Abstract
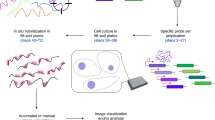
RNA-sequencing (RNA-seq) measures the quantitative change in gene expression over the whole transcriptome, but it lacks spatial context. In contrast, in situ hybridization provides the location of gene expression, but only for a small number of genes. Here we detail a protocol for genome-wide profiling of gene expression in situ in fixed cells and tissues, in which RNA is converted into cross-linked cDNA amplicons and sequenced manually on a confocal microscope. Unlike traditional RNA-seq, our method enriches for context-specific transcripts over housekeeping and/or structural RNA, and it preserves the tissue architecture for RNA localization studies. Our protocol is written for researchers experienced in cell microscopy with minimal computing skills. Library construction and sequencing can be completed within 14 d, with image analysis requiring an additional 2 d.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Rifai, N., Gillette, M.A. & Carr, S.A. Protein biomarker discovery and validation: the long and uncertain path to clinical utility. Nat. Biotechnol. 24, 971–983 (2006).
Jaitin, D.A. et al. Massively parallel single-cell RNA-seq for marker-free decomposition of tissues into cell types. Science 343, 776–779 (2014).
Battich, N., Stoeger, T. & Pelkmans, L. Image-based transcriptomics in thousands of single human cells at single-molecule resolution. Nat. Methods 10, 1127–1133 (2013).
Lein, E.S. et al. Genome-wide atlas of gene expression in the adult mouse brain. Nature 445, 168–176 (2007).
Zeng, H. et al. Large-scale cellular-resolution gene profiling in human neocortex reveals species-specific molecular signatures. Cell 149, 483–496 (2012).
Diez-Roux, G. et al. A high-resolution anatomical atlas of the transcriptome in the mouse embryo. PLoS Biol. 9, e1000582 (2011).
Femino, A.M., Fay, F.S., Fogarty, K. & Singer, R.H. Visualization of single RNA transcripts in situ. Science 280, 585–590 (1998).
Raj, A., van den Bogaard, P., Rifkin, S.A., van Oudenaarden, A. & Tyagi, S. Imaging individual mRNA molecules using multiple singly labeled probes. Nat. Methods 5, 877–879 (2008).
Levsky, J.M., Shenoy, S.M., Pezo, R.C. & Singer, R.H. Single-cell gene expression profiling. Science 297, 836–840 (2002).
Lubeck, E., Coskun, A.F., Zhiyentayev, T., Ahmad, M. & Cai, L. Single-cell in situ RNA profiling by sequential hybridization. Nat. Methods 11, 360–361 (2014).
Choi, H.M. et al. Programmable in situ amplification for multiplexed imaging of mRNA expression. Nat. Biotechnol. 28, 1208–1212 (2011).
Ke, R. et al. In situ sequencing for RNA analysis in preserved tissue and cells. Nat. Methods 10, 857–860 (2013).
Larsson, C., Grundberg, I., Söderberg, O. & Nilsson, M. In situ detection and genotyping of individual mRNA molecules. Nat. Methods 7, 395–397 (2010).
Larsson, C. et al. In situ genotyping individual DNA molecules by target-primed rolling-circle amplification of padlock probes. Nat. Methods 1, 227–232 (2004).
Lagunavicius, A. et al. Novel application of Phi29 DNA polymerase: RNA detection and analysis in vitro and in situ by target RNA-primed RCA. RNA 15, 765–771 (2009).
Merkiene, E., Gaidamaviciute, E., Riauba, L., Janulaitis, A. & Lagunavicius, A. Direct detection of RNA in vitro and in situ by target-primed RCA: the impact of E. coli RNase III on the detection efficiency of RNA sequences distanced far from the 3′-end. RNA 16, 1508–1515 (2010).
Lee, J.H. et al. A robust approach to identifying tissue-specific gene expression regulatory variants using personalized human induced pluripotent stem cells. PLoS Genet. 5, e1000718 (2009).
Shendure, J. et al. Accurate multiplex polony sequencing of an evolved bacterial genome. Science 309, 1728–1732 (2005).
Drmanac, R. et al. Human genome sequencing using unchained base reads on self-assembling DNA nanoarrays. Science 327, 78–81 (2010).
Grun, D., Kester, L. & van Oudenaarden, A. Validation of noise models for single-cell transcriptomics. Nat. Methods 11, 637–640 (2014).
Shapiro, E., Biezuner, T. & Linnarsson, S. Single-cell sequencing-based technologies will revolutionize whole-organism science. Nat. Rev. Genet. 14, 618–630 (2013).
Islam, S. et al. Quantitative single-cell RNA-seq with unique molecular identifiers. Nat. Methods 11, 163–166 (2014).
Lee, J.H. et al. Highly multiplexed subcellular RNA sequencing in situ. Science 343, 1360–1363 (2014).
Adiconis, X. et al. Comparative analysis of RNA sequencing methods for degraded or low-input samples. Nat. Methods 10, 623–629 (2013).
