions and introduction to data export and analysis. (MOV 7006 kb)
Supplementary Data 1: Cell Modeling.
This zip file contains an excel spreadsheet (Supp Data 1 worksheet.xl) and a Word document with instructions for its use (Supp Data 1 instructions.doc). The excel spreadsheet contains the formulas required to determine whole organ estimates and to model population growth and exit from cell niches. (ZIP 33 kb)
Supplementary Data 2: An example OPT z-stack.
This data is from a normal E14.5 C57BL/6 embryonic kidney with ureteric tree detected with Trop-2 antibody. The stack can be used as a guide to test Tree Surveyor usage and analysis. On analysis, this sample should have approximately 327 tips, total length of ∼44 μm, a convex hull volume of ∼0.56 mm3, and tips with an approximate mean length of 56 μm. (ZIP 8367 kb)
Supplementary Data 3: Example confocal dataset.
An example of a high resolution 40× confocal z-stack of a normal E15.5 kidney (Zeiss LSM format, compatible with Imaris). The stack contains data from four channels marking nuclei (blue), cap mesenchyme (red), Cytokeratin (white), GFP (green). This can be used as a guide to test the included protocols in Imaris. (ZIP 51871 kb)
Supplementary Software: R Studio Script to model EdU incorporation.
This file provides code and demonstration data for EdU incorporation curve modelling. An R Studio script (curvefitting.R) calculates cell cycle length estimates and standard errors, and plots modelled curves. The script has example data embedded which can be replaced. Instructions for use are embedded in the script. (ZIP 5 kb)
Rights and permissions
About this article
Cite this article
Combes, A., Short, K., Lefevre, J. et al. An integrated pipeline for the multidimensional analysis of branching morphogenesis. Nat Protoc 9, 2859–2879 (2014). https://doi.org/10.1038/nprot.2014.193
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2014.193
This article is cited by
-
Lin28 and let-7 regulate the timing of cessation of murine nephrogenesis
Nature Communications (2019)
-
The contribution of branching morphogenesis to kidney development and disease
Nature Reviews Nephrology (2016)
-
A morphological investigation of sexual and lateral dimorphism in the developing metanephric kidney
Scientific Reports (2015)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.