Yachida, S. et al. Distant metastasis occurs late during the genetic evolution of pancreatic cancer. Nature 467, 1114–1117 (2010).
Frumkin, D. et al. Amplification of multiple genomic loci from single cells isolated by laser micro-dissection of tissues. BMC Biotechnol. 8, 17 (2008).
Lovatt, D. et al. Transcriptome in vivo analysis (TIVA) of spatially defined single cells in live tissue. Nat. Methods 11, 190–196 (2014).
Schmid, M.W. et al. A powerful method for transcriptional profiling of specific cell types in eukaryotes: laser-assisted microdissection and RNA sequencing. PLoS ONE 7, e29685 (2012).
Islam, S. et al. Characterization of the single-cell transcriptional landscape by highly multiplex RNA-seq. Genome Res. 21, 1160–1167 (2011).
Ramskold, D. et al. Full-length mRNA-seq from single-cell levels of RNA and individual circulating tumor cells. Nat. Biotechnol. 30, 777–782 (2012).
Hashimshony, T., Wagner, F., Sher, N. & Yanai, I. CEL-seq: single-cell RNA-seq by multiplexed linear amplification. Cell Rep. 2, 666–673 (2012).
Avital, G., Hashimshony, T. & Yanai, I. Seeing is believing: new methods for in situ single-cell transcriptomics. Genome Biol. 15, 110 (2014).
Buxbaum, A.R., Wu, B. & Singer, R.H. Single β-actin mRNA detection in neurons reveals a mechanism for regulating its translatability. Science 343, 419–422 (2014).
Hanna, J. et al. Direct cell reprogramming is a stochastic process amenable to acceleration. Nature 462, 595–601 (2009).
Buganim, Y. et al. Single-cell expression analyses during cellular reprogramming reveal an early stochastic and a late hierarchic phase. Cell 150, 1209–1222 (2012).
Itzkovitz, S., Blat, I.C., Jacks, T., Clevers, H. & van Oudenaarden, A. Optimality in the development of intestinal crypts. Cell 148, 608–619 (2012).
Hansen, C.H. & van Oudenaarden, A. Allele-specific detection of single mRNA molecules in situ. Nat. Methods 10, 869–871 (2013).
Porreca, G.J. et al. Multiplex amplification of large sets of human exons. Nat. Methods 4, 931–936 (2007).
Kosuri, S. et al. Composability of regulatory sequences controlling transcription and translation in Escherichia coli. Proc. Natl. Acad. Sci. USA 110, 14024–14029 (2013).
Li, J.B. et al. Genome-wide identification of human RNA editing sites by parallel DNA capturing and sequencing. Science 324, 1210–1213 (2009).
Zhang, K. et al. Digital RNA allelotyping reveals tissue-specific and allele-specific gene expression in human. Nat. Methods 6, 613–618 (2009).
Michael, W.M., Choi, M. & Dreyfuss, G. A nuclear export signal in hnRNP A1: a signal-mediated, temperature-dependent nuclear protein export pathway. Cell 83, 415–422 (1995).
Kaposi-Novak, P., Lee, J.S., Mikaelyan, A., Patel, V. & Thorgeirsson, S.S. Oligonucleotide microarray analysis of aminoallyl-labeled cDNA targets from linear RNA amplification. Biotechniques 37, 580, 582–586, 588 (2004).
Nanda, J.S. & Lorsch, J.R. Labeling a protein with fluorophores using NHS ester derivitization. Methods Enzymol. 536, 87–94 (2014).
Mardis, E.R. Next-generation DNA sequencing methods. Annu. Rev. Genomics Hum. Genet. 9, 387–402 (2008).
Massingham, T. & Goldman, N. Error-correcting properties of the SOLiD Exact Call Chemistry. BMC Bioinformatics 13, 145 (2012).
Applied Biosystems. SOLiD System Accuracy with the Exact Call Chemistry Module (https://www3.appliedbiosystems.com/cms/groups/global_marketing_group/documents/generaldocuments/cms_091372.pdf).
Itzkovitz, S. & van Oudenaarden, A. Validating transcripts with probes and imaging technology. Nat. Methods 8, S12–S19 (2011).
Eliceiri, K.W. et al. Biological imaging software tools. Nat. Methods 9, 697–710 (2012).
Schindelin, J. et al. Fiji: an open-source platform for biological-image analysis. Nat. Methods 9, 676–682 (2012).
Kankaanpaa, P. et al. BioImageXD: an open, general-purpose and high-throughput image-processing platform. Nat. Methods 9, 683–689 (2012).
Pawley, J.B. Handbook of Biological Confocal Microscopy 3rd edn. (Springer, 2006).
Acknowledgements
This study was funded by US National Institutes of Health (NIH) Centers of Excellence in Genomic Sciences (CEGS) grant no. P50 HG005550. J.H.L. and co-workers were funded by National Heart, Blood and Lung Institute (NHBLI) grant no. RC2HL102815, by the Allen Institute for Brain Science and by National Institute of Mental Health (NIMH) grant no. MH098977. E.R.D. was funded by NIH grant no. GM080177 and by National Science Foundation (NSF) Graduate Research Fellowship grant no. DGE1144152.
Author information
Authors and Affiliations
Contributions
J.H.L. and E.R.D. conceived FISSEQ library construction, sequencing, image analysis and bioinformatics. J.S., R.K., J.L.Y., B.M.T., H.S.L. and J.A. provided key feedbacks during the FISSEQ method development. R.T. and T.C.F. assisted with automated microscopy and image analysis. K.Z. and G.M.C. oversaw the project. J.H.L. wrote the paper, and E.R.D. wrote the FISSEQ software.
Corresponding authors
Ethics declarations
Competing interests
Potential conflicts of interest for G.M.C. are listed on http://arep.med.harvard.edu/gmc/tech.html. Other authors have no conflicts of interest.
Integrated supplementary information
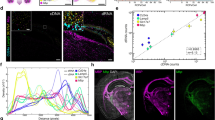
Supplementary Figure 1 The altered distribution of poly A-associated transcripts at different fixation temperatures.
(a) Primary fibroblasts were fixed at room temperature or at 37°C for 10 minutes. The cells were then hybridized to the Cy3 Poly dT(50) or 18S rRNA probe. At room temperature a large fraction of the mRNA is retained in the nucleus. (b) The cold shock-induced nuclear retention of poly A-associated transcripts is cell type-specific. Highly transformed cell lines and iPS cell lines do not appear to be affected by the fixation temperature.
Supplementary Figure 2 Acid-treatment improves cell permeabilization for FISSEQ.
(a) Low amplicon counts are frequently due to poor permeabilization, which can be observed in the axial view of human iPS cells. (b) A brief treatment with 0.1 HCl can significantly improve the uniformity of FISSEQ amplicons throughout the cell, especially in iPS cells.
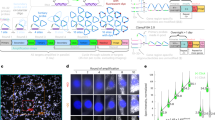
Supplementary Figure 3 Degradation of residual RNA is essential for FISSEQ amplicon generation.
Far more amplicons are observed after the RNase-treatment before circularization in numerous human iPS cell colonies (labeled with mCherry; bar: 1 μm).
Supplementary Figure 4 Image deconvolution of FISSEQ amplicons in fibroblasts.
(a) A high quality confocal image requires minimal image deconvolution, but doing so can dramatically increase the signal to noise ratio by removing many spurious pixels; however, over-deconvolution can actually lead to more background noise or artifacts (bar: 1 μm). (b) A low quality epifluorescence image can contain a significant amount of out of focus light. Using 3D deconvolution to remove such signal can dramatically improve the image and speed up base calling (bar: 1 μm).
Supplementary Figure 5 Image alignment of FISSEQ amplicons in fibroblasts.
(a) One of the adjustable parameter is the block size for local alignment. For large images (i.e. 4000-by-4000 pixels) dividing images into a 10-by-10 grid is essential. Even for small images one can appreciate the effect of increasing the block number. (b) When using local alignment it is important to specify the degree of block overlap for image stitching. A large image correction with insufficient or no overlap will result in image fragmentation. (c) In general sub-pixel registration accuracy will not affect alignment significantly; however, increasing the upsampling factor can lead to better registration in certain instances.
Supplementary Figure 6 Data analysis using RStudio.
(a) RStudio provides a straightforward GUI for importing, analyzing and visualizing FISSEQ data. We provide several sample datasets and R sessions for FISSEQ data analysis. One can view the history window and click through commands to reproduce the following plots. (b) Correlation plot of gene expression frequency in region 1 and region 2. (c) Frequency histogram of RNA class instances. (d) Distribution of cluster sizes. (e) Area plot of non-ribosomal reads and their location from 5 different wound healing regions.
Supplementary information
Supplementary Information
Supplementary Figures 1–6 (PDF 2210 kb)
Rights and permissions
About this article
Cite this article
Lee, J., Daugharthy, E., Scheiman, J. et al. Fluorescent in situ sequencing (FISSEQ) of RNA for gene expression profiling in intact cells and tissues. Nat Protoc 10, 442–458 (2015). https://doi.org/10.1038/nprot.2014.191
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2014.191
This article is cited by
-
Matisse: a MATLAB-based analysis toolbox for in situ sequencing expression maps
BMC Bioinformatics (2021)
-
The spatial RNA integrity number assay for in situ evaluation of transcriptome quality
Communications Biology (2021)
-
Organoids in image-based phenotypic chemical screens
Experimental & Molecular Medicine (2021)
-
Microdroplet-based one-step RT-PCR for ultrahigh throughput single-cell multiplex gene expression analysis and rare cell detection
Scientific Reports (2021)
-
ClusterMap for multi-scale clustering analysis of spatial gene expression
Nature Communications (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.